Environmental DNA: Seeing the Unseen
This week, we’ll be delving into the world of environmental DNA: what did researchers find when they went looking for the Loch Ness monster? Plus, in the news, with an autumn “booster” on the cards, we look at how much antibody you need to be protected against Covid 19, and how sea level change can affect volcanic eruptions...
In this episode
00:60 - How many COVID antibodies are enough?
How many COVID antibodies are enough?
Robert Seder, National Institute of Allergy and Infectious Diseases
Recently, the Joint Committee on Vaccination and Immunisation - outlined plans for an autumn booster programme to "top up" the immunity of everyone over 50 against Covid-19. But one thing that hasn’t been clear so far is how much antibody do I need in order to be protected; and why are some people, despite vaccination, still able to catch coronavirus infection but not become unwell? Now we have a bit more clarity, because as Robert Seder explained to Chris Smith, he’s done some careful experiments on monkeys, which develop Covid-19 infections in a similar way to humans. This has revealed the level of antibody that’s needed to protect both against severe lung disease and also against just catching - and potentially passing on - the infection. The two turn out to be different, and this means we now have a benchmark to aim for, when/if embark on a programme of booster doses this autumn…
Robert - When you give the vaccine, we're trying to measure the type of immune response in the blood that would tell you that you would be protected in the lungs or in the nose. You want to protect people first in the lungs, so they don't get severe disease. And you'd like to protect people in the upper airway, so it might prevent them from getting symptoms of a cold, and then you would not be able to give it to somebody else.
Chris - Was this not known already, given that we have put billions of doses of vaccines into the world's population so far?
Robert - There had been a couple of studies that showed that the higher levels of antibodies you had in the blood, a measure of the immune response, the better off you were for protection. So our study in animals provided greater specificity to really define exactly what the level of antibody response in the blood was to mediate protection in either the lung or the nose.
Chris - So this is sort of a way of standardising. When we give someone a vaccine of some type in particular, what's the threshold level of antibody we probably need in the blood either to protect their lungs or to protect their upper airways and therefore protect them either from just getting COVID or from transmitting the infection?
Robert - That's exactly right. And the reason why, if you have a precise measurement, is it would allow you, if people had their blood measured and you were immunocompromised, or you were older or had a pre-existing illness, if your antibodies were below that level, it might indicate that you're at greater risk and it might indicate that you would need another shot of the vaccine to put you above that threshold. So it can be very helpful in guiding perhaps people at greater risk or over a long period of time, whether your immune response is waning and you need a boost. It also helps you with other vaccine approaches where if they haven't gone through the full process of a large, phase three study, but their vaccine shows very high levels of those immune responses, you might be able to use the metric data to suggest that that vaccine is very likely to work and possibly being able to facilitate approval of another type of vaccine based on achieving that certain metric.
Chris - Does it shed any light on something we're increasingly seeing perhaps in the light of having a very high fraction of the UK population now vaccinated is that we seem to be seeing lots more asymptomatic infections and people are a bit surprised because they think, well, I've been vaccinated. How am I catching this thing at all? Do your results give us some insights into how that's happening.
Robert - Yeah, so I think one of the key findings of the study was that we showed that less antibodies or less immune response is required to protect you from severe disease in the lower airway than what is required in the upper airway. And I think that explains why all of the vaccines are very good at preventing severe disease. But you're now starting to see a differentiation of vaccines for protection against mild infection or asymptomatic infection, in the upper airway. And I think it would just make sense that you just need more of an immune response in the upper airway to limit infections, especially with something like the Delta variant, where there's a lot more virus in the nose. And so the better vaccines that induce higher immune responses, that translates into having higher responses in the upper airway, and so that's why a vaccine could protect you from going to the hospital, but might not protect you from mild infection or transmitting it.
Chris - Is one interpretation of your findings then that actually, if we drove everyone's immune response as hard as possible, we could get everyone to a point where they'd have enough antibody where they just couldn't be infected, and at the moment, we're just not doing that enough?
Robert - That's a very good point. And that is potentially the rationale for giving people another vaccination or a boost, is if you could boost antibodies higher, it might sustain your continued very high level protection against severe disease. Remember all vaccines are basically 90 to 100% protective. And so if you boost, you sustain that, but if you had more antibodies perhaps in your upper airway, might that serve to limit transmission? And when you look at the Delta variant, which is existing at probably a hundred- to a thousand-fold more virus in the nose, might that be the only way that you can accomplish that is with another boost to get higher levels of antibodies? The alternative in the future might be for things like COVID and upper respiratory infections is to have vaccines that might be given to induce immune responses directly in your upper airways, and might that be better? And I think that that will be an interesting area of research going forward.
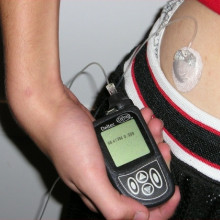
07:33 - Automatic "artificial pancreas" for diabetics
Automatic "artificial pancreas" for diabetics
Charlotte Boughton, University of Cambridge
2021 marks 100 years since the discovery of insulin. This week, there’s exciting news for the millions of people living in the UK with Type 2 Diabetes who already are, or could become, reliant on injected insulin to control their blood sugar levels - a fully automated pump has been trialed in outpatients for the first time to great success. Charlotte Boughton, a leader on the study, from the University of Cambridge is joining us now...
Adam - So Charlotte, why was this study done? Why is, you know, just manually injecting insulin, not enough. Why do we need the pump?
