Almost every single cell of your body is packed with more than two metres of DNA, containing your genes. But not only does it have to be packed up to fit in there, it also has to be organised and read. Plus, how genetic variations are linked to cancer risk,analysing dinosaur DNA, and an adventurous gene of the month.
In this episode
01:06 - Prof Wendy Bickmore - Packing DNA
Prof Wendy Bickmore - Packing DNA
with Professor Wendy Bickmore, MRC Human Genetics unit, Edinburgh
This year is the 60th anniversary of the discovery of the structure of DNA by James Watson, Francis Crick and Maurice Wilkins, aided by Rosalind Franklin. We're all familiar with images of the DNA double helix from school textbooks, newspapers and even company logos. But the reality is a lot more complicated, as more than two metres of this elegant twisted ladder has to be squished into the nucleus of each one of our cells, and then it has to be organised, copied and used.
To find out more about how our cells achieve this incredible feat of packing, I spoke to Professor Wendy Bickmore from the MRC Human Genetics Unit in Edinburgh.
Wendy - We used to know what a gene was. We could identify it from the sequence of bases in the DNA because they obviously coded for proteins. So, we could the start of a gene which we call a gene promoter and then the bit of the gene that codes for the protein. And so, we can say, the gene starts here and it ends here. But now, what we realise all of a sudden is actually, that isn't all the gene comprises. A gene needs that core bit of sequence that codes for the protein and the promoter that's going to drive the transcription of that bit of DNA into RNA, but it also needs an increasingly complex surrounding region of the genome to help control when and where, and how strongly that gene is expressed in development.
Kat - Because obviously, we have somewhere between 20,000 and 30,000 genes. They're not just on all the time in all our cells.
Wendy - No. I think one of the - maybe disappointing things in the sequencing of the human genome was that actually, we didn't have very many more gene than a worm has and I guess we felt affronted by that. We thought we were a bit more complicated. So, the realisation since that time has actually been, it's not how many genes you've got, but what you do with them, how you control them.
Kat - Not what you got, it's what you do with it that counts.
Wendy - Absolutely, yes.
Kat - So, if you think about these 30,000 genes scattered through the millions and millions of base pairs of DNA that we have that it adds up in every cell of our body to about 2 metres of DNA. How is it organised? How do you compress all that into the nucleus of a single cell and then start to switch genes on and off in the right time and the right place?
Wendy - We actually think the two problems are actually intimately connected. So, if you think about the 2 metres, your DNA that's going to fit inside the cell nucleus which is probably about on average 10 millionth of one metre in diameter, that's a kind of a big packaging problem and it's about more than a thousand fold of condensation of the DNA sequence you've got to do. And people used to think, "Well, that's just an issue of space and you've got to get in the space, and you just squish it down."
Now, we realise actually, it's how you do that squishing and folding that helps to control whether a gene is able to be activated and read in a particular cell type and not in another one. So, I think if you think of the - you know, people make the analogy of the human genome as a book - encyclopaedia or instructions of how to make an organisms - you can't read the page of instructions when the book is closed. The book has got to be open to the right chapter and the right page to actually read the recipe for that particular gene and particular cell type. So actually, opening and closing down regions of the genome in terms of the way the string is folded up is actually intricately tied into whether the gene gets switched on or switched off.
Kat - To use that string analogy that you just mentioned, if you have a gene and then you have the enhancers that kind of tell the gene, "Be on. Be off" how far away from the actual gene themselves can they be?
Wendy - In terms of the actual DNA sequence themselves, we know that the controlling elements, what we call the enhancers can be 1 to 2 million base pairs away from the gene. It's an extraordinarily long length in terms of just a DNA sequence. But in terms of real spaces inside the cells, what does that mean? So that's the question that really interests me. And we think that an enhancer can act upon a gene as long as it's within about half a micron distance, so a half a millionth of a meter.
Kat - So, it's got to be kind of close enough to get a sniff of the gene and turn it on.
Wendy - Well, that's one model of how enhancers work. Does the enhancer of the piece of DNA that is the enhancer - I don't know - so it actually have to come over and touch the gene and switch some magic way, communicate with it directly? Or is it some other mechanism that doesn't really require the two bits of DNA to directly talk to each other. I prefer perhaps the model where they talk to each other indirectly by sharing proteins.
Kat - So, it's kind of just a lump of stuff and they're all part of this complex and that's how the information gets transferred.
