This month we’re literally getting inside our genes, as we explore chromosomes through a 3-dimensional virtual reality art, music and science project. Plus, researchers are turning to bees, trees and more in search of new genetic systems, and our gene of the month has been around for a while.
In this episode

01:04 - CHROMOS: Get inside your genes
CHROMOS: Get inside your genes
with Mikhail Spivakov and Csilla Varnai, Babraham Institute
Kat Arney recently headed up to Cambridge to chair a fascinating event entitled CHROMOS, run by the Babraham Institute - a research institute nestling in the countryside just outside the city. This was no ordinary evening - guests were invited to step right inside the human genome, using virtual reality headsets to experience a writhing, dancing world of DNA created by visual artist Andy Lomas using real data about three-dimensional chromosome structure generated from experiments by the institute’s scientists.
In addition, leading electronic musician Max Cooper created an atmospheric soundtrack inspired by the science and the visualisation - that’s the music you can hear in the background. Before the event got started, Kat caught up with Mikhail Spivakov and Csilla Varnai from the Babraham Institute in Cambridge to find out how this unusual collaboration came to life.
Mikhail - The idea was that the analysis of the three-dimensional structure of the DNA and how the DNA is packaged in the nucleus, they are extremely important for understanding how cells use the DNA to do what they need to do. But it’s also extremely visually appealing and we felt that it would be a great opportunity to bridge our science with the art to actually tell people with more and maybe give them a bit more of a feel for how the DNA is and what it is.
Kat - So how did you go about trying to create this art because it’s not just visual art, is it? It’s music as well.
Mikhail - So that started from my reading an interview with Max Cooper where he mentioned that he has a PhD in computational biology. I was really excited and I thought actually, he must be the ideal person to do this kind of stuff with. So I contacted him via Twitter and he got really excited.
We had him over to Babraham Institute and it’s then that we decided to really create a project which is not just music about the DNA, music inspired by DNA, but also, the video and that’s when Max got Andy involved who he’s been working with for a number of occasions. It turned into a music video project about the DNA and how the DNA is arranged in the nucleus.
Kat - Where do the virtual reality component come in because it’s absolutely fantastic - I’ve had a go at flying through all these data?
Csilla - That was entirely Andy’s idea. So we commissioned Andy and Max to make a music video for us and then at one of our meetings, Andy showed up with a VR kit and we could all immerse ourselves in this virtual reality, looking at a molecule in three dimensions.
Kat - It’s like being in the matrix. You’ve got all these twisting things in space. You can put yourself right inside it and see all these coils moving around. What is it that we’re looking at when we’re inside that environment?
Csilla - So what we are looking at is the three-dimensional structure of DNA as it was in the nucleus of the cell. We can see the actual chromosomes moving individually and apart from this, we can also see red hot spots where there are active genes of DNA and they form various clusters, and that’s where all the genes are being read as recipes to make proteins for the cell.
Kat - This is your actual data from the lab. What is that data? Where have you got it from?
Csilla - I got this data from experiments that my colleagues do at the Babraham Institute. What the data comes from is effectively contact information about the DNA. So, we have a list of contacts about which parts of the DNA is in contact with other parts of the DNA. Using that information, my job is to reconstruct what the likely shape of the chromosomes was in the nucleus of the cell.
Kat - As a scientist, how do you feel when you're standing in that virtual environment and looking at your work spinning around you.
Csilla - It’s fantastic! I've seen it in two dimensions before or in various other pieces of software where you could drag and move the molecules around, but it is literally entering a new dimension.
Kat - Csilla Varnai, and before her Mikhail Spivakov from the Babraham Institute.
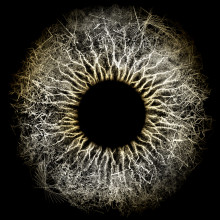
05:45 - Genes in 3-D
Genes in 3-D
with Andy Lomas, digital artist.
