This month we're delving into the junk in the genome - or, to put it more correctly - our non-coding DNA. Less than 2 per cent of the human genome contains protein-coding genes, so what does all the rest do?
In this episode
01:02 - Not just genes
Not just genes
with Wendy Bickmore, IGMM Edinburgh
This month we're reporting back from the Genetics Society Autumn meeting, held at the Royal Society in London. Over two packed days we heard a fascinating array of talks focusing on one of the biggest challenges in biology - understanding how our genes get turned on and off at the right time and in the right place. We know that we have around 20,000 human genes that encode recipes to make molecules called proteins, which are the molecular 'workhorses' that build our cells and keep our bodies functioning. We also know that when genes are read they care copied out in the form of a chemical called RNA, which is similar to DNA. But only a tiny fraction of our genome is made up of these protein-coding genes. So what's the rest? To get a flavour of the talks aimed at shedding some light on the dark matter of the genome, Kat Arney caught up with one of the meeting organisers, Professor Wendy Bickmore, director of the MRC Institute of Genetics and Molecular Medicine at the University of Edinburgh...
Wendy - What we realised was we were in this amazing situation now where thanks to advances in technology, and our computational power to store and analyse the sequence of the genome, we can actually study the whole genome of hundreds, thousands, millions of individuals. They might be individuals from different species and compare in between species, or individuals within a species, and different members of the human population. Or even between different cells within the same individuals so comparing cancer cells to the normal cells in the same individual. So we have that ability to capture that information and then we realise that actually, for the vast majority of that information, we can't do anything with it. So it's a great "stamp collecting" exercise, but it's a little frustrating if all you can do is sit and stare at the sequence. And that we realised the problem was, for the bits of our genome that code for proteins - so the protein coding regions of the genome.
Kat - The "honest to God" genes.
Wendy - Yeah, the proper...
Kat - The real genes.
Wendy - Yeah, the workers of the genome that actually code for messenger RNA that makes a protein that does stuff. We can make a fairly educated guess that if there is a particular change in the DNA sequence - a change from an A to a T, at that position - it'll actually affect the way a particular protein works. And therefore, we can say whether that change is going to matter for the individual, for the species, for the disease. And that's because we understand the triplet genetic codes - so three letters equals one particular type amino acid. So we've kind of got a grammar that we can understand what change means. But that part of the genome is only 2 per cent of our genome. So the 98 per cent of the genome is doing other stuff and we don't have that understanding of what a sequence change means because there's no code that tells us these three letters do this. So that's a huge challenge. So, we wanted to bring together people who were thinking about this in different ways. So people who are looking at evolution and what evolution tells us about the part of the genome that doesn't code for genes, so comparing different closely related species and looking at how sequence changes outside of genes. People looking at disease states for example, people using smart computational and statistical arguments to identify change that matters. And then people that are trying to understand at the level of molecules of what these changes might mean. I think what we've been hearing today in the meeting, I would categorise them into to two kinds of studies. There's those where the DNA sequence change is affecting, not a protein coding gene, but still a bit of the DNA that makes an RNA molecule.
Kat - We would call this like the non-coding RNA?
Wendy - Non-coding RNA is exactly the term they've been given. They do lots and lots of interesting different things in the cell so we know they have functions. We are starting to see that actually single changes in these RNAs can actually affect the way these RNAs work so fantastic examples where they actually change the shape of the RNA. So these RNA molecules fold into these wonderful knotted structures and just little tweak in the sequence can make new bulges and knots in the RNA which presumably affects the way the RNAs work. So, we're making pretty good progress I think in that but the bigger challenge looks like it's the sequence changes in the bits of the genome that don't even make RNA molecules. They don't even seem to make anything.
Kat - So we'll be calling these things like the control switches, the regulatory switches that are involved in turning genes on and off.
Wendy - That's what we think, yes. That's what we think these bits of DNA are doing, proving that is turning out to be really tough I would say, but it's very exciting. There's been some fantastic studies in fish. Let's talk about fish for example that live either in seawater or freshwater and looking at the changes in the shape and the behaviour of those fish, and seeing when you sequence the genomes of those two fish, the changes are in what looks like regulatory switches that are changing the way these fish behave or look. Starting to come out from human genome sequencing projects as well, evidence that there could be changes in some of these switch elements that are causing very severe human disease. They are probably contributing to common disease as well. And people are starting to develop assays in cells, in organisms using computational approaches to try and get some understanding at least of how these changes in sequence might work. But we're obviously a long, long way away from being able to really understand what these are. So there's progress being made. It's really exciting, but we know there's a big challenge ahead.
