From whole populations to individual cancers, we're taking a look at genes and evolution. We're also talking about dogs and their diseases, shining a light on Van Gogh's sunflowers, and wondering whether the USB-sized DNA sequence is hope or hype. Plus we've got our gene of the month - whether it's Sonic, Desert or Indian, we'll be carefully getting to grips with the prickly persona of the hedgehog gene.
In this episode

01:14 - Human evolution - Professor Mark Thomas
Human evolution - Professor Mark Thomas
with Professor Mark Thomas, UCL
Mark - Up until recently, when we talked about evolution, we meant genetic evolution. Because what we were talking about was changes that are inherited. If they're not inherited, if they're not passed on from one generation to another, then evolution can't act on them. Processes can choose or natural selection can choose something as being a good idea or not a good idea. But if it's not passed on from generation to generation then it's not an evolutionary process. And of course, when we think about Biology and passing on from generation to generation, we're firmly in the field of genetics there. And in fact, you know, many people would argue that in fact evolution is just genetics plus time. So you add genetics, the rule of genetics to time you'll get evolution. It's just inevitable.
Kat - And so what do we mean exactly by evolution? What are some of the pressures that are acting to change genes over time?
Mark - When we think about evolutionary pressures, of course the first thing that we think about is natural selection. And of course, Darwin first proposed this is the means by which species change over time. And natural selection is a wonderful idea. And of course it is real, it works. And so - but we have to remember that there are other forces that change the frequency of genetic variance over time, with different gene types over time.
And one of the really important things is just randomness, just the noisiness of inheritance. So there are many, many different genetic variance in a population, for example, if you imagine a population of people alive today. And some of those are getting passed on, some are not. So of them get passed on because they're disadvantageous, because they've caused their carriers to die or fail to reproduce. But some just won't get passed on by chance.
Kat - So you can imagine for something like humans to be able to run away from a predatory animal, it would be advantageous to them to select for genes that make you run faster. What are some of the other characteristics that humans have been selected for?
Mark - Ten years ago, you would have got somebody waving their arms around saying "well I think it could be a bit of this or a bit of that". But now we know exactly what's been selected. They are genes involved in environmental adaptation. So one obvious, clear cut example of environmental adaptation you see in humans today is skin color. And that's obviously a response to the amount of sunlight, particularly the amount of UV.
And then things involved in diet. When you think about it, that makes a lot of sense because you don't eat or you don't eat and you don't digest or you don't deal with the food in the right way, you're not going to survive. You need food for it's one of those things like sex that we need in order for the species to keep on going. And it turns out that in fact if we look that, say at Europeans and we dig into the genome of Europeans and look for these signatures of selection that we can see nowadays, we can actually see in the genes the signature of selection. We can see -- we can get an idea of how strong they are.
One of the very, very strongest signatures of selection in Europeans is actually an adaptation to be able to digest the sugar in milk. And that only occurred in the last seven-and-a-half or 8,000 years. That should be obvious why that's the case because prior to, when we had farming, we didn't have animals. And if we didn't have animals, we wouldn't have a supply of fresh milk to drink. Well, we could have got it in other ways but that would have been a bit rude. So you know, we really need domesticated animals to get the milk. And sure enough, that's the time when we see this adaptation to be able to digest the sugar in milk.
There are also adaptations in our immune systems. That's exactly what you expect because of course pathogens are evolving themselves all the time. And we are in an arms race with pathogens. We're in this constant battle. You know, we don't - you don't win that war. You just keep on fighting it and you keep on doing as well as you can. And the pathogens will try to do well as they can as well. So the genes that are involved in our immunity to pathogens, you can see really clear cut signatures of natural selection have an action on those.
Perhaps more surprisingly, another set of genes that seemed to be under strong natural selection but it's maybe a little bit less obvious, were genes involved in sperm motility and in sperm production. And the theory behind that, why that's the case is that, in fact sperm are competing with each other. And if you think about it, I mean if you have a multiple mating event, then if one set of sperm could kill out the other set of sperm, they're more likely to produce offspring. And so therefore there's going to be some - that maybe reflected some kind of competition between sperm.
Now the ones of course that everybody would really want to find out and be really excited about is, what about the ones that make us brainy? Because of course, you know, when we require brainy species as species go but you know, you go back about five million years, I mean we the same as a chimp, with a similar size brain. So you know, the really kind of cool questions are, you know, what has changed in terms of our genes that makes us brainier? Unfortunately, we don't really know a lot about that one.
There are few genes that people have suggested that are involved in brain size. There are genes that people have suggested are involved in linguistic ability, language ability. But what it beginning to look like is actually what made us start doing really smart things, was not natural selection evolving our brains to be cleverer. I mean ultimately, they must have evolved to be cleverer at some point. But doing the clever stuff seems to be another form of evolution. And that form of evolution is cultural evolution.
Kat - So you've talked about these five pressures. We've talked about food, we've talked about sex, we've talked about the environment, our brains, and the immune system. And you say you can see these genetic signatures of evolution. How do you do that now? How can we actually look in our genes now and our genes in the past and see how this has evolved?
Mark - Imagine we're back 10,000 years ago and some new mutation arises. And that mutation is - when it arises, only arises in one person. So it's very, very rare initially. Imagine that new mutation makes absolutely no difference to anybody. It doesn't help them survive but it doesn't lead them to dying either. By chance, that mutation may be passed on to one child or to no children or to two children or three children. And then by chance, the next generation might be passed on again and again. So you can imagine that the frequency of that variant might change a little bit in each generation. It might be go up, it might go down.
Now imagine another scenario. And so new mutation occurs 10,000 years ago and it happened in one individual but it gives a big advantage. So it's much more likely to be passed on to two or three or four children and to increase in frequency in the subsequent generations. So if it's increasing in frequency, then it's going to go up and up and up until it's very, very great common, right? So the difference between a gene that's been under natural selection and a gene variant that's just bouncing around, just neutral, is that the ones in the natural selection, they go from low frequency to high frequency quickly. And it's the quickly that's the essential thing, okay?
So we can tell whether a gene has got from low frequency to high frequency, by for example, looking in a population from ancient DNA and then looking in a population today and seeing what the difference in frequency is. And if that change is simply too quick to explain by random processes, then we have to invoke natural selection instead.