Visual artist Andy Lomas used Unreal - a virtual reality gaming engine usually used to make life-like shoot-em-ups and other games - to turn all this scientific data into 3-dimensional virtual reality. But how did he actually start doing it? Kat Arney found out.
Andy - That’s a good question. I was wondering the same myself when I started this project. So we got all these files, just genuine files and simulations that Csilla has been running. I haven't used Unreal that much but I just thought, okay, this game engine technology, it should be able to read in this data. It should be able to write a code that can directly read the simulation files and what can we then do with it and if we put it in that environment, we can hopefully look at things in different ways, try things out and basically, we’ve almost got like an environment to experiment with things.
So, I was learning to be honest, doing it. I thought it should work. I wanted to have something that interactive. I guess most of the work I've done in the past has been in film and visual effects where you really know what you're doing – you’ve got a storyboard, you know exactly what the target is, and you're often doing things that take very, very long amount of time.
I wanted to have something that allowed us to try sort things out really quickly which is why we wanted to try to use this game technology. So it’s a very big experiment and we wrote all the codes which just reads these files, reads them into memory, and builds the geometry based out of the data.
Kat - So, this isn’t like you’ve got a string of DNA and you're animating it to do what you want. You're actually taking real data from the experiments and just throwing it into this computer engine and seeing what it throws back at you?
Andy - Yeah, pretty much. It’s real data from Csilla’s research work on trying to reconstruct the shape of the nucleus, the DNA, and the chromosomes. So it’s actually real files from her which amount from the simulations she’s running which is three-dimensional data. It’s like positions of various locations on the chromosomes and positions over time and space. So it is already three-dimensional but I was having to turn that into visible geometry and also animated geometry.
I guess when we came to the institute and had a look at their work, what we they were showing us was the final shapes which is what most researchers are really trying to reconstruct. What we’re trying to explore in this work was, how were they getting to those shapes? How were they reconstructing this geometry? Actually, I don’t think before they'd actually looked at the data during the simulation, they were always looking at things at the end of the simulation.
Kat - It’s sort of, how did we get here?
Andy - Exactly, yeah – the sort of the process, as if it was almost like musical because they're doing these simulations that go through different phases. So just visually, it’s interesting. I mean, it just tells a story of almost like a discovery. You're going from something which is just this perfect mathematical, but almost this arbitrary structure to start with, because we really don’t know what the real shapes of the nucleus and the chromosomes are.
And then through all these forces, these constraints – things pulling things towards each other which need to be close to each other by the end – and just simulating with those literally, you heat it up to a ridiculous temperature so everything moves around with all this thermal noise and then gradually cools down. As it cools down, it solidifies hopefully into the right shape and so, there is beauty to that. You'll see it as a work in different movements.
Kat - It is an incredible thing to see because when you look at the video or indeed in the virtual reality simulation, you see this big column of almost like strings stacked up in a spiral, and then it starts to go crazy – it’s dancing all over the place and then slows down and solidifies into a sort of a ‘fluffy blob’, I guess would be the best way of putting it. How did you decide to turn that into a virtual reality environment where people could actually step inside and explore?
Andy - It was almost just because we could. I mean, our big thing with this was, “Let’s try this out in Unreal so we can experiment with it quickly, so that we can actually try things out, so we can look at what's in the data in a three-dimensional space, interactively.”
Unreal just has the opportunity to do things in virtual reality because it’s one of the two main game engine pieces of software you use to create to top end games including virtual reality games. So, it just has a VR preview mode and I have a virtual reality headset which I've been playing with and trying things out with and we just tried it out. Just like, “Okay, it should work. We should be able to hit VR preview and see.” Just how much you spatially understand the structure by being in VR actually really surprised me, even though I've done quite a number of things in VR before.
And then we worked it and turned this into a more interactive thing thinking like, how would you want to maybe visualise things, understand things, look at chromosome colours, which can influence colours off, look at activation zones? It was like, what would I want to explore in this set? But the first thing was, we could just hit that button, VR preview and have a look at it, and we looked at it and okay, that’s great!
