Synthetic biology - engineering life - is set to revolutionise the world, but how? We'll be hearing about some of the most exciting applications for synthetic biology, and how it's being commercialised. Plus, our gene of the month has got itself all in a twist.
In this episode
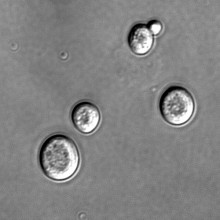
01:02 - Paul Freemont - Biosensors hacked yeast
Paul Freemont - Biosensors hacked yeast
with Paul Freemont Imperial College London
Kat - Back in October we visited the world of synthetic biology - using genetic engineering techniques to cut and paste DNA together to create exciting new biological components that can do all kinds of things. Now we're returning to that theme with a look at some of the potential applications being developed using synthetic biology approaches. Paul Freemont is co-director and co-founder of the Imperial College synthetic biology hub in London, and he told me about some of the exciting approaches he and his team are exploring.
Paul - We have all this huge antimicrobial problem and we have a lot of need to have very good, possibly point of care diagnostics that could detect infections, bacterial infections or whatever quickly and cheaply and reliably. So, one of the great projects I think that my lab is working on is to develop a whole series of in vitro biosensors. What I mean by that is that you could use a living cell as a biosensor. But because of the application space, people feel slightly uncomfortable having an E. coli as a biosensor.
Kat - Yeah, bacteria sensing bacteria seems weird.
Paul - And which one is the good one and which one is a bad one, and all that stuff. So, we early on decided, well okay, some people might feel uncomfortable with that. So, let's think about, "Can we use the cell extract which would have all the machinery to be able to run our genetic programmes that we've been designing?" So essentially, what we use is the cell-free extracts. So they're non-living. They don't have any DNA in them. And then we put into that extract our designed DNA which would then make some proteins, which would then detect various signals that are analytes or small molecules or whatever you want to call them - chemical molecules which would indicate the presence of a pathogenic bacteria. And when that molecule bound that little chemical entity, it would then activate the production of some colour or some fluorescent protein or something else. But it's all genetically encoded if you see what I mean.
Kat - So, it's kind of almost the same idea with like pregnancy test stick or something like that where you pee on it, it changes colour but this is a biological system that's changing colour.
Paul - Perfect example, perfect analogy. I think what's exciting about this type of approach is that there's been some recent work from colleagues in the United States to indicate that these cell-free extracts can actually be freeze dried onto filter paper potentially. We've also been working on other kinds of materials that we can sort of freeze dry these extracts. So, it's essentially quite a complicated extract and it's got a complicated piece of DNA in it. But actually, this could end up being an incredibly simple device.
Kat - So, you could just pee on it?
Paul - Yes, theoretically. That's kind of exciting because it would be very cheap, easy to use, and it would be quite safe. It wouldn't be quite safe - it would be very safe.
Kat - So, that's using this kind of technology to detect bacteria, pathogens in the environment. Are there other things that you could detect?
Paul - There are and I think people are very interested in detecting sort of potential diagnostic biomarkers that could indicate if someone is unwell or if they've got a continual condition like diabetes or whatever or if they might have had a heart attack, there'll be some people who are very interested to have early detection systems or whatever. You can imagine the kind of scenarios. So, there are a whole bunch of other people and ourselves working on biosensors that would detect protein biomarkers. So maybe, we'll end up with sensing protein markers and protein markers in the urine or in the blood which could indicate either cancer or some particular early stage physiological dysfunction. So, those are very exciting. And then the other ones I think which are also very exciting is developing sensors that could be used in more global healthcare scenarios. So, one other sensor we're working on in my lab is to detect the schistosomiasis parasite. This is a very debilitating disease. It's a waterborne disease. It affects 200 million people in the world. It's a tiny little parasite that actually uses humans as part of its lifecycle, so it's a really interesting bit of biology. But it would be really good if we could have a very local, very cheap, simple diagnostic sensor to test water samples. Not necessarily to prevent people from going into water but just also to understand the disease, to understand just the epidemiology of the disease, and just also to have a tool that local water management people could use in that kind of area. So, we're working on that as well.
Kat - How about another thing that's really exciting? Tell me about something else that's really cool right now.
Paul - The other really amazing project which we're involved in is called the synthetic yeast project and this is a really, really exciting project because what the project is about is to essentially synthesise chemically all of the chromosomes of Saccharomyces cerevisiae. Now, Saccharomyces cerevisiae is better known as baker's yeast and you use it to make bread and make beer or whatever you ever use it for. And it's also very well-known. It's a very well-studied organism and we've been studying it for hundreds of years, thousands of years, we've been using them. So the idea is that can we replace the natural chromosomes in that cell - and it's a eukaryotic cell, i.e. it's not a bacteria, it's a much more complex cell - can we then replace it with synthetic semi-designed DNA?