Wendy - Yes. They both hang out in the same part of the cell nucleus where there's high concentrations of goodies to turn genes on.
Kat - Another aspect of your work that always interested me is that in all our cells, we have 23 pairs of chromosomes. That's 46 lengths of DNA string. How are they organised? Are they just all mixed up like 46 bits of string in a bundle, or do different chromosomes hang out in different places?
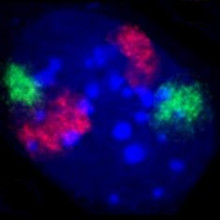
Wendy - Yes, our genome is not a plate of spaghetti. If we take the pasta analogy a bit further, it's probably a bit more like gnocchi. So each individual chromosome occupies its own space in the nucleus which we call a chromosome territory and we can see that because one of the great things about the human genome is, because it's so complicated and large, we can actually see it with and light microscope. And we can use fluorescent tools to actually see individual genes and individual chromosomes so we can paint chromosomes in different colours, so we can see that the chromosome territory for chromosome 18 which you might paint in red is in a very different and distinct place from the chromosome territory for chromosome 19 which you might paint in green. So, we can kind of do colour by numbers of the human genome.
Kat - And these pictures are very beautiful, but they're also - at least some of them are very static. What do we know now about how our DNA is moving around in the nucleus of our cells during normal life?
Wendy - So, if we step back and look at the chromosome territories that we just talked about, they're quite static. So, a chromosome is not able to move say, from one side of the nucleus to the other side of the cell nucleus within the lifetime of that cell. But an individual gene within that chromosome territory might be able to pop in and outside of the chromosome territory. So, we can actually see that happening in living cells. So, individual genes are pretty mobile, but again, constrained. We think of it as kind of dog on a leash. So, it can run around at will, but only so far. You're tied to the lamppost so you can't get to the other side of the road.
Kat - And is this tied into how they might be interacting with these enhancers, these switches as well?
Wendy - Absolutely, so if we watch a gene moving in a living cell, it's able to freely move. The dog on a leash can move over the space of about half a micron which is exactly the distance in the nucleus that an enhancer can work on a gene. So, for an enhancer to be able to contact a gene, it's got to be on a leash that's long enough to kind of wander over that nuclear space and find it.
Kat - Where do you think this kind of research is going to go next? What are the real questions that you want to try and get handle on?
Wendy - I'm interested in both the basic mechanism of how something a million base pairs away can communicate to a particular gene at the right time and place of development. So, what are protein factors involved? Does it really involve the chromosome folding over and touching? But also, interested in how this plays out in human genetic variations, so a lot of the differences between us are in the bits of DNA that don't code for protein sequence and the non-coding bit which we now know as full of this regulatory enhancer DNA. So, how do these very subtle changes in our DNA sequence between individuals affect the way we look, risk factors for various diseases, how high we are for example, or the colour of our eyes?
Kat - Or even things like how we behave?
Wendy - And of course, behaviour as well. I mean, the brain is a biological organ. It's controlled by gene expression so no reason to think that some behavioural traits aren't going to be due to single changes in these enhancer elements.
Kat - And what are you really excited about that seems to be just coming to focus now?
Wendy - I'm really excited by technology developments actually. They've really opened our horizons. We used to be constrained, looking at one gene at a time and doing very small scale experiments. Now, we can scan across whole genomes between multiple individuals and different types of cells, and really bring a much more comprehensive view of the genome.
Kat - That was Professor Wendy Bickmore, from the MRC Human Genetics Unit in Edinburgh.

10:00 - Mark Henderson - Genes and society
Mark Henderson - Genes and society
with Mark Henderson, Wellcome Trust
Now it's time to take a look forward to the upcoming Genetics Society Spring Meeting, which is focusing on genomics for health and society. Scientists from around the world will be looking at the potential benefits of the genetics revolution, but also exploring what this explosion of information and technology might mean for our society. To get a taste of what we might expect, I spoke to one of the speakers at the meeting, Mark Henderson. He's Head of Communications at the Wellcome Trust and author of the books The Geek Manifesto and 50 Genetics Ideas You Really Need to Know.
Mark - I think we're at a really interesting point. It's now 10 years since the publication of the final version of the human genome sequence. In the 10 years since then, we have in certain areas of medicine particularly, really progressed our understanding to the point that genetics and genomics is no longer just about research. It's actually about treating patients.