Kat - That’s cool! As an ex-scientist whose field was this kind of thing, I was really fascinated by going from the straight line of DNA to the writhing library of genes and how they're all interacting, thinking about it in a three-dimensional way. When I was a scientist, there weren’t these kind of tools and what you’ve done here is an art project. But could we develop this kind of tools for scientists? I think it would be incredible to sit inside my data and virtually pick it apart, see it in space around me?
Andy - I think absolutely, yeah. I mean, we have started talking with people at Babraham Institute exactly about that. First, wouldn’t this be great to take much further as an art and science project for science fairs and things like that? And also, for all the genuine research – if you're actually generating structures and particularly interesting questions like, where are the locations of activation? Is the location really on the interphases between the different chromosomes and things like that?
Actually, spatially understanding the structure when it’s really complex – it’s this fluffy, really complex shape. It’s a very, very difficult structure to understand by just looking at like a picture on a computer screen. If you see it in VR, it feels like it is there in the room with you. It gives you just intuitively so much deeper understanding of that structure. I think particularly, it’s those really complex forms like these folded chromosomes where that’s really important.
Kat - Of course, the sci-fi nerd in me loves the idea. If you see the films, the scientists kind of go, 'bzzz' and the data appears like hovering over this table. Are we going to get there? Are we going to get to be able to probe our scientific data in three dimensions? I’d love that.
Andy - I guess so. I guess one thing is because we are genuinely using the real science data, it does take about 2 minutes to start up at the beginning as it loads all the data files. So, big data takes quite a long time and game technology, and these graphics cards in computers now have incredible amount of power to visualise really, really complex difficult spatial datasets. So it’s almost like you're riding on top of the people playing first person shooters to help you do real science.
Kat - Andy Lomas, and the piece you heard earlier was Chromos, by Max Cooper - the title track of his new EP inspired by the project, which is available now on iTunes. There are also a couple of amazing videos on Youtube, combining Max’s music and Andy’s visuals, so head to the podcast page at nakedcientists.com/genetics to find the links or search online for Max Cooper Chromos.
Videos:
https://www.youtube.com/watch?v=oyxwoLHFv_U
https://www.youtube.com/watch?v=gq7YwOIVW1Y

13:60 - Buzzing about bee genes
Buzzing about bee genes
with Paul Hurd, Queen Mary University of London
Regular Naked Genetics listeners will remember that a couple of months ago Kat Arney went to the joint spring meeting of the Genetics Society, the British Society for Developmental Biology and British Society for Cell Biology, held at the University of Warwick. One strand of talks explored so-called newly tractable systems - new model organisms that geneticists are finally able study for the first time, flippantly summarised as ‘bees and trees’. To get the buzz about our insect friends, she chatted to Paul Hurd from Queen Mary University of London.
Paul - So what fascinates me I guess are honeybees. They fascinate me for a number of reasons. They're important pollinators, they're important for maintaining genetic diversity in flowering plants, and plants of course are required for all animal life because they fix carbon from the atmosphere and produce carbohydrate. So they occupy a pretty crucial place in ecosystems.
From a genetic point of view, they're interesting because a bee genome, a bee larva doesn’t just have the capacity to become a bee – they can become three different types of bee. The question is, how is it that one genome can become three different animals? What is it that enables that to happen?
Kat - So this is more complex than in humans, we have males and females. This is much more complicated than that.
Paul - I think it probably is more complicated. So obviously in humans, we all come from a zygote and that zygote has the potential to form 200 or so different cell types. For bees, that same zygote has the potential to form 200 different cell types put that up in three very different organisms. So, I think it is more complicated, yes.
Kat - So introduce me to the different characters in the honeybee hive. Who do we have in there?
