The mystery of orphan genes
Interview with
Orphan genes - sections of protein-coding DNA that appear to have no detectable homology with peptides from genomes of other similar species. So, how do new or orphan genes crop up in a population and where do they come from?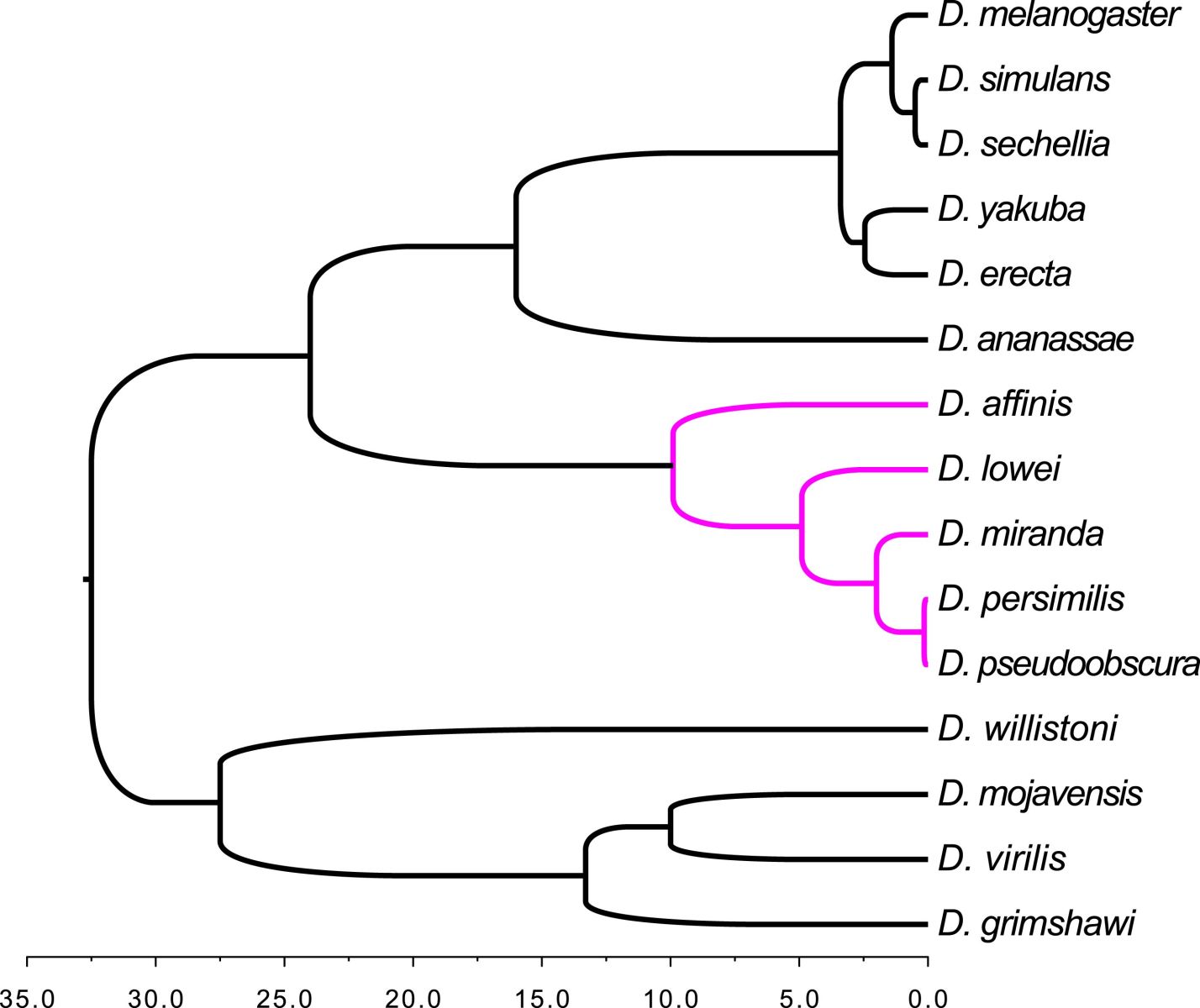
Christian - My name is Christian Schlotterer. I work at Vetmed University in Vienna. I'm a population geneticist and my interest is understanding adaptation and innovation that helps populations to adapt to changing environments. That novel genes can arise at a high rate has been well-documented in the literature, but if new genes arise at a high rate, why doesn't the number of genes increase over time? Because that's not what you see.