Kat - So, when the human genome sequence first came out back in 2000, 2001, I think there was this wonderful idea that once we have the human genome sequence, we would know all the genes and then we would know how it all works. And now 15 years later, are we like "Oh crumbs!" We're only scratching the surface.
Wendy - We are but we are excited by that because we're realising now how complicated our genome was. I guess one of the other disappointments I suppose from the human genome project was, everyone was expecting that we would have many more genes than flies and worms. We were all a bit deflated when we realised we only had approximately the same number of protein coding genes as a fly.
Kat - We're just sophisticated fruit flies.
Wendy - We are sophisticated fruit flies. We are more complicated than that and it turns out the reason why we're more complicated is we have more types of cells than a fly and we use a set of genes in much more sophisticated ways in those different types of cells.
Kat - So it's not what you've got. It's what you do with it that counts.
Wendy - It's what you do with it, indeed. That's true.
Kat - I think that sums up the whole meeting, right?
Wendy - Absolutely!
Kat - Wendy Bickmore, from the University of Edinburgh.
08:06 - Emma Farley - Sea squirts and switches
Emma Farley - Sea squirts and switches
with Emma Farley, UCSD
Kat - One of the most exciting talks of the conference came from Emma Farley, assistant professor at the University of California San Diego, whose work focuses on understanding exactly what the switches in our genome look like, and how they switch on genes - known as gene expression. And her studies have led her towards a rather unusual - not to mention squirty - organism.
Emma - So I'm trying to figure out how the instructions for development and cellular integrity are encoded in our DNA sequence. We have a set of genes in our genome. We kind of understand these genes, but we don't understand how these genes are turned on in different cell types. And this is the key for building all of the different cells of our body and for maintaining their health during adult life. So that's a huge question in biology at the moment.
Kat - I know that we have 20,000 genes but of course, we have lots of different types of cells. You can't have all genes on in all cells or we'd just be a blob, wouldn't we?
Emma - Exactly, yeah. We need like heart-specific cells in our hearts and so we need to turn these genes on with specific switches. We need cells that have expression of particular proteins leading to neural functions so that they're neurons and these genes need to be switched on by particular enhancers or switches in the genome.
Kat - This is interesting - the switches - because the actual genes is only 1 or 2 per cent of all the DNA that we have in all our cells. What is the rest of it? How much of our genome is these switches that turn the genes on and off?
Emma - So, we really have no idea. We have methods to detect these switches in the genome. So, we can detect them by the proteins that bind to them and particular marks on the DNA. Using those methods, we estimate that there are somewhere on the order of a million enhancers in the genome. And so, this really provide the instructions for when and where all the genes are deployed in the human body.
Kat - A million switches to 20,000 genes. That starts to make more sense now because I remember people saying, "What do you mean? We've only got 20,000 genes." But a million switches!
Emma - Yeah. Each cell needs to turn on different genes at different times and this gives them a key identity and allows them to become say, heart or nervous system. It's really the diversity in these switches and deploying these genes that allows us to have all these different cell types.
Kat - Let's zoom in a little bit more. So what is a switch? What does it look like? Is it a stretch of DNA? What do we know about them right down on that DNA level?
Emma - So, switches are pieces of DNA in our genome. They're on the order of a hundred to a thousand base pairs, so a hundred to a thousand letters of DNA sequence. They contain within them sequences to which transcription factors, these proteins, bind and the binding of these proteins to the switch is like somebody with a finger turning a switch on and turning a light bulb on, and that leads to expression of particular genes.
Kat - So, what are you doing to try and unpack these switches and work out exactly what they look like and how they work.
Emma - I'm trying to understand how the sequence of DNA that makes up a switch encodes the information for a particular expression pattern. So during development, we have genes expressed in the nervous system or in the heart or in our skin cells. So, I want to understand how the information to turn them on in those locations is encoded in our DNA sequence within these enhancers. And so, we know that there are proteins in our cells known as transcription factors that bind to these switches. But that is not the whole story because the same transcription factors bind to different switches and turn on different patterns. So for example, there's one called ETS and one called GATA - these two transcription factors - they turn on expression in the heart, they turn on expression of genes in the gut, and they turn on expression of genes in the nervous system. But the switches are slightly different. There are different sequence identities of the particular binding sites that allow different expression and I'm trying to understand the regulatory principles that can translate enhancer sequence into expression patterns and find principles that can help us much like the relationship between coding DNA leading to protein sequence - what is the relationship between enhancer sequence and expression pattern.
Kat - So it's not as simple as say, AAGTC, that transcription factor always sticks to that sequence and always turns on a gene. So, it's more subtle than that?