Charlotte - We've previously shown that a closed-loop system or an automated system can actually improve people's glucose control in those with Type Two diabetes when they are admitted to the hospital. And so in this study, we wanted to see if the same system can be safe and effective compared to people's usual insulin injections in the home setting, as you mentioned. And we did this study in people with Type Two diabetes who also have kidney failure and need dialysis, and that's because that's a time when diabetes can be particularly challenging to manage
Adam - What are people wearing? What does it look like when they put it on? And how does it work?
Charlotte - A closed-loop system comprises three separate parts or bits of kit. So people wear a glucose sensor, which is about the size of a two pound coin, and that's worn on the arm or stomach, and that measures the person's glucose levels in real-time. And that then communicates with an algorithm or a controller, which in our system is on an app on a normal Android smartphone. And the algorithm calculates the right amount of insulin and tells a small insulin pump, which is a light, slightly smaller than a mobile phone-sized pump that delivers the insulin to the person. And it repeats this cycle about every 10 minutes, constantly adjusting the insulin dose in response to the glucose levels. And the person doesn't need to do anything, they just need to be wearing the kit.
Adam - And how easily do people tolerate this? Because I imagine at least part of it is going to have to be, you know, at least under the skin
Charlotte - Young children who have Type One diabetes wear very similar technology to this, so it's been designed in a way that it's highly acceptable. So the sensor is inserted every 10 days and it's just a tiny little wire that sits under the skin. It doesn't hurt at all. And then that sticks on with a bit of adhesive. And similar to the pump people are used to doing insulin injections, where with a pump, you just need to change the needle every three days, rather than doing injections several times a day. So people really like wearing the device. It takes some getting used to but overall is pretty well tolerated.
Adam - And is that then the big advantage that it takes the thinking, I suppose, out of your head? You can't forget. You can't make a mistake.
Charlotte - Yeah, absolutely. So in this system, it's a fully automated system. For people with Type One diabetes, they use a slightly different system. But in this case, you can almost forget about your diabetes because you don't really need to do anything. You don't need to measure your glucose levels because that's done automatically and you don't need to calculate the right dose of insulin to give, and provided that you change over the devices every sort of few days, then actually the system will work and allow you to get on with your day to day activities.
Adam - Now, to me, that sounds brilliant. The thing I can forget about until something, say, goes wrong. So what if my phone dies or I were to forget it and I needed my insulin. What would happen then?
Charlotte - We rarely leave our phones places, but absolutely it's something that we have to think about when designing these systems. So if the phone's out of Bluetooth communication or the battery runs out, then the pump still gives insulin. It's just it won't give it in an automatic way. So you lose the brains behind the system, but you can still measure the glucose levels and the insulin is still given. So it's sort of a safety net really.
Adam - And it will be effective enough until they can get their phone, or recharge?
Charlotte - Exactly, until it's recharged, yeah.
Adam - That's why it's potentially great for the patient, but what about on the other side, the hospital system, the NHS? Are there benefits for them, for people using this system?
Charlotte - So our study that we've just done was quite small and too short to demonstrate any clear benefits to the NHS. So this would need a much larger and longer study to see if that can translate into improved long-term outcomes, which may have cost savings for the NHS in this population.
Adam - Okay, that makes sense. You mentioned earlier that there are previously pumps for Type One diabetes, and this has been developed for Type Two. Why is Type Two coming later? What's different about it that makes it harder to get a grip on?
Charlotte - It was more where the priorities lay. Some of this research started off in children because of the concern about the effect of high and low glucose levels in young children with Type One diabetes, where they can't often communicate that they've got high or low glucose levels. And these closed-loop systems are transforming the management of Type One diabetes. I think it's come later in Type Two diabetes because in Type One, people don't have any insulin, and so rely on insulin completely to manage that glucose. Whereas in Type Two diabetes, we do have other treatments, so diet and exercise and oral medications and other injections as well. So there are more options available.
Adam - What state are you in? Could you see this being rolled out more widely?
Charlotte - So we already do have similar systems being rolled out in Type One diabetes. And our study was a sort of early proof of concept study that this is potentially a treatment that could be rolled out more widely, particularly in a follow-up study that we're doing in people with Type Two diabetes who don't have kidney failure. And that will hopefully provide evidence to support wider adoption of this technology.

13:52 - Moon magnetism mystery solved
Moon magnetism mystery solved
John Tarduno, University of Rochester
Walk almost anywhere on Earth armed with a compass, and the magnetic field issuing from inside the planet will point you in the direction of the North Pole. But what would happen if you went for a similar walk on the Moon? Does it have a magnetic field? Some claimed that - at least at one point - it did, others were doubtful. Now Rochester University’s John Tarduno has analysed samples of Moon rock brought back by the Apollo missions and confirmed that the Moon never did have a magnetic field. Instead, he’s shown, rocks around craters can become magnetised by meteor impacts, and it’s this that tripped up earlier researchers, as he told Chris Smith...
John - Since the return of the rocks from the Apollo missions there has been this idea that the moon once had a magnetic field, just like the earth, but it lost its magnetic field, but it only lost the magnetic field relatively recently. That's a paradox because the moon doesn't really have a power source to drive a magnetic field. The Earth's magnetic field is created in the earth's core, the moon has a core, but the core is really small. So how could the moon's core have created a magnetic field? That was a question that we're really after.
Chris - What did people think it did have one, then?
John - The Apollo astronauts, part of the missions of course were to return lunar rocks. And when these rocks were analysed in the 1970s, it was found that some of them had strong magnetisations. And on the basis of that scientists concluded that the moon once had a magnetic field as strong as the earth's.
Chris - So what would you speculate caused those magnetisations in the bits that the Apollo astronauts brought back then?
John - What we found is that a very young sample - young on the moon means 2 million years old - collected by the Apollo astronauts from an impact crater had a really strong magnetisation. So actually impacts themselves can create magnetic fields, and if there are rocks nearby, they can be magnetised.