10:18 - Epstein-Barr virus in dogs
Epstein-Barr virus in dogs
with Nell Barrie
Nell - So this is about dogs and Epstein-Barr virus, which is type of virus that can cause some types of cancer. And it can also do that in dogs as well. And this could maybe mean that we've got a new model for studying this type of relationship in humans.
Kat - Because there's quite a lot of human diseases that are actually in dogs. I remember long time ago, there was a paper about I think narcolepsy in Dobermans and there were all these dogs just falling over asleep. At the moment, there's no other animals that get Epstein-Barr virus. Why do you think this is going to be useful?
Nell - Well, one interesting thing about this virus and a lot of other types of viruses is that we know that many, many people are affected with them, infected with them rather. But they don't always lead to particular types of cancer and everyone who's got the virus inside them. So we need to figure out why that is, why some people are susceptible to this and others aren't. Perhaps we can figure this out in dogs. It will give us some clues for what is going on in humans. And that might lead to new ideas how to treat the disease or maybe prevent it.
Kat - Because Epstein-Barr, this is really, really common particularly in places like Africa where it really causes a lot of cancers.
Nell - Exactly. So we don't really know what's going there, why are some people susceptible to this and others aren't. We just don't know what's happening inside those cells in the moment. And if we have an experimental model in dogs for example, that could really give us powerful idea to figure out what's happening.

11:36 - Epilepsy in dogs
Epilepsy in dogs
with Nell Barrie
Kat - Another story that I noticed involving dogs and their diseases is about epilepsy. And actually a quite number of different breeds of dogs are affected by epilepsy. There's that Belgian Shepherd dogs, about 1 and 5 of them, get some type of epilepsy. And there's some research in Finland of tracking down the gene that might be involved in this. What have they found here?
Nell - So they're looking an area of particular chromosome that maybe the key to what's going on in these dogs. And it's again, it's another funny one where it seems like quite a lot of dogs have this particular area and have mutations in this area of the chromosome. But they're not all getting epilepsy. So again, it's kind of, you know, what's happening there? Why is this affecting some dogs and not others? Can we figure this out? And could that maybe give us clues for disease in humans as well?
Kat - Because the thing to point out about dogs is that they're quite highly bred, so they got less genetic variation than humans. And I noticed as well the research group that has done this, they've been looking all sorts of other dogs as well. And they have a canine DNA database in Finland. They've got DNA from 40,000 samples from more than 250 different breeds. So I think that's quite a useful resource there.