Emma - Yes. So, if you look in the genome for a particular transcription factor binding site, there's probably 10 million locations where that binds and not all of those are functional. The question is, where in the genome do proteins bind and mediate function. A key component of this is combinatorial control. By this, I mean when you have two different transcription factors and the combination of the two leads to an expression of something. This added complexity allows this small set of genes to turn on expression in many different cell types.
Kat - So it's got to be the right kind of looking set of letters. You've got to have the right factors, right time, right place, and then you get the right genes turned on to make the right stuff.
Emma - Exactly. It sounds simple but we're still trying to work out exactly why certain sequences turn on in certain locations and not in others. And then the question beyond that is, what happens when people have mutations in their switches, and how do mutations in switches change gene expression and cause disease?
Kat - You're using quite an unusual organism to study this kind of question. Tell me a bit more about that because I guess people think, "Human genes, let's use humans." I guess you can't do these experiments in people.
Emma - Exactly, so I used an organism called Ciona intestinalis. The common name for it is the sea squirt because when you pick them up, they squirt water out. So they live in the ocean. They're sessile so they're attached to rocks and the bottom of boats. They're a long see-through sort of object and they have water that pumps in and out of them through two siphons. But they have a heart and when they're embryos, they look very similar to us, developing chordates. They have a notochord which is really important for development of the nervous system and they have a dorsal nerve chord which is the key definition of a chordate embryo. And they're actually the sister group to vertebrates. So they're basically our closest invertebrate relative.
Kat - So, our little sea squirty sisters. How are you studying these switches in these organisms?
Emma - So, these switches as I said are pieces of DNA sequence in our genome that can be somewhere on the order of a hundred to a thousand base pairs. We want to understand within that region, how does that region code for a precise expression pattern. It's really challenging to understand and look at this question. We've taken an approach where we want to change the sequence of this switch and see how changes in sequence of the switch impact gene expression, and then work out how you code for precise expression. If you think about a small sequence of DNA, you've got your A, C, T, and G letters. So, if you want to change each position to any four other positions, even on a small piece of DNA about 70 base pairs long, you're talking over 10 to the 30 possible sequence variants.
Kat - Wow! That's a lot.
Emma - Exactly. So, if we want to understand how a switch encodes tissue-specific expression, we need to be testing hundreds of thousands, millions, maybe even one day, billions of sequence variants. If we want to understand how variation in the sequence impacts where the expression is, we need to be doing this in intact developing embryos. And so, Ciona intestinalis offers a system where we can obtain millions of fertilised eggs and then we can introduce millions of variants of a particular switch and then see where these switches are turning on. And from that, we can start to find principles about how switches turn on in precise locations and what sorts of violations in these regulatory principles lead to disease.
Kat - What seems to be the indications so far? What do we know about what makes a good switch?
Emma - So, we had some really surprising results. If you think of a switch to turn on in the nervous system, you might think this switch is built to be really great at turning on in the nervous system. But what we found was actually, you want to make sure you don't turn on anywhere else. So you want to be bad at turning on in all other tissues and to do that, you have to be quite poor at turning on in the nervous system. So we found that you don't use optimal features in the enhancer. You're actually using these suboptimal features and this ensures that you don't turn on in the wrong place.
Kat - This is putting me in mind of the light switch in my hallway where you just have to really smack at it to get it to work. but in the end, it will go on when you want it to.
Emma - Exactly. You want something to turn on only in the right place because turning genes on in the wrong place is known to cause disease. So we think that those mechanisms within the genome that have allowed these switches to be turned on only if you press them in exactly the right way.
Kat - Emma Farley from the University of California San Diego. And now it's time to find out about the latest news in the world of genetics.
18:00 - Shooting the messenger
Shooting the messenger
Huntington's disease is caused by carrying a faulty version of the gene encoding the Huntingtin protein. Multiple extra copies of a short repeated sequence in the gene lead to an expanded mid-section of the protein, which causes major problems in nerve cells.
Now scientists at the Centre for Genomic Regulation in Spain have shown that it's not just the faulty expanded protein that causes disease, as was previously thought, but also the RNA messenger itself that is read from the gene when it is switched on.
Writing about their findings in The Journal of Clinical Investigation, the team showed that interfering with the expanded section of the gene at the level of the messenger RNA could make a difference to the signs of disease in mice with a model of the illness.
Huntington's disease is currently incurable, so the team hopes their findings will open the door to new approaches for treatment based on targeting the Huntingtin RNA as well as the faulty protein.