Paul - So top of the pile is queen bee. Queen bee is the largest insect in a hive. She’s the reproductive female - she can lay fertilised or unfertilised eggs. If she lays an unfertilised egg, the resulting larva becomes a male which is also reproductive. If she were to lay a fertilised egg, which is what she tends to do most of, that larva will grow and develop into a non-reproductive female worker bee. So two females bees in a hive, one can reproduce – the queen - one can't reproduce – the worker.
Kat - What do we know about what's going on in terms of the control of the genes that are doing this switch and controlling these different types of bees?
Paul - So this is what we’re interested in looking at. Honeybee caste determination whether you become a queen or a worker is dependent on diet. So what the larvae eats within the first three or four days of it hatching actually determines what it will become as an adult. If the larva eats royal jelly, a specialised substance, a nutritional substance produced ironically by the workers, the larvae develops into a queen. If the larva is fed boring carbohydrate-rich nectar pollen, then she’ll become a worker.
Kat - This is a switch that’s affecting the outcome basically based on diets. That seems quite strange to determine something as important as, are you going to be able to reproduce in your life or not.
Paul - Absolutely. I mean, the bees really are what they eat. There has never been a clear example of an organism really is what it eats. The reason why we’re interested in this is because the two bees are genetically identical, but what they eat changes what they become, we think this is happening at an epigenetic level, the layer of information on top of the genes which controls genome output.
Kat - So there's increasing interest in humans about how the environment thinks we do, the things we’re exposed to, the things we eat influence how our genes turned on and off this sort of layer of tags and switches. What do we know about how that’s happening in bees that’s making the switch between queen or worker?
Paul - We don’t really know. That’s one of the things that we’re looking at, but to some extent, the components of royal jelly do at least give the impression that it’s a pretty good epigenetic diet. So there are things within that diet that we think will be directly affecting gene expression, the switching on and off of genes, and it’s that that we’re trying to tease apart.
Kat - There are some health food shops that will sell you royal jelly and stuff like that, and people will eat it and say, “It must be good for my health.” That could actually be concerning if it’s making that sort of switch?
Paul - I think royal jelly should definitely come with a health warning. I would certainly not eat it. I've seen how it can change bee development from one organism into another so I would treat it with caution.
Kat - Here at the Spring meeting, we’ve just had a series of talks about people who were looking at other unusual insect systems where there are different varieties or castes of insects. Tell me about some of those.
Paul - So there are a number of Hymenoptera that have this caste based system. One obvious one are ants. Ants work in a slightly different way in that there's no nutritional difference. It’s thought that its pheromone chemical signalling that determines choice of caste in ants. Bumblebees – it also seems to be a chemical based system rather than a nutritional based system. Finally, termites and as we saw in the talk today, nobody really knows the ecology of termite caste differentiation.
So, one thing they all have in common is that they're all eusocial. So eusocial insects have this caste determining system because of kin selection. They're all genetic. They're related and so, their hive as a whole looks after the one reproductive queen for the sake of the whole colony.
Kat - So it’s all these organisms – they're all genetically the same, but they're putting themselves into different roles and jobs to support the overall structure of that society.
Paul - Yeah, so it’s a classic case of E.O. Wilson’s kin selection theory where, because of the genetic relatedness of all the organisms within the hive, they act as one super organism for the good of the one reproductive animal within that colony.
Kat - I love JBS Haldane’s quote that he’d be prepared to lay down his life for something like two brothers or eight cousins. I guess it’s on a grand scale.
Paul - Absolutely, on a kind of 50,000 scale!
Kat - What do we still really need to know? What are the really burning questions?
Paul - I think the really burning question is still a fundamental one which is, if the genome is the book of instructions, how is it that that book of instructions can be read and interpreted in multiple different ways in response to various environmental cues? I think that’s the fundamental question in honeybees.
The other question in honeybees that’s kind of related is, what happens to honeybees when their food is contaminated with pesticides and how pesticides that have ingested by honeybees change their physiology and behaviour? I think that’s also a good question in the field.
Kat - This is such a big issue in the environmental world with the neonicotinoid pesticides and the concern that if we run out of bees, we’ll run out of food.