Chris - And how do these orphan genes or new genes come about? What's their origin?
Christian - They arise from previously non-coding DNA. Basically, there is DNA there and is not coding for protein and then suddenly, it can acquire a few mutations and then it gets an open reading frame and it suddenly has a function.
Chris - So, at what rate are these orphan genes appearing? If you look in fruit flies for example, in each generation, how many do you expect to see?
Christian - You don't see them in generations. You need hundreds of years...
Chris - So, how did you actually approach this problem then, given that you've got that long time scale, they're appearing over hundreds of years to thousands of years? How did you do the study you've done?
Christian - Well, I should say, maybe taking one step back, how have the orphans been detected beforehand? People were looking at basically isolated phylogenetic images. They didn't look at closely related species. So, what is novel in our study is that we looked at closely related species that were close enough that we could detect these novel genes there as well. So, what we did, we used the set of related species, we looked at relatives within increasing divergence age. So, they were species that split from a common ancestor at different time points in the past. The key thing we were not so interested in was the origin of these new genes. We were interested in basically asking what is the persistence time, how long do they stay around.
Chris - In other words, by looking at an orphan gene that's cropped up, but then subsequently disappeared, you can ask, well how long do they persist in each given arm of the population or each individual species' branch. And this gives you some insight into the persistence time of those orphan genes, and you can then ask whether some persist for longer or not.
Christian - Exactly. What we found is that there are several indicators that give you a hint when genes are persisting for a longer time, one will see expression intensity. So, if a gene is more highly expressed at a higher level then this gene is more likely to be persisting. At the same time, we also find that if the gene has more expression in males than in females, then this gene also has a higher chance to persist over longer evolutionary timescales than genes that have no bias.
Chris - Well, how do you account for that then, given that you have seen that and if you think that it's a surprise that those genes should cling on? Why do you think they do?
Christian - Well, that's a good question. I wish I would have a good answer to this. One possibility could be that we can distinguish between spurious transcription and genuine transcription because the way we identify whether a gene is basically a real gene is based on transcription only. And then we look at parts of the gene that are transcribed and have an open reading frame, this means they have a capacity to encode for a protein, and then these are basically classified as orphans. Nevertheless, it's well-known that parts of the genome are just randomly transcribed, and so, it could be that if you have a strong expression, this indicates that you really look at the genuine signal rather than maybe a spurious signal. I'm pretty sure, what we have classified as orphans, there will be some noise in there that may not actually be real orphans.
Chris - What about the sex bias though as well because that's also very intriguing? Why should there be a favour on the male's side to keep these things going?
Christian - Obviously, you caught me on the wrong foot. I have no good explanation. There wasn't an explanation why you could think that new genes, when they originate among male-biased because the idea was that you need simpler architecture of the regulatory module that's basically is responsible for the expression pattern of a gene. But our data basically focuses upon this hypothesis. So, we're left with an observation that's very difficult to understand.
Chris - Broadly, what do you think the implications are of what you've discovered? How would you - in a nutshell - say, this is important because...?
Christian - The importance of novel genes has for a long time been neglected. And so, I think, really to understand novelty in evolution, these new genes are probably one of the key factors that we need to understand better. Our study has made a major step in showing that these genes can persist for some time, but not all of them do. These novel genes, we applied population genetic tests, we see that there is selection preventing mutations occurring in these genes, and those are the best indication that there's something really functional.










Comments
Add a comment