19:03 - Evolving Ebola
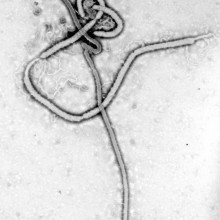
Evolving Ebola
Scientists in the US and UK have discovered that during the 2013-2016 Ebola outbreak, the virus picked up genetic mutations that increased its ability to infect human cells, publishing their findings in two papers in the journal Cell.
Normally Ebola hides away in animals - possibly bats or monkeys - and only rarely infects people, and outbreaks are short-lived. But by the end of the most recent prolonged outbreak in West Africa, more than 11,000 people had died.
By tracking through genetic data from samples of these infected patients, the scientists were able to discover that the Ebola virus had picked up mutations in the gene encoding a protein on its surface - known as a glycoprotein - making it more infectious to human cells, although not to cells from other animals such as bats.
This might have contributed to the viruses' rapid and deadly spread. By studying the patterns of evolution in outbreaks like this, scientists hope to figure out how to beat viruses at their own game, making us better prepared to avoid such huge death tolls in the future.

20:11 - Monkey business
Monkey business
It's now well-established that humans and Neanderthals lived alongside each other and interbred for thousands of years, and genetic researchers have found that up to around 4 per cent of our modern human genome came from our Neanderthal cousins.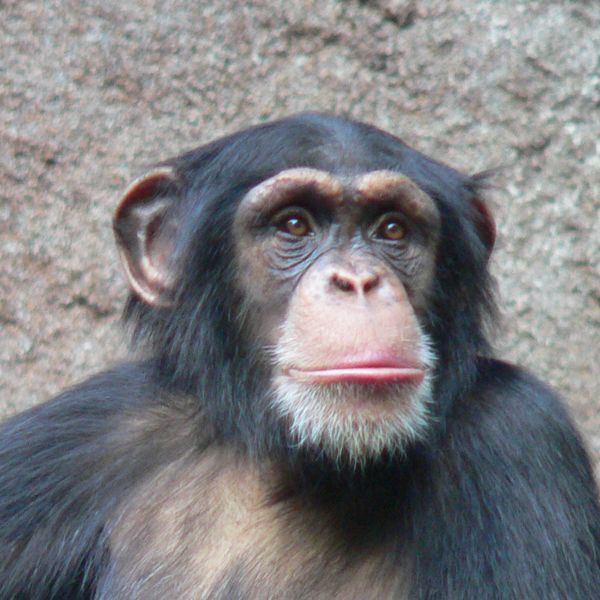
Now scientists from the Wellcome Trust Sanger Institute in Cambridge have found that the ancestors of chimps and bonobos also got up to the same monkey business back in their evolutionary history.
Writing in the journal Science, the team analysed DNA from 75 chimps and bonobos in 10 African countries and showed that one percent of chimpanzee genomes are derived from bonobos, even though the two species officially split between 1.5 and 2 million years ago.
There seems to have been at least two periods of subsequent interbreeding, with the last happening as recently as 150,000 years ago. The scientists also discovered a strong link between a chimp's DNA sequence and its country of origin, which could be useful for conservationists trying to work out where chimps are being illegally captured.

21:31 - Duncan Odom - Swapping switches
Duncan Odom - Swapping switches
with Duncan Odom, CRUK Cambridge Institute
Kat - It's time to return to the Genetics Society autumn meeting. Every year the Society awards a number of medals to some of the leading geneticists around the world, who are invited to give a guest lecture at one of the society's meetings.
This time around there were three worthy winners - Felicity Jones from the Max planck Institute in Tubingen, Germany, who picked up the Balfour medal for 2016; Ben Lehner from the Centre for Genomic Regulation in Barcelona, Spain, who won the same award last year; and Duncan Odom from the Cancer research UK Cambridge Institute, who walked away with the society's Mary Lyon medal. I asked him to give me a run-down of the work he and his team are doing to figure out the differences in the control switches between different species and how they contribute to the incredibly diversity of life on the planet.
Duncan - The key piece that we're trying to exploit is the fact that nature has done a lot of the experiments that we in the laboratory would want to do anyway. So, there's a huge amount of sequence turnover which clearly is driving a huge amount of regulatory turnover as well. My laboratory specialises, along with our collaborators, in trying to connect these two things - by actually doing experiments in tissues from a lot of the species where we have genetic sequence information but we don't necessarily have any functional genomics data and then connect those two things.
Kat - So what kind of species are you looking at and what are you looking at when you get their DNA sequence and line it up?