I think there are three particular areas where that's already having a real impact. One of them is rare disease that's children born with syndromes often causing mental retardation, that kind of thing, but really difficult to diagnose. What is increasingly happening is that it's becoming possible to diagnose at least some of those children using genome sequencing. The second area is infectious disease. That's actually sequencing bugs rather than people. Using that, it's now possible to for example track down the precise origin of now break of MRSA - the hospital superbug in a hospital. And the third area is cancer where cancer is a disease of the genes. It's a disease that happens when genetic mutations accumulate and as a result of that, it's been possible to develop some targeted medicines that actually attack the specific mutations that go wrong in cancer.
Where we are I think is with those areas and a couple more where we're now at a point where genomics is not a technology of the future. It's a medical technology of now. The big question is how we exploit that technology properly on the NHS and also, how we answer some of the deeper questions for society that genomics raises.
Kat - Because it does seem to me that the technology and the science has raced on far faster really than we can cope with it. Who do you think are the most important people to get across the stage of progress where we are and understanding these ideas and talking about them because this is happening now?
Mark - So, I think there are at least three key audiences that really need to understand this better. The first one is policy makers - people within government who will be taking decisions over the coming 2, 3, 4 years about how we fit up the NHS in particular to take proper advantage of these technologies. At the moment, I think understanding is improving and there's certainly a great deal of enthusiasm. But it's important that that enthusiasm is matched by realism as to what genomic medicine can but also can't deliver and that we don't end up with expectations raised too high, only for them to be dashed. That's the first audience.
The second audience is actually the medical profession where beyond certain specialist kinds of doctors who have always been very well-versed in genetics - the clinical geneticists, the oncologists - I think something that will happen increasingly is that genomics becomes embedded in the practice of medicine throughout the clinical care pathway. And even those going those going through medical school right at the moment are not really taught in a systematic way about how complex genomics might influence things such as drug prescribing, some of the options that might be available for cancer patients, rare disease, infectious disease.
Kat - And I guess the final group is the public.
Mark - That's right and I think it's very important here that we try to achieve the communication of genomics in a more realistic way. The way in which genomics and genetics are often covered seems to suggest that they're deterministic sciences - we read a lot about genes for being fat, genes for being tall, genes for getting cancer. It's actually very rare that genes work in a deterministic way like that. There are one or two conditions - Huntington's disease would be the prime famous example - where there is a one to one relationship between inheriting a certain mutation and getting a certain disease. But actually, that's not how genetic risk works more normally. More normally, there'll be 10s, 100s, even thousands of genes working in concert with the person's environment to raise or lower their risk of often a large number of different disease.
And I think that as soon as you start to think of genetics like that, it actually starts to become a little bit less scary that your genetic code is a very important factor in your health and well-being, but it's only one factor. It certainly doesn't mean that your diet is unimportant, how much exercise you take is unimportant. Other lifestyle and environmental factors work together with the genome to form outcomes, not in isolation from them. And your genome very rarely determines either who you are or what diseases you have.
Kat - That was Mark Henderson. The Genetics Society spring meeting will be held at the Royal Society in London on Friday 19th April. For more information, go to genetics.org.uk
16:17 - Dr Nick Gilbert - Twisting DNA
Dr Nick Gilbert - Twisting DNA
with Dr Nick Gilbert, NRC Human Genetics Unit, Edinburgh
Now it's time to take a closer look at the twisted world of DNA. Inside our cells, our DNA is wrapped around ball-like packing proteins called histones, forming something known as chromatin. But what does chromatin look like? And why doesn't our DNA get all tangled up? Dr Nick Gilbert, also at the MRC Human Genetics Unit in Edinburgh, is trying to find out.
Nick - If you isolate chromatin from cells and unfold it and look at it under a microscope - a very high-powered microscope like an electron microscope - what you would find is that it looks like sort of beads on a string-like structure. But if you then fold up this chromatin into what you'd normally expect in a cell, it has a sort of much more folded appearance and'sthickened out and we actually call it, something called a 30 nanometre fibre. So, not a very imaginative name, but that's the sort of the dimension you see under the microscope.
Kat - It strikes me as an incredible feat of packing. You've got these long, long strings, you wrap them around beads, you coil them up, and coil them up, and coil them up, and shove them all into the nucleus. How does it not get twisted up? How does it not end up as just a massive mess?