Chris - You've mentioned that you did this by taking samples that had been brought back to earth by various Apollo missions and so on. How did you analyse the samples you got?
John - We have a magnetics laboratory and we have a special magnetometer - a device to measure magnetic fields - but we can measure the trace magnetic fields, and we also use a special laser to heat the samples. What we're trying to do is we're trying to reproduce the process where, during its formation, the rock actually acquired magnetisation. And by reproducing that process in the laboratory, we can then gauge the nature of the magnitisation contained by the samples. Using these techniques we were able to demonstrate that most of the samples that we measured, the ones that were not associated with impacts, had no magnetisation, but most importantly, they have the ability to acquire a magnetisation if the magnetisation had been in place. So essentially what we're doing is we're evaluating the reliability of these ancient rocks as tape recorders. So we were able to demonstrate that they were actually are good tape recorders and they're recording no magnetic field.
Chris - And what do you think the implications are then? How does this rewrite the story we have assembled so far about where the moon came from?
John - Well, first of all, it resets the idea of the interior evolution of the moon. This has been a paradox for decades and that problem goes away now. But I think the bigger importance here really is the history of the surface of the moon. Without this magnetic shield, this means that solar wind elements, things like helium three, which is really interesting and could be a potential power source for exploration, and hydrogen - hydrogen can actually combine with other elements in the lunar soil to form water. So we could have helium three and water, much more of it than we might've thought in the past, in the lunar soils useful for future exploration.

17:58 - Sea level influences volcanic eruptions
Sea level influences volcanic eruptions
Chris Satow, Oxford Brookes University; David Pyle, University of Oxford
We normally think of volcanoes as unchangeable - we couldn't stop one erupting if we tried, and we have tried! - but it turns out that some environmental factors like high sea level can prevent volcanoes from erupting. Eva Higginbotham reports...
Eva - If you live in the UK, you may not think that often about the risks of volcanic eruptions, but the 800 million people around the globe who live within a hundred kilometres of an active volcano surely do. Others like geologist Chris Satow from Oxford Brookes University just really likes studying them.
Chris - The question we were trying to answer was "What effects does sea level change have on volcanic activity, in particular at volcanic ocean islands?" And we were able to discover that low sea levels are the times when the majority of eruptions are concentrated and at high sea levels, the high sea level itself has the effect of reducing the likelihood of eruptions. And we used the volcanic island of Santorini as our case study.
Eva - Scientists had previously seen that in places where there had been large ice sheets in the past covering volcanic areas, like Iceland for example, there had been a big increase in volcanic eruptions when that ice had been removed.
Chris - So it got us thinking that perhaps the same effect might occur in the oceans, but of course in the oceans, it's not ice that's being removed and added. It's water that's being removed and added as the sea level has risen and fallen over many tens and hundreds of thousands of years. And the sea level goes up and down globally by quite a substantial amount. So the difference between the sea level today and the sea level during the last ice age is around a hundred meters. That's a huge change in the sea level. That's essentially saying that the English Channel would be entirely dry and you could walk from England to France at that time. That's a huge amount of mass to remove from the surface of the Earth or the top of a volcano. And that gave us an idea that the sea level change might have some influence on the activity of the volcano.
Eva - But how do they know when eruptions happened anyway? David Pyle is another volcano enthusiastic this time from the University of Oxford.
David - So Santorini is a fantastic example of a volcano that shows what we'd call a layer cake. You can see different layers of rock making up the cliffs, and some of those layers of rock, they weather in different ways, they crumbled in different ways. They've got different colours and different patterns on the surfaces. And as a geologist, what we do is essentially work our way down through those layers.
Eva - By analysing these layers of pumice and ash and solid grey lava, geologists can come up with a relative time or stratigraphy of when eruptions took place, which is what they did for Santorini. The next step was comparing that timeline with the sea level record.
Chris - By taking samples of sediments from the bottom of the oceans, and these come in huge tubes - cores as we call them - which might be even up to a kilometre in length. And we know that the mud that's higher up in that core is very young. And the further down you go, the further back you go in time. And if you can process them, if you can split it up into its constituent parts, you start to find fossils and most useful fossils for us to work out what the sea level was, are fossils of these creatures called 'foraminifera', or 'forams' if you're on first name terms with them.
Eva - Forams are these microscopic sea creatures who live almost everywhere on Earth, and they produce these shelves using the materials from the seawater they live in, including oxygen. Like all elements, oxygen comes in different varieties, called isotopes. And luckily for scientists, the ratio of various oxygen isotopes in the foram shells reflect what the sea level was when they were alive.
Chris - So these little creatures are recording what the sea level has been at various points in the past into the composition of their shells. And then we can extract them from their fossilised mud, analyse them and look back through time at how the sea level has changed over hundreds of thousands of years.
Eva - Chris, David and their colleagues then used computer modeling to marry up the data on Santorini's eruption history with the established sea level record and found that, indeed, during times of low sea level, there were more eruptions and times of high sea level, there were fewer. So does this mean, contrary to what we normally think, that having high sea levels like we do now is kind of a good thing?
Chris - The truth is we don't quite know what's going to happen to all the magma that's being stored underneath the volcano, if it's not being erupted. So it could be of course, that the magma gets stored and then more magma gets added and more gets added and then over time. So it could be that in fact, you build up a much larger volume of magma underneath the volcano, which then in the future could results in a much larger eruption than would otherwise have been anticipated
Eva - Interestingly though because the sea level rises all over the world at the same time, does this suggest that in some ways, all the underwater volcanoes around the world are sort of linked?