Duncan - The species that we tend to focus on are often ones that the entire community has chosen in the sequencing space to be important. These include a number of species like dog and cow, rat, mouse that a number of laboratories have a long history of working on both for agricultural reasons as well as for evolutionary biology reasons - their evolutionary separation might be almost the same length as human and mouse for example. Those species, we didn't choose actually. The community chose them to sequence to the highest standards to start with. What my laboratory specialises in is adding on another layer of functional genomics data that allows us to go in and connect the sequence that the community has developed with functional data that we can then analyse to try to connect the sequence evolution with functional evolution. Does that make sense?
Kat - Let's unpack this a little bit so, if we think about just the genes that make proteins and we think about them across lots of different species of say, mammals.
Duncan - Those are largely conserved.
Kat - They're pretty much the same so that's what we know. So then you're looking at the - what is probably wrongly called the "junk DNA", the "non-coding DNA" all the other stuff of the genome.
Duncan - Yes.
Kat - So, how do you then start looking at that? What sort of things are you looking for when you're looking for these important control switches?
Duncan - It's much straightforward than you might imagine. There are very specific histone marks which are known to associate with active promoters for example. So, by simply mapping where those are in the genomes within specific tissues, we often use liver, you can identify what regions in the non-coding genome as you accurately put it, are functionally deployed or at least prepared to be functionally deployed in the cells of interest.
Kat - And by histones, these select the wrapping, the packaging proteins of DNA.
Duncan - Yes. Those are the highly positively charged proteins that DNA coils around in the nucleus to take what's 2 metres worth of genetic sequence and compress it down into the micron scale.
Kat - In your studies, was there anything particular challenging to figure out how to do?
Duncan - So, we obtained whale and dolphin material from a British cetacean stranding investigation programme. Those species have very poorly sequenced genomes. One of the challenges that we had was to try to align sequence information that we got from our functional assays back to a very fragmented genome. That was one serious challenge that a lot of the more unusual mammals that we were able to obtain post mortem tissue for posed.
Kat - What's kind of the take home message from the research that you found when you're looking at these control switches between species and comparing? I mean, we know that the genes are all roughly the same pretty much. What about the switches? Can we say they're the same too?
Duncan - That was one of the major discoveries that we've made is that most of the switches in fact are quite highly divergent and they change very, very rapidly. Even ones that are near genes that are thought to have conserved control, it turns out that oftentimes, this control is not conserved - in particular switches that have to be turned on have changed between species. It would be sort of like having one light in the centre of a room and in human, you flip the on switch from the bathroom door. But in mouse, that's migrated to the hall door. It's still the same switch. It might even be on the same side of the room but it's nevertheless in a different place.
Kat - I love the idea that we have very, very similar genes in different species of mammals, but different mammals look different - you've got things like whales, through to mole rats, to humans, to dogs, to cows - are these switches really evolution's playground I guess?
Duncan - That's a good way to put it and that's definitely true.
Kat - Here at the Genetics Society Autumn Meeting, we've heard a lot of stories about these control switches, how they work, people searching for them, people trying to define them. What do you think are really the big questions we still need to answer here?
Duncan - Ultimately, the tools that we had developed based on the very high conservation of protein coding genes are largely inadequate to understand the functional constraints and how they express themselves through non-coding DNA. That to me is one of the key messages from this meeting - that a lot of the hopeful and understandable approaches that we were thinking would naively solve some of our questions around what human genetic variation does, a lot of these tools are simply inadequate and we really have to develop a different framework for understanding them. That to me is one of the major take homes from this conference.
Kat - Duncan Odom, from the Cancer Research UK Cambridge Institute.

27:59 - Gene of the Month - Ouija Board
Gene of the Month - Ouija Board
with Kat Arney
And finally it's time for our Gene of the Month, and because we've just passed Hallowe'en, I've picked the scary-sounding fruit fly gene Ouija Board. In fact, this isn't the only creepily-named gene - it's one of a family of so-called Hallowe'en genes, with names such as spook, spookier, phantom, disembodied, shadow and shade.
Although Ouija Board was discovered in 2015, the rest of its terrifying companions first turned up in the late 1990s, when researchers noticed a series of fruit fly embryos carrying genetic mutations that made them look pale and ghostly, lacking the solid cuticle that normally forms around the developing larvae. Digging down into the detail revealed that the Hallowe'en genes are all involved metamorphosis and moulting in insects and other arthropods, working through hormones called ecdysteroids, which are - as the name suggests - steroid hormones.
It turns out that Ouija Board is responsible for switching on the spookier gene, which encodes an enzyme involved in making ecdysteroids. And although humans obviously don't have sturdy cuticles or exoskeletons we do have steroid hormones that are involved in controlling all kinds of aspects of our health, reproduction and metabolism, and there are similar genes involved. But then they go wrong, they can cause serious illnesses - making this a real life human horror story.










Comments
Add a comment