Nick - Now, that's a very interesting question and it's something that's very important for the cell to avoid and the cell has some fairly complicated enzymes which were actually responsible for removing a twist that's actually introduced into the DNA. And a lot of these enzymes are topoisomerases ,so these are enzymes that can chop the DNA like a molecular pair of scissors, remove some of the twist and then stick it back together again. There's also another class of enzymes called helicases and these helicases are like little molecular motors that can twist the DNA. And interestingly, there are a couple of human genetic diseases where there are mutations in some of these helicase proteins, and these are very, very rare genetic diseases. For example, Bloom syndrome and Werner syndrome, and in these patients, the little helicases don't work properly and then there are problems with packaging the genome.
Kat - So, how do these helicases and these topoisomerases work out where they need to go and what they need to be doing? Do we know how they sense whether DNA is too twisted or not twisted enough?
Nick - So, that's something we really don't have much of an idea about. So, we did a study recently where we did some experiments on topoisomerases and one of the things we concluded from these experiments is that these enzymes must have the ability to sense twist. But we've got no idea as to what that mechanism is, whether they're actually recognising a change in structure in the DNA or if there are other sensing proteins that can signal to the topoisomerases that they need to be active in different parts of the genome. What we did find though is that there are - in mammalian cells, there are two main classes of topoisomerases and from our experiments, they appear to have different distributions in the genome, implying that they might have different functions in different regions. But that's really a very broad scale and we need to do more experiments to actually understand how these enzymes detect changes in twist.
Kat - It's an amazing image to think about that you've got these little beads of protein being shuffled about on our DNA. You've got little machines pushing them around. You've got things untwisting DNA, twisting it back up. It's very dynamic idea of what's going on in the nucleus of a cell.
Nick - Well, I think historically, people have thought of chromatin as a very static structure, but this is probably a reflection of the techniques people have used. Historically, people have used microscopy to look a lot of these processes, but microscopy is used on samples that have been fixed, so they're sort of static samples.
Kat - They're dead stuff basically.
Nick - Essentially, but what we have found now is that using a lot of new techniques that actually a lot of the proteins in the cell as you say are very, very dynamic. And we need to think of the organisation of DNA and its packaging in the cell as a very dynamic process that's constantly responding to for example, gene transcription or DNA replication or just other external stimuli from outside that the cell needs to respond to so it can express certain genes. So, it's very important for us to develop new techniques to actually look at these dynamic processes as they're actually occurring in living cells.
Kat - So, tell me a bit about the work that you're doing, how are you trying to unravel this complicated problem?
Nick - So, my lab is really interested in looking at the structure of DNA and how DNA is then folded into the sort of fundamental chromatin fibres. So, we've spent quite a lot of time looking at the structure of these chromatin fibres in cells using a combination of sort of biophysical techniques and very high resolution imaging. And what we find is that in different parts of the genome, these chromatin fibres have slightly different structures. What we've been doing recently which is very interesting is to actually look at the structure of DNA in cells and there's really so much still we don't know about it.
For example, we know the structure of DNA in a test tube on the lab bench, but we don't really have an idea of the structure of DNA actually inside the cells. So, what my lab has been doing over the last few years is try and develop new techniques for analysing the structure of the DNA whilst it's still in the cell. And we've recently developed a new sort of molecular probe that can distinguish between DNA that has a very twisted structure and DNA that has a less twisted structure. And we are able to show that the genome is actually organised into large domains of twist and I think this is something really very interesting that we're going to carry on studying.
Kat - You mentioned earlier that there are some human diseases where there are problems with the helicases that twist up DNA and the topoisomerases that untangle them. Do you think that DNA twisting could be important in some other human diseases?
Nick - Yes, I think absolutely. For example, in diseases like cancer, you find that topoisomerase enzymes are often upregulated and what we imagine is that this might have an effect on the DNA structure so that maybe the chromatin is not packaged properly and it enables other genes to be misregulated. And cancer really is a disease of gene misregulation. And if you start messing around with the proteins that are responsible for packaging the genome properly then chances are, then genes will be expressed at the wrong time and in the wrong cell type. And that really could be a very important initial cause of cancer and might also well be important in the evolution of diseases like cancer.
Kat - That was Dr. Nick Gilbert from the MRC Human Genetics Unit in Edinburgh.

How is dinosaur DNA purified?
Kat - Now it's time to look at your burning genetics questions, with the help of Naked Scientist Martha Henriques.