David - If you allow the system to persist for long enough, then indeed you'd expect that volcanoes would come into sync or there'd be some sort of synchronisation or synchronicity between volcanic systems that are physically independent, but they're all kind of responding to the same external processes.
Eva - So just as everyone around the planet is likely to be affected by the changing climate due to global warming, it seems volcanoes may be going to experience a similar phenomenon.

23:46 - Babylonian artefact reveals history of maths
Babylonian artefact reveals history of maths
Daniel Mansfield, The University of New South Wales Sydney
But first, we probably all remember learning Pythagoras’s theorem in school, and there was always one kid who asked, “but when are we ever going to use this in real life”? Well it turns out that not only did ancient Babylonians learn about Pythagorean triangles 1000 years before Pythagoras was born, they also used them in real life to measure out land. An old clay tablet inscribed with symbols found lying in a museum was the clue that led researchers to this ancient form of maths, as Sally has been hearing...
Daniel - We always knew the ancient Babylonians were aware of Pythagorean triples, but we had no idea why. We had no idea what they were doing with these objects.
Sally - That’s Daniel Mansfield from the University of New South Wales, Sydney, who discovered that the ancient Babylonians were using Pythagorean triples. So, what is a Pythagorean triple? We all remember right angled triangles from school and calculating the length of the longest side, the hypotenuse. Because of all the square roots involved, even if two of the sides are nice whole numbers, like 5 and 6cm long, the third side is usually a horribly complicated number with lots of decimal places, which in our case as you will have all worked out already from remembering Pythagoras’s theorem, is of course the square root of 25 plus 36, otherwise known as root 61, or the rather unpleasant 7.8102496... However, for a special group of right angle triangles, all the sides are lovely round numbers. For example, 3, 4, and 5. And if you make a triangle with sides that measure 3cm, 4cm and 5cm, it’s impossible for it to be anything other than a perfect right angle. These special triangles are called Pythagorean triples. . Mathematician Daniel was looking at markings written on palm sized flat discs of clay from 4000 years ago…
Daniel - You know, this is about a thousand years before Pythagoras was even born.
Sally - Two tablets in particular caught his eye in the museum. One, the Plimpton 322 had already been deciphered in 1945 which was simply a long list of these Pythagorean triples, much like the trigonometric tables you may have looked up in school. But the ancient Babylonians probably weren’t studying for their Maths GCSEs, so why did they need these triples? That’s when a small, round clay tablet drew Daniel’s attention, with rectangles and measurements that he realised was a map of field boundaries.
Daniel - Now, thanks to this new tablet, we know they were using them to solve problems about land. In particular, they were using them to create perpendicular boundaries.
Sally - Why did Babylonians need to measure out their land so accurately with such precise right angles?
Daniel - Let's slow it down. Because I want to tell you a story about lands and gods and math. So right at the beginning, Babylonian society is based on these small cities built around the local temple and all the land around is owned by the local temple or maybe a king or a palace. And they have surveyors, but the surveyors, unlike our surveyors, they don't measure boundaries. They're just there to tell the local prince or the local temple administrators how much land they've got. They can decide for themselves what they own and they practically own everything. They don't need no surveyor to tell them where their boundaries are. Then at about this time is old Babylonian period, something changes. And we now see land in the hands of private individuals and private individuals can't decide between themselves necessarily where their boundaries are. You need a surveyor to do that for you.
Daniel - Now we see surveyors change what they do instead of just saying, "Oh, prince, your land is this large". They say "Two people, here's your land. Here's where your land stops. And here's where your land stops". And they start making boundaries. Now to maintain good boundaries between neighbours, you need to have some confidence in those boundaries and to a Babylonian that means you get the right angles just so. In modern society, that's easy. You just use trigonometry to make the right angles, but this is a thousand years before trigonometry would have been invented. So instead the Babylonians are using Pythagorean triples.
Sally - Why should we care about old maths when we've got perfectly functioning new maths?
Daniel - It's so different. Their whole approach to mathematics is just unbelievably different. We tend to think of mathematics as this kind of universal language where everyone can agree on what at least what mathematics is. And if aliens came from out of space, then we could, we may not be able to communicate them in words, but we could, you know, we could share math with them.
Sally - Didn't we send maths on that golden disc out into space.
Daniel - Yeah, I'm afraid we may, we may have overestimated how universal that was going to be.
Sally - Has anyone tried using Babylonian maths to solve modern problems?
Daniel - We're still trying to figure out what Babylonian maths is. This is all, this is all happening very quickly. Actually the original computers used a Babylonian method of performing division because it's very, very simple. I like to think there's still a place in the world for them though, for fantastically simple calculations that sure, aren't as accurate as what you can do now with a modern computer. But geez, they're efficient, just so very efficient. So I like to think there's a place in the world for those kinds of things like in computer graphics, where speed is more important than accuracy. Maybe this Babylonian approach might have some kind of advantages.

What is eDNA?
Beth Clare, York University in Toronto
Sally Le Page has spent the last couple of weeks with Naked Scientist Phil Sansom digging deep into the exciting new world of environmental DNA, or eDNA for short. It's the same technology you might have heard being used during the pandemic to detect COVID DNA in sewage water. But, what actually is this technology? Sally spoke to Beth Clare from York University in Toronto...
Beth - Environmental DNA is any DNA you collect from an animal that you didn't get directly from that animal. So, some people would argue it has to be effectively naked DNA that's out of itself, floating around in the environment. Other people would say, well, it's just sort of anything. It’s bits of sloughed off cells and bits of hair that are out there that you collect, but you don't actually see the animal when you get them. You're just taking them out of some substance, like water or soil.
Sally - But, why would you want to get the DNA from the environment, the water, the soil, and not just take the DNA directly from the animal?