12:44 - Sequencing the microbiome
Sequencing the microbiome
with Nell Barrie
Kat:: Another story that we also noticed this one is about the microbiome. Now they've sequenced the human genome, they're trying to move on to sequencing all the bugs in our gut. Is this possible now, do you reckon?
Nell:: It certainly looks like that. I mean we've got so much more powerful genetic sequencing technology compared to, you know, 10 or even 2 or 3 three years ago. And it's just really exciting the amount of data you can get out of this. And I supposed, the funny thing about this research is we don't really know what we're going to do with the data. We've got this amazing potential for finding out all kinds of cool things but we don't know what those cool things are going to be. So a lot of the coverage is a bit like as this hope, is it hype? What are we going to do with that? How is it going to help people? And we don't know yet but I don't think that's the reason why we shouldn't be doing it. I think it looks really exciting and it could have a lot of interesting applications for a lot of kinds of diseases.
Kat:: Yes. Some of the coverage I was seeing was saying, "Well we sequenced the human genome and you know, it hasn't solved everything." But I don't think that's a reason not to do this because actually, when you think about it, manipulating the bugs in your gut is probably easy than trying to develop drugs to target gene faults. Because have you seen the stuff about poo transplants? This is great, we've seen this.
Nell:: It's really cool. Yes. That just sounds amazing. So it's kind of taking all the bugs from someone else's gut and transplanting them into somebody who's maybe affected by autoimmune problems or they might have something wrong with their digestion. And in some people it seems to have really amazing effect. And again, you know, we don't really know what's going on there. But if we can figure out how these little bugs are interacting, how they affect people, then we could maybe work out a nicer way to do that that doesn't involve transplanting poo.

14:26 - "USB" gene sequencer
"USB" gene sequencer
with Nell Barrie
Kat - And another story that hit the headlines, the USB-sized gene sequencer. And I think people just went nuts for this. Did you see this one?
Nell - I think it's one of those ones when you look at the picture, and just go, "Oh my god, it's amazing. It plugs into your computer." And then...
Kat - The future is here.
Nell - Yeah. And then you think actually what would I personally do with it, I can't do anything with that.
Kat - I'd sequence anything!
Nell - But one point they made as part of the story was, you know, yeah, you get all this data out of it but then what you do with it? You've got your USB plug in but you might also need a supercomputer to go with it to analyze all the stuff that you've got. That might be bit more expensive than the sequencer itself.
Kat - Yeah. What's interesting is that it's called I think GridION technology, works at a different way to normal sequencers. And that there's basically all nanopores that are full of tiny holes. And the DNA gets pulled through it like pulling a necklace through a hole. And each nucleotide, each letter of the DNA gives out a slightly different electrical signal and they monitor those. So those were going, pop-pop-pop-pop, ACTG, whatever. That's what they say. Though there is quite a lot of controversy about how accurate it is.

15:28 - Evolution of cows
Evolution of cows
with Nell Barrie
Kat - And speaking of sequencing and things like that, earlier in the program we heard from Mark Thomas, who's looking at human evolution and cultural evolution. And is there a nice story I noticed about cows. So what's going on here?
Nell - I really like this. So they were looking at DNA from ancient cow ancestral fossils from Iran, in fact. That's quite interesting because they've taken these bones from a very hot desert environment and still somehow managed to get DNA out they can use to figure out what the DNA sequences were back in those ancestors of cattle. And this kind of made people go, you know, "Ooh Jurassic Park, you know, we can recreate prehistoric cows". I got very excited. They're not quite there obviously. But it's interesting to see how you got this very small population of cattle that were initially domesticated and that has given rise to all the kinds of cows we have today. So it's quite cool.
Kat - Also it's cool in as well the point that they're making the, in the story also that these were Aurochs, big ox things. Now even when you start domesticating things like that, it's going to be tricky. So I think it's probably a number of years before they really got them domesticated. So, the early life of a cattle herder...
Nell - Yeah. It might have been more scary.

17:24 - The Gorilla Genome
The Gorilla Genome
Never mind Gorillas In The Mist, how about gorillas in the lab? Scientists at the Wellcome Trust Sanger Institute have sequenced the whole genome of the gorilla - the last of the great apes to have its genome read. This means we can now compare the genomes of humans, chimps, gorillas and orangutans, shedding new light on primate evolution.
This means we can now compare the genomes of humans, chimps, gorillas and orangutans, shedding new light on primate evolution.
The results show that gorillas diverged from chimps and humans around 10 million years ago. And although chimps are still our closest primate relative, there are some intriguing similarities between human and gorilla genes - for example, in the evolution of genes for hearing, which cast doubt on theories that the evolution of human hearing is linked to language development.

18:07 - Zebrafinch singing
Zebrafinch singing
Scientists at UCLA have identified around 2,000 genes in zebrafinches linked to singing - more than 1,500 than were previously known about. All 2,000 show changes in activity levels when birds sing, and a number of them are also found in humans, making the researchers think they may also play a role in human speech and speech disorders such as autism.