Kat - So, can you engineer a yeast to be exactly what you want it to be?
Paul - Yes and also, the great thing about this project and it's a world consortium. It's led out of the US by a guy called Jeff Boker who's in New York and Jeff set this consortium up. So there are people in China, people in Australia, people in Singapore, and in the UK Imperial College working on different chromosomes because there 16 chromosomes. It's a massive project and we're working on chromosome 11 and this is led by my colleague Tom Ellis in our centre. He's a fantastic synthetic biologist. And so, Tom has been leading on the synthesis of this and I've been involved in the project as well. The idea is that not only can you get a yeast cell which has been essentially controlled by designed synthetic chromosomes - that's what we're aiming for - but also, we've built into the designs, bits in the DNA sequence that allow you to essentially scramble it. So, this is the really cool bit. So, we can induce production of a protein in this synthetic yeast cell which will essentially start recombining differentparts of these synthetic chromosomes. So, we could end up scrambling the whole genome and then plating the yeast off and looking for different phenotypes, like a yeast that produces much more alcohol might be interesting or a yeast cell that...
Kat - Woohoo! Or a nicer bread?
Paul - Or a nicer bread or a yeast that grows at higher temperature. From an industrial biotechnology point of view, it's a terribly important project because yeast is a very good cell for manufacturing chemicals or pharmaceuticals or drugs, or whatever. And so, if we can build these kind of interesting yeast strains that have particular properties that are very useful for manufacturing, are very useful for whatever, that could open up a whole plethora of different applications. And finally, we will also learn a lot about biology because by scrambling all these genomes up, we're going to learn all sorts of weird stuff about why is a cell growing at 42 degrees for example. It shouldn't be. It should be dead. The wild type strain doesn't - so, what have we done to create those properties? It's going to be very interesting.
Kat - Does it feel a little bit strange to effectively be evolving yeast or I suppose, to use a terrible phrase, to be playing God to yeast
Paul - Well, I don't consider playing God. I mean, I consider it to be more engineering. As you probably realise, we're very focused on the sort of engineering approach to synthetic biology. Yes, we're exploring accelerated evolution and I think that's right. You were right in saying that. You can't select naturally different strains but it takes a long time. It doesn't always work. This is an acceleration of that process, but it's also something we've never been able to do before which is explore the boundaries of chromosome organisation, how genes are regulated and turned on and off. Just by scrambling everything and then look and see what's happened and what it produces is a really interesting experiment, just on its own right.
Kat - Paul Freemont from Imperial College London's synthetic biology hub.
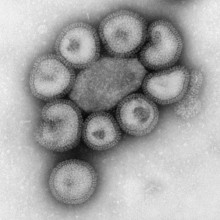
09:39 - Paul Kellam - Blocking the flu
Paul Kellam - Blocking the flu
with Paul Kellam, Wellcome Trust Sanger Institute
Kat - We're heading into the flu season, and people at risk such as pregnant women, the elderly and those with certain health conditions are being advised to get a flu jab to help them stay free of the virus this winter. But could doctors soon be adding genetic risk into the mix when deciding who might benefit most from a jab? Paul Kellam, from the Wellcome Trust Sanger Institute outside Cambridge, has been investigating whether screening for variations in a gene called IFITM3 could form the basis of just such a test.
Paul - A particular gene, and it's known as interferon inducible transmembrane 3 or IFITM3 for short, has a variant in it and that variant is associated with more severe influenza disease than if you don't have that gene variant.
Kat - Is that a straight one to one relationship? If you've got that gene variation, you are going to end up in the hospital when you catch flu?
Paul - So genetics really doesn't work like that. There are modifiers. What you're looking at is an increased risk so an increased relative risk of developing disease given your genetics. This gives you a four or so fold increase risk of developing disease. So, these aren't absolutes. These are modifiers. Not everybody that ends up with severe disease, whether it's influenza or other viruses, will have this variant. So, it's not a variant that is universally everywhere, causing everything that we see, but nonetheless, in the subset of people, this has a very clear association.
Kat - It certainly seems to be important as you say, but what's it actually doing. If genes are the instructions that make proteins in cells, what's going on at this cellular level?