Beth - Well, because some animals are hard to find. So, it's a good non-invasive way of collecting things from an animal that you can't get, or you shouldn't get in contact with. So, it's a way of collecting things that doesn't cause you to interact specifically with the individual that you are targeting.
Sally - When you say you shouldn't get in contact with the animal, what are we talking about here?
Beth - Well, something that's really sensitive, something that is highly endangered. You don't want to interfere with it in any way that might cause harm or cause stress to that animal. It's a really good non-invasive technique.

32:19 - How was eDNA discovered?
How was eDNA discovered?
Eske Willerslev, University of Cambridge
Phil Sansom and Sally Le Page heard from founder of eDNA Eske Willerslev from the University of Cambridge about his discovery...
Phil - Eske, am I right that you wrote the very first paper on environmental DNA and sort of established this whole new field of genetics back in 2003?
Eske - I believe so. I mean, it was part of my doctorate, so I couldn't get my hands on any of these super famous fossils, you know, bones and teeth and stuff like that. So, I kind of thought I have to do something else. I thought, what can I do where people don't care about the material? And then I was sitting, looking out of the window, leaves falling down from the trees. It was autumn and there was a dog that took a c*** on the street?
Sally - Or to put it a little more delicately Eske, the dog defecated.
Eske - And I thought, well, we know that there's DNA in these things right from the animal or the plant. But we also know that after the next rainfall, the dog faeces will be gone. We also know after a few years the leaf will be gone, but the question is what will happen to the DNA? I thought, well, maybe it could survive in sediments. I went to my supervisor and said, I have this idea. Could DNA survive in sediments and both him and the rest of the professors laughed and said, we've never heard anything so stupid, but luckily I didn't give up actually. So, in the past, before I became a scientist, I was a trapper in Siberia and also did a lot of expeditions there.
Sally - Because all scientists have a background as fur trappers in Siberia.
Eske - But it became actually quite helpful because obviously I'd seen these permafrosts where it's frozen soil and we knew already then that cold temperatures are good for long-term DNA preservation. So, I got in touch with a Russian guy with some of these permafrost cores. Actually from the same areas where I had been running around as a trapper. I thought one would need an enormous amount of material to obtain DNA because what is the chance of an animal passing just across that particular area? So, I tried to work with very large, like kilograms of soil, but obviously you couldn't extract. There were no DNA extraction kits fitting that because everything people were doing at that time was retrieving DNA from bacteria. So, I tried to freeze-dry it and all kinds of stuff, and it didn't really work. And in the end, you know, I said, well, I'll just try with half a gram. And I took out this DNA and I remember it was like Christmas Day. Everybody had gone home from work and there, I got the results coming out of the sequencer and I blasted them. And, you know, I could see woolly mammoth DNA, horses, reindeer.
Phil - Woolly mammoth?
Eske - Yeah, yeah. This was ancient soil. It was like 20,000 years old.
Phil - You immediately found woolly mammoth DNA, which is pretty incredible. Did you expect it to work so quickly or was it a complete shock?
Eske - I didn't expect it to work that well. I thought that you would need a larger amount of materials to even then be lucky. Finding a mammal for example, would be like looking for a needle in a haystack. So, that's what I thought, but realised just from those early experiments, that was not really the case. We are in fact walking around on DNA from animals and plants, both in our contemporary settings, but also from the past. When those sediments were part of the surface 20, 30, 40,000 years ago. But nobody believed it for 10 years. I was almost the only one doing this stuff, for the following 10 years. And I remember they came to my dad and they just didn't believe it. I mean, they said, well, you know, it's not real. And until they went to the lab themselves, got a sample and tried.

36:14 - Sampling the river for eDNA
Sampling the river for eDNA
Kat Bruce, NatureMetrics
Just 20 years ago, it would have taken months of work to get just one eDNA sample. Luckily, nowadays the process is so simple that even primary school kids are doing it. Sally Le Page met up with Kat Bruce, founder of NatureMetrics, to discover what animals had been visiting the river, Great Great Ouse, at Over in Cambridgeshire...
Kat - So we're going to collect an eDNA sample, so we are going to filter some water. And that filter will be sent back to the NatureMetrics lab and we will analyse it to see what species of animal we can find from the river.
Sally - It's so exciting. How do you collect eDNA? What are we going to be doing?
Kat - Over the years, we've designed this really simple kit. The first thing we ought to do is put gloves on. Again, this is to stop us from introducing our own DNA into the river. In reality, we actually are able to use human blockers in our analysis, and that really sort of reduces the amount of human DNA, because to be honest, this river is full of human DNA.
Sally - And out of this plastic bag with all of the different ingredients, I mean, that just looks like a sandwich bag to me. Have you just brought your packed lunch?
Kat - I have, this is a very sterile pack, so you can see it seals. So, we have to rip the top off to open it, take the top off this bag, reach down and scoop up some of the surface water. I'm going to hold it so the bags facing upstream and the water can just flow into it.
Sally - We now just basically have a very clean sandwich bag full of water.
Kat - I mean, so you can think of the water in here as a sort of soup of genetic material of all of the wildlife that's been in and around the river. So it will contain the DNA of the fish and things that are living in the river, but also a surprising amount of the land-based species that have been either drinking or swimming or whose DNA has been washed off the bank into the river as well. And then we've got a hundred mil syringe. We literally just go to use the syringe to pull up the water and then the syringe screws onto the filter housing with a filter membrane inside it, when we're dealing with eDNA, we're dealing with really tiny traces of DNA. That makes it very vulnerable to contamination either from ourselves or from sort of other things that we've brought in from the environment. So, the membrane is kept inside that housing.