18:30 - Light-sensitive stinging cells
Light-sensitive stinging cells
Researchers studying hydra - tiny sea-creatures related to corals and jellyfish - have made the surprising discovery that their stinging cells, called cnidocytes, are sensitive to light.
Although it's known that these cells sting in response to touch and taste, this is the first time they've been linked to light.
The scientists, led by Todd Oakley, found that the stingy cells contains opsins - light-sensitive proteins that are used widely in light-sensing organs and eyes across the animal kingdom.
Oakley thinks that his findings may help to shed light on the evolution of eyes - well, we'll see...

19:06 - Salt-tolerant wheat
Salt-tolerant wheat
Writing in the journal Nature Biotechnology, scientists in Australia have bred a new strain of salt-tolerant wheat, by crossing in a gene from a more ancient strain of the plant with a higher tolerance for salt. Around 20% of agricultural land around the world is affected by high saltiness, or salinity - which can seriously affect plant growth - and it's only getting worse as the climate changes. At the moment, the researchers have tested the gene in durum wheat, used to make pasta and couscous, and they hope to cross it into bread wheat next. Importantly, they didn't use GM techniques to get the gene into the wheat, so they're hoping this will speed up the process of testing and commercialisation.
19:48 - Fungi for biofuel
Fungi for biofuel
And finally, researchers in the US have sequenced the genome of particular species of fungus that can turn cellulose into biofuel. The fungus, called Ascocoryne sarcoides was found to have more than 80 clusters of genes that can convert cellulose into useful fuel - many of which have only previously been found in plants. The scientists hope their work could pave the way for further explorations of the potential of fungi for making biofuels.
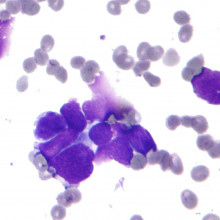
20:33 - Tumour evolution - Professor Charles Swanton
Tumour evolution - Professor Charles Swanton
with Professor Charles Swanton - Cancer Research UK London Research Institute
Charles - We have to determine how representative a single tumour biopsy was of the entire tumour genomic landscape. And by that I mean if you identified mutation in one part of the tumour, how likely are you to identify that mutation in another part of the tumour? Because clearly, if we're going to identify new ways of predicting drug response, we have to go for genetic events that are common throughout all parts of the tumour so that a random biopsy could pick those genetic events up.
And what we found actually is that, in the study when we took multiple biopsies from several primary renal cancers, and in some cases their metastatic sites, that the number of genomic aberrations or the pattern of genomic aberrations very often wasn't shared across all biopsies. And in fact, when we counted the number of mutations, about two-thirds of the entire mutational load in the first two tumours as we sequenced were not shared across every biopsy taken from those two tumours.
Kat - So this is quite a big difference from the original tumour. You find differences in the gene faults within the tumour and also, in secondary cancers where it spread. Presumably this almost blows personalized medicine out of the water, what do you think is going on here?
Charles - No, I definitely don't think this blows personalized medicine out of the water. I think what this does is actually begins to inform us that if we consider tumours as trees where we have the common mutations represent the trunks of the tree, these mutations are present at every site of the disease. And the mutations that differ from one region to the next represent the branches.
Now, in terms of personalized medicine, one can perhaps think of HER2 or the BRAF mutation as the mutation or the amplification event that occurs at every sites of the disease and is therefore a good drug target.
Kat - So that's the trunk?
Charles - The trunk, exactly. That controls the disease effectively because that mutation or genetic aberration is present at every site of the disease.
The model taken through is sort of logical conclusion also helps to perhaps think about how drug resistance is acquired, that low frequency events within the tumour that may confer drug resistance upon the tumour. Our equivalent to the mutations perhaps that occur in the branches of this tumour to take the tree analogy further. And during treatment, those resistance mutations may be selected out enabling the tumour to evade drug treatment.
Kat - So when you cut off some branches it just keeps growing.
Charles - Exactly. That's right. It's like cutting off some branches.
Kat - What does this mean for the field of genomics research in cancer?
Charles - First and foremost I think we need to sequence fewer tumours in greater depth and at multiple regions, comparing primary with multiple metastatic sites to really truly identify those driver trunkal mutations that are present and responsible for the disease biology at every site of the disease. Those can be very efficient drug targets one would imagine.
It also tells us that actually understanding what's driving the diversity, in terms of the tree model, what's driving the changes between the trunk and the branches, actually maybe very important for trying to restrict underlying tumour diversity, to stop, if you like, the target from moving. Because it's the diversity within the tumour that is likely responsible for tumour adaptation, resistance to treatment, and potentially the acquisition of mutations that allow tumours to grow at sites distant from the original tumour.
I hope in the next 10 to 15 years, considering tumours as trees, trunks and branches, it might help to improve the development and discovery of new drugs and enable us to understand how resistance to treatment is acquired by studying how these branches are cut off.
Kat - You mentioned that you're sequencing the whole genomes of several samples of a single cancer, how have advances in technology allowed us to do this kind of research?
Charles - I mean we wouldn't be able to do this research two or three years ago. I mean the advances in sequencing machines, in computer technology, have been immense over the last two to three years and continue to be dramatic and will continue in the future to change and enable us to sequence tumours and process the data at an unprecendented rate
We're already seeing in some centres, in the US and elsewhere offer patients access to whole genome sequencing facilities as part of their treatment. So already we're seeing human genomes sequencing impact upon patient care.
Kat - And finally for you, where next for this research and for your team?
Charles - What we're very excited about is the implications of this research to our understanding of how resistance is acquired during treatment. Because it's only by understanding resistance to treatment that we will really make progress in prolonging patient benefit from the drug treatments we already have in the clinical setting.
So it's really understanding what's going on in branches of the tumours, it's understanding how the branches changed through therapy, and most importantly I think, understanding what initiates the change from the trunk to the branches, what is basically creating the underlying tumour diversity.
And when we've addressed to all of those problems, I think we'll have a much greater understanding of kidney cancer as a human disease and our, hopefully ability to target it much more effectively.
Why does cancer have multiple causes?
As we heard from Professor Swanton earlier, at its heart cancer is a disease driven by faulty genes - either faults that lead to genes being switched on that drive cells to grow and spread, or that lead to protective genes being switched off. In fact, one of the most potent causes of gene damage is reactive oxygen molecules produced by our own cells as they make energy. We can also inherit gene faults that make our cells more likely to pick up damage or fail to repair it properly. Chemicals in tobacco, our diet and environment can also cause damage, as can UV light. And certain viruses - such as the human papillomavirus that causes cervical cancer - can also hijack the genetic programme of cells and make them grow out of control. Different cancer types often have different risk factors - for example, UV light from the sun or sunbeds damages DNA in skin cells and can lead to skin cancer, but it can't penetrate inside the body to cause cancer internally. So while the fundamental root of all cancers is the same - cells growing out of control - individual cancers may be caused by different factors, and usually by a combination of many different things that it may be impossible to separate out on an individual basis.
Do genes delivered by gene therapy get inherited?
Most types of gene therapy are what's called somatic gene therapy - they're designed to only deliver genes to certain non-reproductive cells of the body, such as delivering a healthy version of the damaged cystic fibrosis gene only to the lung cells of someone with the disease. This is the main approach being used by scientists working in gene therapy. It's very unlikely that these genes could transfer into the sperm-producing cells in men, and even less likely they could get into the egg cells in women. Germline gene therapy, which changes the genes in sperm or eggs so they would be passed on to children, is actually banned in most parts of the world at the moment, for both technical and ethical reasons.