Martha - This month, listener Steve Cherry from Lethbridge, Alberta in Canada wanted to know how researchers extract dinosaur DNA from fossils and separate it from contaminant DNA from other organisms. Firstly, the bad news is, that no one has yet successfully isolated DNA from dinosaur fossils. Dr. Greger Larson from Durham University told me how DNA is isolated from fresh biological specimens and why doing the same for ancient specimens like dinosaur remnants is so difficult.
Greger - The way that we usually do it is to simply remove everything that isn't DNA and filter it off and that leaves us with the small tube that only has the DNA left in it. Of course, DNA exists in every single living organism, so we can't discriminate between all the different possible sources of DNA. All we can do is isolate DNA from everything that's non-DNA. And once we've got that in a tube then we can start to do all kinds of things including sequencing it or going after specific markers, or amplifying specific places, but the act of isolating it is a relatively straightforward process and we can do it from any number of things including skin cells or feathers or bone, or teeth, or anything that's biological and that is alive or was recently alive, we can isolate DNA from.
Martha - But when it comes to ancient specimens like dinosaur fossils, the chance of success with this method becomes vanishing small.
Greger - As soon as something dies, everything about that thing starts to degrade. So, the analogy would be, the bowl of fruit that you probably have in your kitchen and if you were just to leave that bowl of fruit there for a day or two, it'll be fine, but if you leave it there for a month, it will probably be no longer something that's even remotely appetizing. And if you leave it there for a year, it would be hard to differentiate the different kinds of fruit you had in that bowl a year before and if you leave it there for 20 years, there probably won't be anything even remotely resembling fruit in there and DNA is no different.
The longer that an organism is dead, the more of the DNA degrades and it degrades at a rate which is dependent upon lots of different factors including how long it's been, what temperature it's being stored at, or what sort of other organisms are eating it, or any number of other things. But even in the best case scenario where you have something frozen say, in the tundra where it's very cold, the half-life of DNA is somewhere in the order of maybe 100 or 200,000 years which means, half of the amount of DNA that you start with will be gone after about 100 and 200,000 years.
So, the reason it's difficult to get DNA out of dinosaur bones is simply because there is no DNA left. The same reason that if you were to leave the bowl of fruit on your kitchen table for 65 million years, the chances of you being able to come back and get any fruit out of that would be virtually nil. So, any reports that there's been of dinosaur DNA, they're either not reporting dinosaur DNA or they're reporting other bits of DNA that they've isolated that don't have anything whatsoever to do with that dinosaur bone.
Martha - This contaminant DNA is a problem for all work on ancient biological specimens. Dr. Mim Bower of Cambridge University worked from isolating DNA from prehistoric horse remains preserved in icy environments. These well-preserved specimens are much younger than dinosaur fossils and still have some of their original DNA present. Dr. Bower describes how she distinguishes any contaminant DNA from the horse DNA that she's interested in.
Mim - So, for every sample of bone that we try to extract DNA out of, we also examine a number of controls so that each time I do a DNA extraction, I work with a set number of 6 control reactions which have exactly the same components in except for the bone particle and that helps me control whether there is a DNA coming from me, or from the environment, or from the reagents that I'm using because if I see a positive DNA peak in my control samples which should have nothing in them then I know that I have made an error somewhere and I go back and I start again, and I'd clean everything down. So, we have a very strict contamination protocol nowadays and I have to say that most DNA researchers who are experienced can work very cleanly. And when we don't work cleanly, we can immediately see what has gone wrong and put it right.
Martha - That was Greger Larson from Durham University and Mim Bower from Cambridge University.

28:13 - Gene of the month-Scott of the Antarctic
Gene of the month-Scott of the Antarctic
with Kat Arney
Our gene of the month is the adventurous Scott of the Antarctic, which was first discovered in fruit flies back in 1996. It was named because the protein encoded by the gene was originally thought to be involved in forming a cellular scaffold called the spindle, which helps separate chromosomes as they divide, by pulling them to each pole of the cell by means of structures called centrosomes. In Scott of the Antarctic mutant fly embryos, the centrosome of one cellular pole becomes detached and doesn't work properly, in a not-very-good analogy to the doomed South Pole expedition by Scott and his team in 1912.
It now turns out that Robert Scott's namesake gene is actually an overactive version of another gene called Greatwall, which is important for helping cells to divide as well as helping to coil up DNA in preparation for division. Not such an evocative name though.










Comments
Add a comment