Paul - Absolutely. With the current decline in honeybee colonies then there will be at some stage, the inability to supply humans with food. And so, it’s a crucial question. A lot of great neonicotinoid workers has been done out of the UK from Jeri Wright in Newcastle that showed, ironically, bees are actually addicted to neonicotinoid-treated plants because nobody ever thought that a neonicotinoid which is an agonist of nicotine, may cause bees to become addicted to the pesticide.
And so, she showed in a great piece of work that bees will always feed on neonicotinoid-treated pollen versus non-neonicotinoid treated pollen because the poor bees were addicted to nicotine.
Kat - One final question, your lab keeps honeybees, you work on honeybees. Do you get honey out of them?
Paul - We do get honey out of them! East London honey is the best honey - I can recommend any East London honey you see is definitely worth buying. Everybody has flowers in their window boxes, in their nicely manicured gardens. So in actual fact, city bees are probably the happiest bees in the UK.
Kat - Paul Hurd, from Queen Mary University of London.

22:30 - Can genes save our trees?
Can genes save our trees?
with Richard Buggs, Royal Botanic Garden, Kew
From flying insects to the trees they buzz around, scientists are turning to genetics to solve one of the most pressing problems affecting UK trees today - infectious diseases, and particularly a nasty fungus known as ash dieback. Richard Buggs - lead researcher at the Royal Botanic Garden in Kew - explains the ‘root’ of the problem to Kat Arney
Richard - In 2012, ash dieback was found in woodlands in Britain for the first time and it had come here from Europe, and we’d seen it actually slowly progressing across Europe over several years. When it arrived in the UK, our government decided to fund quite a lot of research into what we could do about ash dieback.
One of the things that I've been doing is working on the ash trees themselves and sequencing their genome to try to find genes that could be responsible for their interaction with the fungus. We’re hoping that we might be able to breed trees in the future that have a resistance to the fungus.
Kat - Let’s backtrack a little bit – so what causes ash dieback and how is it spread?
Richard - Ash dieback is caused by the fungus Hymenoscyphus fraxineus which is native to Japan and Eastern China. We’re not quite sure how it got over here but here, it’s much more pathogenic against our trees than it is against ash trees in Japan and China, which is why we have a problem.
Kat - Trees don’t get up and move around. They don’t transmit pathogens in the same way that animals might. How is it spread then from tree to tree?
Richard - It’s spread by its spores which infects as they're growing and then when the leaves drop off the trees on the forest floor, you get little mushrooms coming out of the decomposing leaves, and those spread the spores that then go often and infect other trees.
Kat - So how are you trying to map the genetics of the ash trees to try and figure out how they are interacting with this fungus?
Richard - It’s relatively easy now to sequence a new genome. Actually, it took us a couple of years. We’ve published that very recently. We’re taking two approaches to identify genes for low susceptibility to ash dieback in ash trees. One is to look at other ash species from around the world, particularly one that’s from China and Japan which seem to have natural resistance to ash dieback, and we’re using a phylogenetic approach with them.
So, we’ve sequenced the genome of every tree – every ash species around the world that we can get hold of and we’re drawing a phylogenetic tree for every gene in the genome and looking for those genes which have a gene tree that fits with the pattern of susceptibility or zero susceptibility in the whole genus, the whole ash genus among all of these species to try to identify our genes for low susceptibility.
Kat - So you're trying to look for kind of suspect genes that always turn up in the populations that are resistant to the fungus and seem to be somehow missing in the populations that are susceptible to it?
Richard - Yes, variants of genes in different species of ash.
Kat - What's the other approach that you're trying to take?
Richard - The other approach is that Forest Research back in 2012 planted out loads of ash trees – actually, 144,000 ash trees - in trials across the southeast of England. Those trees are now being screened for 5 years against inoculum pressure from these spores of ash dieback. Lots of them have died but some have survived.
So that enables us to pick out British ash trees that seem to have low susceptibility. What we would like to do, if we get funding, is to do a genome wide association study on them to try and find the genes within British ash populations for low susceptibility to ash dieback.