29:52 - Gene of the month - Sonic Hedgehog
Gene of the month - Sonic Hedgehog
with Kat Arney
If you ask most people what they think of when they hear the words "Sonic Hedgehog", they'll probably describe a spiky blue video game character. But ask a biologist, and they might at least pause for a moment, because as well as being the main protagonist of the 90s Sega games, Sonic Hedgehog is also an important gene found in mammals, along with the similar genes desert hedgehog and Indian hedgehog - both of which were discovered before their showbiz counterpart and are named after real species of hedgehogs rather than cartoons. But why hedgehogs at all?
Like so many genes with wacky names that we'll cover in this series, the original hedgehog gene was first found in tiny fruit flies called Drosophila Melanogaster. Healthy fly embryos are torpedo-shaped, with neat rows of spiky scales called denticles. But fly embryos with a faulty version of the hedgehog gene are short, dumpy and rounded, and completely covered in denticles - much like the spikes that cover a real hedgehog.
In fruit flies, hedgehog is responsible for defining the front from the back of the individual segments that make up a fly embryo, and it's also involved in helping flies to grow wings and other organs. And although mammals like humans don't have wings or make embryos in the same way, the mammalian versions of hedgehog - Sonic, Desert and Indian - are all involved in helping to shape an animal as it grows in the womb. For example, Sonic hedgehog sets up a front-to-back pattern in something called the neural tube - the precursor of the brain and spinal cord - and also helps to form fingers and toes. So despite the comical name - which is probably the closest thing that passed for cool in the scientific world in the 90s - Sonic hedgehog and the rest of his genetic family are vitally important.










Comments
Add a comment