Sally - It does just look like a plastic coin almost.
Kat - Yeah. It's about sort of twice the size of a 50 pence coin, I guess. So, then we just push the water through any traces of particles, sediments, DNA, cells, et cetera, that are all stuck inside the filter membrane now. So, we'll do this about 20 times. I think this is pretty much my last one and I'm just going to press the preservative liquid inside. I've got the cap on each end, he seems to have a yellowish colour on the inside.
Sally - That's the only thing that people have to post to your lab.
Kat - Exactly.
Sally - So we have just samples, potentially dozens of species just in like 10 minutes. That's just by sitting on a jetty and squirting some water through a glamorised water pistol with a filter on the end. That's it, that's it.
Kat - Yeah. Expedite that delivery back to the lab and see what we can find.
Sally - It was just unbelievably simple and because it's so simple, they can send these kits off to researchers wherever they are. They can stick them in their backpacks and they don't need a lab. And the company nature metrics is now working with the conservation organisation, the IUCN to sample water from around the globe to create eBioAtlas documenting all of the world's aquatic species.

40:32 - Looking for animal eDNA in the air
Looking for animal eDNA in the air
Beth Clare, York University in Toronto
Phil Sansom and Sally Le Page have been speaking with Beth Clare, who is looking to push the field in a whole new direction by looking for eDNA in the air...
Beth - Well, I was writing a report for the government on how we should look at environmental sampling into the future. And I was writing about all the sources of DNA, and I was summarising papers like Eske’s, and thought we should look in water, and we should look in soil, and we should look in leaf litter, and in rain, and in snow, and I said “and in the air!”. And like a good scientist, I went to find the papers to back up all these statements. And I got to the point where I couldn't find one for air, except for some science fair students. And I tried really hard to get in touch with them and I couldn't. And so I said, we should look in the air. And then I thought, well, okay, I'll look in the air.
Sally - How do you go about looking for DNA in the air?
Beth - We've done it a few different ways, but the one we actually published, you know it's the middle of a national lockdown. You can't go anywhere. You can't really do much experimenting, but we had to go into our naked mole rat room to take care of the animals, and they'd been there in their colonies
Sally - In the lab, or just in your house, you have a naked mole rat room?
Beth - No. In the lab, in the university lab. So, we've got the animal care facility and we've got these naked mole rats. They live in a colony. They're really confined. They've been there for many generations. They live in these burrow systems and we thought, okay, if we can get DNA out of the air, that will be the place you can get at first because they have been there for a long time and the DNA is building up in the environment. And a lot of people said, well, that's not going to work because it's too diluted in an air volume. And we sucked air through filters for periods of time from inside the naked mole rat colonies, and then out in the room where those colonies existed, but they weren't actually open. And to our real surprise, it worked on the first try. So, the first filter we sucked air through produced naked mole rat DNA. It also produced human DNA. It produced dog DNA.
Sally - I'm guessing you don't have dogs in your lab?
Beth - No, but one of the animal facility caretakers looks after his mother's dog. And so he was tracking, we think, his mum's dog's DNA into the room with the naked mole rats, and we were sampling it in these forensically trace amounts.
Sally - That is incredible.
Phil - So, it's almost frightening, Beth, how much you can find just from the air, the amount of information you can glean!
Beth - I think so, but we should think of this as a massive opportunity. That idea that you can actually go and survey that way opens up a whole new world of how we might track biodiversity, and the aquatic eDNA people have led the way showing that you can go into oceans and pick up DNA. You know the volume is less of a problem than we thought. And so what we're trying to do now is show that we can replicate that on land and that we don't have to worry too much about terrestrial mammals washing into the water in order to get them, just go after the air they live in and see if you can find them there.
Phil - And would it work outside?
Beth - Out in the wild, that's the next big challenge and something that we're working on right now. It's a big challenge, but it's not an insurmountable one. And, I think we can do it.

43:48 - What animals did we find in the river?
What animals did we find in the river?
Kat Bruce, NatureMetrics
Sally Le Page was very keen to find out what animals in the river Great Ouse were revealed by our DNA test with Kat Bruce from NatureMetrics...And Naked Scientist Phil Sansom, Eske Willerslev from the University of Cambridge, and Beth Clare from York University of Toronto weighed in...
Sally - We've got our sample, we popped it in the envelope. We mailed it to your lab. What happens in the lab?
Kat - It goes in the oven which heats it up to about 56 degrees for a few hours. And that just bursts open any cells that are still intact and releases the DNA into solution. Then we need to make millions of copies of the DNA of the particular group of animals or organisms that we're wanting to survey. And that gives us enough to be able to sequence.
Sally - So, you're only making the copies of the animals you care about?
Kat - Yes, exactly. Otherwise, if we just sequenced the DNA as it is in the sample, it would really just be dominated by bacteria and things. So, once we've done that, it goes onto the DNA sequencer. At the end you have a data file which has 30 million lines of As, Ts, Cs, and Gs. And of course that doesn't mean very much to most people. So, it then goes through a computational process, which sort of summaries that information. And then it runs each unique sequence against a reference library of known DNA sequences of species. And that lets us put the names onto the species that we've found.
Sally - And the big question is what did we find in the Great Ouse?
Kat - So, it takes a few weeks to analyse a sample from beginning to end for meta barcoding. So, in this case I've got some results from last summer when I actually came down here with a group of citizen scientists.
Sally - In true Blue Peter style here's some we made earlier!
Kat - Absolutely. And in total, so we did a vertebrate analysis on the samples and in total, we've got 23 species. Most of these are fish. So, 14 of the 23 species are fish, things like roach, perch... quite a bit of human in there as well. It was a busy day on the paddle boards! But then, you know, even things that are much lower level things like spined loach, this is quite an important species in this area, actually it's really hard to survey because it just sits around at the bottom of the river and is not sampled with electrofishing.