Paul - So, the gene variant unfortunately, we do not know its mechanism of action and that's true of a lot of human genetics. What we do know, what this protein does and therefore, by implication, we can start to infer what the gene variant does. So, the protein is expressed in the cell. It decorates what are known as endosomes and these are internal vesicles that viruses use to get from the outside of the cell to the inside. So, the virus gets taken up at the cell surface, ends up in an endosome, the endosomes lower their pH as they get deeper and deeper into the cell, and that low pH causes the virus to trigger and to breakout.
Kat - So, they're becoming more acidic.
Paul - They're becoming more acidic, that's right. The endosomes acidify as they go deeper into the cell. What this protein seems to do is make it much more difficult for the virus to break through the endosome membrane. Whether it makes it more sticky, less fluid is not really clear, but they seem to create an extra hurdle, an extra barrier for the virus to get out of the endosome. This is enough to really, really change the disease course.
Kat - We've talked about flu and about swine flu. Is that the only kind of virus that this protein is working against?
Paul - Well, no and that's one of the interesting things about this gene and this class of genes. Because they are blocking common features of what viruses need to get into cells or use to get into cells, then they have a broader activity spectrum. So this particular gene will work against viruses such as dengue virus, other influenza viruses, yellow fever virus, Ebola virus, to name but a few. So, we're starting to look at broad spectrum antiviral molecules.
Kat - A lot of drugs that we have target molecules in the cell and molecules in pathogens and stop them from working, they block them. I'm thinking of some of the cancer drugs that block the molecules that tell cells to grow. But what you're trying to do here is boost the level of this protein. How can you do that then, because that's kind of hard?
Paul - It is kind of hard. Fortunately, biology helps us in this way. Proteins turn over in cells. some proteins turnover very quickly, some turnover very slowly, and have much longer half-lives.
Kat - So, that's kind of creating it, destroying it, creating it, destroying it?
Paul - That's right. Some recent work that has come from a group in the US has identified the mechanism for turning over this protein IFITM3. What they found is that there's a particular pathway in the cell that helps to destroy IFITM3. Now, we know the proteins that interact together then the theory is if you block that interaction specifically, you'll leave more IFITM3 on the endosomes and that will increase your antiviral effect.
Kat - And that sounds potentially like, "Wow! This is an antiviral drug that could work for lots of people against all kinds of viruses!" So, when are we going to get it?
Paul - Well, it's always a difficult question. This is very much at the early stages. This is understanding the basic biology and mechanism. And then you've got to start on that long and hard road of drug development. In the end, you've got to be able to find a molecule that really works, a small drug, and that that drug has a good therapeutic effect. That means the beneficial side is much, much bigger than the detrimental side of any toxicity associated with the drug. As yet, we don't have any evidence that you can do that very easily in these sort of systems.
Kat - Turning more to the genetics angle, are there any ways that we could use this genetic information that people with certain gene variants respond in different ways to the flu? Could that be useful as well?
Paul - Well, I think that's a very interesting way of thinking how you can get quick wins from this sort of information. One of the things that people can relate to very easily is how infectious diseases feel. People have gastroenteritis, they have respiratory tract infections. As you say, you've had flu. You know what it feels like. Therefore, if it's possible to understand the genetics that changes how you respond to a pathogen and identify people that are at higher risk of severe disease then you have a way of stratifying them if you like, for treatment. Now for something like influenza, that treatment is very straightforward. It's identify those individuals and encourage them to have a vaccination because the vaccine tends to protect against the strain of flu that's coming. So, that's a very easy way of thinking how genetics can go through actual information that people can really relate to. The other way of thinking about it is in severe influenza seasons when people are coming in to accident and emergency wards and the clinicians are faced with who is going to have flu but get better as opposed to those that are going to have flu and get worse and really need to worry about them as a clinician. At that point, stratifying based on genetics starts to give you that extra information that hopefully becomes clinically useful.
Kat - That was Paul Kellam from the Wellcome Trust Sanger Institute.

16:15 - Stephen Chambers - Selling synbio
Stephen Chambers - Selling synbio
with Stephen Chambers, Imperial College London
Kat - Now it's time to return to our theme of synthetic biology. Stephen Chambers is CEO of SynbiCITE - a national centre set up to turn bright ideas in synthetic biology into real life commercial applications, by partnering researchers with companies. I started by asking him what kind of organisations he works with.
Stephen - It's the complete range of companies that we talk to. It's everything from multinational companies, right the way down to start-ups and a lot of researchers in the lab that are thinking about commercialising. So, they haven't even set up a company. They're just thinking about setting up a company.
Kat - In many areas of biology, perhaps the most obvious way of commercialising something might be to develop a drug for a disease or cancer or something like that. In terms of synthetic biology, what are the kind of applications that people are starting to look at? Where are these engineered organisms potentially being used?