Kat - You're starting to find these British ash trees that are a bit more hardy, can withstand the infection. Can't you just start breeding them and breed them all over the place?
Richard - That is exactly what we’d like to do. But we can accelerate that process if we’ve identified the particular genes involved and then we can do marker-assisted breeding. Another approach is genomic prediction where we don’t have to find the actual genes but we can look at the genetic profile of trees which have higher resistance, and that can accelerate breeding. So of course, breeding trees is a very long process because they live such a long time.
Kat - What about some of the new techniques that I started to see like CRISPR – these genome editing techniques? If you find the gene variations that make a tree more resistant, can't you just cut and paste those in?
Richard - Yeah, that could be a very rapid way to develop trees with resistance to ash dieback. We actually thought about doing some kind of genetic manipulation on ash trees and we surveyed the public. We found that most members of the public didn’t really want genetically modified ash trees in natural woodlands. They might be happy with them in plantations but not in woodlands. So, even if we do take a GM approach, we also need to have a more natural approach just using traditional breeding methods, perhaps accelerated using genetic knowledge but not involving genetic manipulation.
Kat - Will it be fast enough? If this fungus is spreading, if British ash trees are dying, are we going to head for a fallow period where there are no ash trees where we’re trying to breed enough resistant ones to fill the gaps?
Richard - We are likely to see a very significant reduction in our ash populations but given that there are British ash trees that do seem to be able to survive, I think in the long run, we will still have ash in our landscape. One thing we have to be particularly careful about is it’s very tempting for people to chop down ash trees when they don’t need to be chopped down.
So for example, if there's an ash tree that’s dying next to a road, the local council will need to cut down that tree because it could be a risk to life. But when they do that, when they’ve got the road closed, they’ll be very tempted to chop down all of the ash trees on that stretch of road, because they’ll think they’re saving themselves money in the long term. But in fact, they could be taking out some of the trees which have resistance which would actually survive and continue to enhance the environment in their lifetime but also have offspring which maybe have even more resistance.
So, one thing we’re thinking of trying to do is develop a tool so that we can use genetics to predict which trees shouldn’t be cut down so that before a local council, or the highways agency, or network rail start doing very expensive felling, we could actually tell them which trees they don’t need to fell, and they could stay there enhancing the environment and surviving to pass their genes on to the next generation.
Kat - Tie a little yellow ribbon around it or something?
Richard - Yeah, that’s right!
Kat - Richard Buggs, from the Royal Botanic Garden in Kew.

29:32 - Gene of the Month - Methuselah
Gene of the Month - Methuselah
It's time for our gene of the month, and this time it’s Methuselah.
Named after the biblical character who was claimed to have racked up an astonishing 969 years of life and is also named as Noah’s grandfather, the functions of the fruit fly gene Methuselah are rather more based in scientific evidence than mythical hearsay.
The gene itself encodes a protein that helps to send signals between cells and is part of a family of similar genes that were thought to be only found in insects, although similar signalling molecules are found across the tree of life.
Flies with a shortened version of the gene live around a third longer than their normal counterparts - although not anywhere close to the roughly 12 times lifespan attributed to their human namesake.
These flies also have additional superpowers - they can flap their wings faster, and are thought to have enhanced sensory abilities and are resistant to various forms of stress, including starvation, high temperature and the insecticide paraquat (the bible does not reveal whether Noah’s grandad had any of these characteristics too).
Somewhat ironically, there is a debate over the actual age of the Methuselah gene itself. Some researchers claim it is a relatively new gene, only popping up in the past 10 million years (that’s quick in evolutionary terms).
But, more detailed genetic analysis has revealed a relative of Methuselah in crustaceans, which separated from insects more than 400 million years ago - so the gene itself seems to be much older than previously thought.










Comments
Can't wait to get into this!
Can't wait to get into this!
Add a comment