Sally - That's incredible. And looking down the list, we got a whole bunch of fish. You've got your dace, roach, chub, but we've also got super exciting things like water voles. They are incredibly rare now in the UK and very hard to spot.
Kat - Yeah, absolutely. They are a protected species. A lot of effort goes into trying to survey and confirm the presence of water vole.
Sally - Just looking around, we're not even in a particularly quiet part of the river, there's boats going past, we've had lots of dog walkers. I can see the reeds, but I never guess that we've got water voles that we've got all of these fish species. We can't see down to the bottom of the river and they're just there and you can tell with their DNA.
Kat - Yeah. And what we really want is people to go, oh my gosh, now we know that these species are here, we're going to come in and look for them and get interested in them and then come to these places with more of a sense of what the place is and what you're sharing, what other species you're sharing the places with.
Phil - Sally, I've never even seen a water vole.
Sally - I've only seen one. And I was incredibly excited when I first saw them. They are, of course, Ratty from Wind in the Willows and their population has dropped. They're so hard to see and they were in the water.
Phil - Is that in line with your predictions, you think?
Eske - I think it's pretty well what you would expect from this.
Beth - Nobody is ever going to look at water or air or soil again once we start talking about what's in it, because these things are just soups. It's just everything that's been through leaves behind a trace of itself. And what eDNA is doing is letting us look at those traces, unlocking that little mysterious door. And as you said, see what's invisible.

47:41 - Making whole genomes from eDNA
Making whole genomes from eDNA
Eske Willerslev, University of Cambridge; Beth Clare, York University, Toronto
Phil Sansom and Sally Le Page spoke with Eske Willerslev from the University of Cambridge and Beth Clare from the York University, Toronto, about combining these tiny fragments of eDNA into entire genomes...
Phil - Eske, part of what you've been doing is moving from fragments of DNA from each animal to sequencing entire genomes, all the DNA from one creature. How do you get an entire genome, a lot of DNA, from these samples?
Eske - For many years, it was what is known as metabarcoding, l ike what you just experienced here, where you are pulling out specific regions of the genome that you know are quite informative in terms of doing some species identification, but obviously just look at the human genome, right? Most of the information that is being used to understand, I mean, not only what species it is, but also this particular individual, how is that related to other individuals? And also we can start looking at selection, for example, what genetic regions are important for survival. But for all these things you need other parts of the DNA. So what we started doing was to go from the so-called metabarcoding to simply shotgun sequencing. That's where you just sequence randomly all the pieces of DNA in your environmental samples. I mean, the challenge is you need a proper reference for comparison, right? And in some situations they are not present and you have to go out and create those references yourself. But what we did this year was that we managed through these many pieces of DNA to assemble it together into genome-scale sequences. And we did that for two bear species that lived 12,000 years ago.
Sally - That DNA must be very old and broken apart. The pieces must be tiny. This is like the hardest jigsaw puzzle in the world. You've got these tiny, tiny pieces and you're sticking them together and you don't even know what the overall picture looks like. How'd you do it?
Beth - Metaphorically using the puzzle metaphor, you've got a picture of what you think it should look like. And you've got all these pieces and you assemble it on the picture to try and fill in the gaps to see what's there. And it's enough to do things like population genetics to figure out the diversity of a population in an area, that's a relatively easy challenge for eDNA. So, you may not get the complete picture, but you get enough of it to be really scientifically useful for understanding how fragile a population is or how important a landscape is, in terms of what is living on it. And that's where the real application of eDNA comes in. It's monitoring populations effectively in real time when you can't and don't have the manpower to go out and sample them by looking for the individuals. You just sample the environment they're in to learn a whole lot about who they are, where they came from, how many there are, what their diversity is, what their origins are. I think we can do that with modern day live animals that are still breathing in their environment, the same way that Eske can with historical records.
Sally - How would this tie in with DNA from the air? Do you think being able to get whole genomes from the air; is that going to be possible?
Beth - Oh, I think so. There's different challenges with air. So, things like UV light are probably going to be damaging DNA in a way in air that you don't have in soil, but it's not insurmountable.
50:59 - What is the future of eDNA?
What is the future of eDNA?
Eske Willerslev, University of Cambridge; Beth Clare, York University, Toronto
Phil Sansom and Sally Le Page spoke with Eske Willerslev from University of Cambridge and Beth Clare from York University, Toronto about the future of eDNA...
Phil - Can you give me an idea of the limits of this?
Eske - I think the biggest limitation right now is, you can say, reference material or reference sequences because if you take an insect out, right, you can identify, in many cases you can say this is this species of insect. With environmental DNA, whether you are doing metabarcoding, or shotgun, you're relying on databases, you're relying on somebody else having sequenced that species before. And if they haven't done that, you're kind of in a situation where you can say, well, this is an insect, but I don't know exactly what insect I'm dealing with. Of course these databases are exponentially growing. So people are sequencing more and more and more. So, my prediction would be within that 10-year period, we will really have that reference. But at the moment it is a limitation.
Sally - And if we can get to the point where we've got a reference genome for every species on the planet and we can get genomes from animals without even seeing them, that's just going to revolutionise everything!
Beth - It already is. You've got entire groups of people now using eDNA in the water. Everybody from research scientists to conservation management organisations that are using that substance to help them track and monitor populations for different applications.
Sally - Just like what Kat's doing with the aquatic surveys around the world!