Stephen - The obvious ones are always like the biomedical area. And they are always there. The exciting thing about synthetic biology is it's much broader than that. That's the exciting thing about synthetic biology. So, it's not just the biomedical technology. It's also agritech, it's chemical industry, it's remediation, it's manufacturing. It's every kind of production you can think of. So, it's much, much broader than just the space that biotech would have had in the past.
Kat - Has it been a challenge trying to get the scientists working in this field to go, "Actually, I have potential commercial applications for my research." Sometimes scientists do think, "Oh no! This is research. This isn't industry."
Stephen - Yes, it is a challenge. But that's one of the roles that we have here to educate the scientists to give them confidence that they can start up their own business and that it is possible. We have a large number of mentors to help them through that process. We have educational programmes, we have funding programmes as well. So, it's all geared to enabling those researchers with the confidence to step outside the lab and commercialise their ideas.
Kat - What sort of different areas of commercialisation are there for synthetic biology technologies?
Stephen - The way I think about it and the way that people typically break it down is that there's two areas. They call it enabling and enabled. So, the enabling is a large number of tools and reagents, things that - you know, it's almost like foundational technology behind synthetic biology.
Kat - The kind of the Lego bricks that people can take and build things with?
Stephen - Exactly, almost the picks and shovels. So, there's that element to it and that's what we call enabling. And then there's the enabled. That's the end product so that's the interesting thing. That's the fine chemical that's being made, that's the drug that's being made. I have a lot of companies that come through me and it's amazing the types of different things they're making, everything from super cosmetics, right the way through to medical devices, apps. So, it's really, really broad. It's very hard to define. It's one of the challenges but also, what makes it so exciting.
Kat - What sort of products and processes are people starting to use this technology in?
Stephen - Well just to talk about my own ecosystem and the companies that I'm interacting with, I have companies that are making hard to make chemicals. I think you'd probably describe them as that. They're making new drugs. I've got one company that's launching a product in a couple of months which is a cosmetic, and then I have another company that is launching some reagents and tools in synthetic biology. So again, it's very broad and full of different applications.
Kat - Do you think that there's kind of a killer app? There's something out there that synthetic biology can solve for us that nothing else has been able to solve yet?
Stephen - Obviously, as a technology, we're always looking for that killer app that's going to get the public's attention. But I don't think I've seen one yet. There are some very exciting tools out there, but I don't think we've seen the killer app yet.
Kat - How do you see this technology expanding, changing, developing over say, the next 5 to 10 years? Where would you hope that we would end up?
Stephen - I think one of the big pushes for synthetic biology is going to be around sustainability where synthetic biology has its roots in. synthetic biology is a very interesting phenomenon. It's very much kind of grassroots based. It's not a top down thing. There's this large community of very enthusiastic participants in synthetic biology and they are pushing it. They have a very utopian idea about synthetic biology and they are really pushing the green side of things. I think where we're going to see a lot of movement in synthetic biology that's going to be attractive to the public is around the bioremediation, around environmental kind of concerns.
Kat - Can synthetic biology save the world?
Stephen - Definitely.
Kat - Stephen Chambers from SynbiCITE.

21:35 - Geoff Baldwin - Building nanocages
Geoff Baldwin - Building nanocages
with Geoff Baldwin, Imperial College London
Kat - One person who's working hard on a new biomedical application for synthetic biology is Imperial College's Geoff Baldwin. He's working on new ways to target cancer drugs specifically to tumours, leaving healthy cells unharmed, with the help of some specially-constructed nanocages.
Geoff - Normally, when you take a drug whether it's a tablet or it's injected then that drug disperses across your entire body and that's why you have toxic effects from drugs where they are acting elsewhere other than the site you want them to act. So, the general proposition is, can we make drug action more specific by targeting that drug to the site where we want it to go in the body?
Kat - So for example, if you have a cancer to make it go to that cancer and not anywhere else?
Geoff - Exactly. So, a lot of the problems associated with anticancer drugs is that they're very toxic. So, they're very good at killing cancer cells, but the problem is they're also quite good at killing other cells in your body too. That's why a lot of chemotherapy treatments for cancer have pretty nasty side effects. So, our idea is to shield these drugs by encapsulating them in a nanostructure so that these drugs, once they're administered into your body, don't have the same toxic side effects.
Kat - So, it's almost like smuggling them in to get them to just the right place.