Beth - Exactly. It's become a commercial industry. It's a research industry. It's a management and regulatory industry. All of these different sectors have taken up eDNA as a way of doing their jobs more efficiently and understanding all of those factors, which make that possible is what we really need to do to push that field forward. And if we can push it into other realms, terrestrial life, through air and soil, the most unusual sources like honey, tracking eDNA and different things like that, that have been used allows you to really expand this field in a way that I don't think anybody could've predicted 10 years ago. So, Eske’s ten-year prediction for genomes from substances: I'll give him five.
Sally - Wow, five years. That will be very exciting.
Eske - It's really just the imagination that are the limits of how you can use it.

53:27 - Can eDNA identify the Loch Ness Monster?
Can eDNA identify the Loch Ness Monster?
Neil Gemmell, University of Otago
Neil Gemmell from the University of Otago went hunting for the Loch Ness Monster using eDNA tech, as he told Sally Le Page and Phil Sansom, and Beth Clare from York University and Eske Willerslev from the University of Cambridge weighed in...
Sally - Talking about imagination. We promised you that we would unveil the mystery of the potentially imaginary animal Nessie, the Loch Ness monster.
Phil - Don't say that, no, you're going to break my dreams in half.
Sally - But we don't know Phil that's the thing, but maybe with eDNA we can find out if Nessie is imaginary or not. Geneticist Professor Neil Gemmell took water samples from all over Loch Ness in Scotland, to look for the Loch Ness Monster. Phil, let's put you out of your misery and work out one way or the other who or what, or even if, is Nessie.
Neil - We went to Loch Ness to prove the power of environmental DNA for studying the biodiversity of our natural world. And the sort of big hook was that, of course, if Loch Ness has a monster, environmental DNA may give us some insights into what might explain the monster and the myth that has been so pervasive in our recent history. But, really it was just a great way to showcase the power of environmental DNA and how we could better understand the biodiversity of places that are special like Loch Ness. And, you know, if you like, we might've found a monster. And we didn't find it. We found all the fish that we expected to find in Loch Ness. Found a whole bunch of other species that we knew were in the area, their total species count. And it's hard to know if these are species, but I guess the distinct DNA sequences that we could identify, there's about 3000 of those. So, if you like then, about 3000 species, and most of those are tiny little things like nematodes. So, roundworms, rotifers, phytoplankton, small crustaceans, those sorts of things. And then of course the bigger things like the fish, salmon, pike, eels, sticklebacks, those sorts of things that we knew were present in Loch Ness. We found all those. We went looking to test a variety of ideas that people had considered. So, the plesiosaur was one, some giant catfish and other giant fish species, sturgeons. And one that we actually can't put to bed at the moment, was the idea that there might be a very large eel or some eel-like creature present because we do find eel sequences in Loch Ness, unsurprisingly. But, whether they are a giant eel or just an ordinary European eel I couldn't tell you. You know, people say to me, does that mean the Loch Ness monster idea's dead. And the short answer is no, people are going to believe what they want to believe. And we joined a very long list of projects that have gone to Loch Ness with the intention of showcasing the latest technologies to see if they could find the monster and found actually no evidence of it, but that doesn't stop people going there.
Phil - It could be a giant eel.
Eske - It could be a giant eel, who knows.
Sally - Phil there's still some hope there.
Phil - The hope's alive and I'll take it.
Beth - But, he was also right when he said it's, those kinds of stories are a really good way for us to showcase how these technologies can be used. And so what he did was real science, just because the motivation is something that's fun and gets people's attention. He's able to demonstrate the value of what he's doing.

56:51 - QotW - Why do ladybirds differ in their spots?
QotW - Why do ladybirds differ in their spots?
This week, Eva Higginbotham's been going dotty trying to answer this question from listener Ruomei...
Ruomei - Why do ladybugs have a different number of spots on their backs?
Eva - Now, I have always wanted to know the answer to this back in the playground everyone said it was about which branch of a tree the ladybird was born on, which I must admit, I totally believed for a really long time. But to get the real answer to this question, I enlisted the help of ladybird enthusiastic Helen Roy from the UK Centre for Ecology and Hydrology.
Helen - Some insects are spotty, some striped and some simply quite blotchy or not patterned at all. And it actually can take a day or two for the colours of an adult ladybird to appear after it first emerges from the cozy confines of its pupa. It's really fascinating to watch this happen. And I can remember as a six year old, seeing some bright yellow and translucent ladybirds on the vegetable patch and within hours, the red colour and seven black spots began to appear. These were seven spot ladybirds, and they would have exactly seven spots throughout their adult life. Ladybirds do not get more spots as they age, but interestingly, the spot number in some species can vary a lot, so the two spot ladybird can have two or four or six spots.
Eva - Largely it comes down to what species of ladybird you're actually looking at and kind of like our eye colour, whatever you have once the pigment has come in is what you are set with for life. But why do they have these colourful patterns in the first place?
Helen - The bright colours and contrasting patterns of some ladybirds are called warning colours. The lady bird is giving a clear message to other species that might be tempted to eat them, that they taste horrible. Perhaps you might be thinking, but why don't all ladybirds look the same and so give the same warning message that predators could learn just one message. It's most likely that this is the contrasting colours that are important, and whether it's black on yellow or red on black or two spots or 24 spots, it really doesn't matter. But also different colour patterns can provide benefits beyond warning predators, ladybirds with bigger or more spots can warm up more rapidly than those with smaller or fewer spots. And sometimes it can be beneficial for the lady bird to hide. And so the colour pattern might provide camouflage in some situations, ladybirds are just amazing!
Eva - I'll agree with that. Thanks Helen. Next week, we're racking our brains for the answer to this question from David.
David - How much of your brain is actual memory?










Comments
Add a comment