Geoff - Indeed, yeah sort of a Trojan horse type affair where a nanocage targets them to the cell and we can direct those nanocages to specific cells. And then once they're at that site then what we want them to do is to release that toxic cargo at the site where you want the treatment.
Kat - Surprise! It's a drug!
Geoff - Yeah, indeed.
Kat - So, how are you using the tools of synthetic biology to do this? How do you make these nanocages and trap the drug inside them?
Geoff - What the approaches of synthetic biology have given us is the sort of a modular approach to biology and by treating these nanocages as modular reformable structures where we can take them apart and then rationally put them back together.
Kat - Effectively, like biological bricks.
Geoff - Yeah, because we've developed quite a lot of foundational tools in synthetic biology around assembling DNA, we can actually use that to recapitulate different modifications and different formulations of these nanocages very quickly. That has made us able to refactor and reform these nanocages and sample lots of different variations of them to pick out the ones that work best for our purposes and we can do that much more quickly and much more efficiently than we could previously.
Kat - How do you make them? Is it the kind of approach where they're being made in bacteria or yeast? How do you actually manufacture these little cages?
Geoff - We just express them in bacteria. And so, our nanocages are made of protein. So they are natural proteinaceous material. We take these nanocages from a variety of sources. Some of them are naturally human proteins but we can express them and purify them as individual proteins and then reform the nanocages from these purified proteins.
Kat - How do they know where to go when they get into the body? How do they know to go to a particular tumour or to the site of a disease?
Geoff - There are two ways that nanocages can be effective in terms of targeting. One of the ways that a number of people are exploiting in treatment of cancer cells with nanostructures is just that a lot of cancers have quite leaky vasculature.
Kat - That's the blood vessels that feed them?
Geoff - Yeah. So these small nanostructures are able to naturally leak out of the blood vessels at the site of tumours and they will have naturally an increased residence time at the sites of many cancers within the body. That's not true of all cancers, but it's certainly true of some solid tumours. The other way we're looking at is to combine these with antibodies and then use the specificity of antibodies against specific cell markers that are known to be associated with cancer cells and so that we can have more active targeting in that way. So, this is something which is currently being explored as to what are the best routes for targeting of these species.
Kat - Where in the process are these nanocages? It's always that, "How long is a piece of string" kind of question, but how far through the process from bright idea in the lab to "Here's the treatment that patients could receive"?
Geoff - We are a long way from having this as a treatment because any drug that you're going to administer into the body has a lot of regulatory hurdles. We've currently looking at developing this as an in vitro drug testing platform so to use these for delivering drugs in cells in the lab. The other area of interest we're looking at is whether we can use these within the gastrointestinal tract. So, there's a lot of difficult to treat tumours that are associated with the digestive tract - pancreatic cancers, liver cancers, oesophageal cancer - that are currently very difficult to diagnose and catch early, and treat effectively. And so, the idea of having a nanocage which is very visible, we can make these things highly fluorescent, and then you have the potential for endoscopic delivery rather than delivery through the bloodstream.
Kat - So you literally put a tube into the organ or the tumour and just like, pop! In you go.
Geoff - Yes. So, you'll be able to kind of spray the surfaces where you're trying to investigate for cancers within the patient. If they light up and stay lit up then you will know that the cancer is there and then what we are also hoping is that you can then perhaps combine that with directed therapy where once you see the tumours lit up, switch phasers to blast and then lead to drug dissolution where you want them.
Kat - Geoff Baldwin from Imperial College, London.

28:03 - Gene of the Month - Curly
Gene of the Month - Curly
with Kat Arney
And finally it's time for our Gene of the Month, and this time it's Curly. First described almost a century ago and a firm favourite with fruit fly geneticists everywhere, the Curly mutation gives flies characteristic upturned, curved wings, rather than their normal straight ones. Because these are easy to spot, it's often used as a marker when doing breeding experiments, so researchers can easily find the flies they're interested in. But for nearly a hundred years, researchers haven't known exactly which gene is responsible. Recently a team of scientists in New York pinned it down to a gene called duox, which makes an enzyme that creates very reactive chemicals inside cells, helping to stabilise the structure of the developing wing.
Pleasingly, Curly works together with another gene known as Curly Su, which helps to tie together protein molecules in the developing wing to give them a strong, straight structure. If either gene is missing, the wings don't form properly, creating the curly shape. In humans duox is found in the thyroid gland and the gut, although exactly what it's doing there is unclear. And it's also found widely across other species, suggesting that whatever it does do is pretty important for it to be preserved so widely through evolution.
- Previous What causes sleepwalking?
- Next What Keeps the Sun Burning?










Comments
Add a comment