We find out how modern genetic techniques are helping to reveal more about our past - how we have evolved and moved around the planet, and the diseases that evolved to infect us...
In this episode

01:30 - Predicting the Asian Monsoon
Predicting the Asian Monsoon
Researchers this week have presented, for the first time, a record of Asian Monsoon data stretching over 700 years. The Asian Monsoon affects nearly five billion people each year but it involves a huge weather system and it's very hard to predict how it will change each year. Until now, there's been very little climate data available on the monsoon.
 Edward Cook and colleagues from Columbia University have measured tree ring data from over 300 locations and they've compiled it into what they call the 'Monsoon Data Drought Atlas,' or MADA. Published in the journal Science, they've been able to reconstruct how the monsoon varied from the end of the Medieval Warm Period, through the Little Ice Age (where Louis XIV's wine supposedly froze on his table) and during the more recent period of human-induced climate change.
Edward Cook and colleagues from Columbia University have measured tree ring data from over 300 locations and they've compiled it into what they call the 'Monsoon Data Drought Atlas,' or MADA. Published in the journal Science, they've been able to reconstruct how the monsoon varied from the end of the Medieval Warm Period, through the Little Ice Age (where Louis XIV's wine supposedly froze on his table) and during the more recent period of human-induced climate change.
The researchers plan to compare this record with others available so they could, for example, see how sea surface temperatures alter the monsoons. And occasionally, the monsoon fails completely - leading to droughts which destroy crops and cause all sorts of species devastation so finding the reasons for these events could help in their prediction in the near future.

02:54 - Sleep on it – and dream about it – to remember it
Sleep on it – and dream about it – to remember it
Have you ever found that the advice to "sleep on it" turns out to be true, whether it's solving a problem or trying to learn something? We've known for some time that sleep helps us to remember things, by helping the brain to file away and strengthen memories. Now new research from Erin Wamsley at Harvard Medical School, published in the journal Current Biology, provides more evidence that the best way to remember something is, indeed, to sleep on it - and, more importantly, to dream about it.
Wamsley and her team asked 99 volunteers to memorise the layout of a complicated computer-based maze. Then they were tested to see if they could get to a specific place in the maze after being dropped at a random starting point. Five hours later, the volunteers were tested again. But in the intervening time, some of them had been to sleep while some of them had stayed awake.
The scientists found that people who had an hour and a half shut-eye in between tests managed to get through the maze and average of around 3 minutes faster than the first time, while the people who'd stayed awake only managed to navigate it a mere 26 seconds faster.
As well as seeing whether the volunteers had had a cheeky nap, Wamsley also asked the nappers whether they dreamed about the maze. She found that people who had dreamed about doing the task during their nap improved in the second test far more than people who didn't. So it suggests that dreaming is a powerful form of mental 'rehearsal' for a task.
In Wamsley's experiments, her volunteers also had some pretty wacky dreams. For example, when the volunteers described their dreams, they didn't talk about specific things in the maze, such as certain points or router. But some of them did mention similar but related situations, such as different mazes, or being stuck in a cave.
And , intriguingly, they found that people who found the maze task most difficult were more likely to dream about it. So maybe their brains were more likely to be processing the information about the task - or worrying about the upcoming re-test - while they napped.
It's important to point out that the researchers don't think that the actual dreams themselves improve our memory - they're more like a side effect of the underlying brain 'filing' process that goes on while we sleep. But based on this research, you might draw the conclusion that it's best to study right before you go to sleep. Or, alternatively, this is a brilliant way to justify having a nap after a hard revision session.
05:50 - Following the squeeze on pills
Following the squeeze on pills
This week researchers in an international team from Switzerland, the Czech Republic and the US have managed to measure the forces felt by a small pill as it travels through the intestines.
 Bryan Laulicht and colleagues developed a technique using a dummy, magnetic pill. This was fed to both human and dogs and they tracked its progress using an array of magnetic field sensors held over the abdomen. Publishing in PNAs this week, not only could they detect the direction of gastric forces exerted on the pill but also the magnitude of those forces. And they could see how these changed through the stages of digestion.
Bryan Laulicht and colleagues developed a technique using a dummy, magnetic pill. This was fed to both human and dogs and they tracked its progress using an array of magnetic field sensors held over the abdomen. Publishing in PNAs this week, not only could they detect the direction of gastric forces exerted on the pill but also the magnitude of those forces. And they could see how these changed through the stages of digestion.
They found that on a full stomach both humans and dogs exerted similar gastric forces on the pill. But when both volunteers were fasted the dogs' innards exerted, on average, five times the force of the humans'. So we now know that dogs' gastric system is only similar to that of humans after they've been fed.
But this research is important because, in order to make tablets as effective as possible, we need to know how long they can last for inside the gut. For some pills, at least, the longer they last inside you the more effective they are at delivering their medicine. So the researchers hope that by modelling the gut forces in this way they can design much more sophisticated pills than those currently on the market.
07:29 - Stretching the limits of stem cells
Stretching the limits of stem cells
Stem cells research is a really exciting area of science, and one we often cover on the show. Now new research published in the journal Nature reports an important step forward in our understanding of stem cells, and how we might be able to use them in the future.
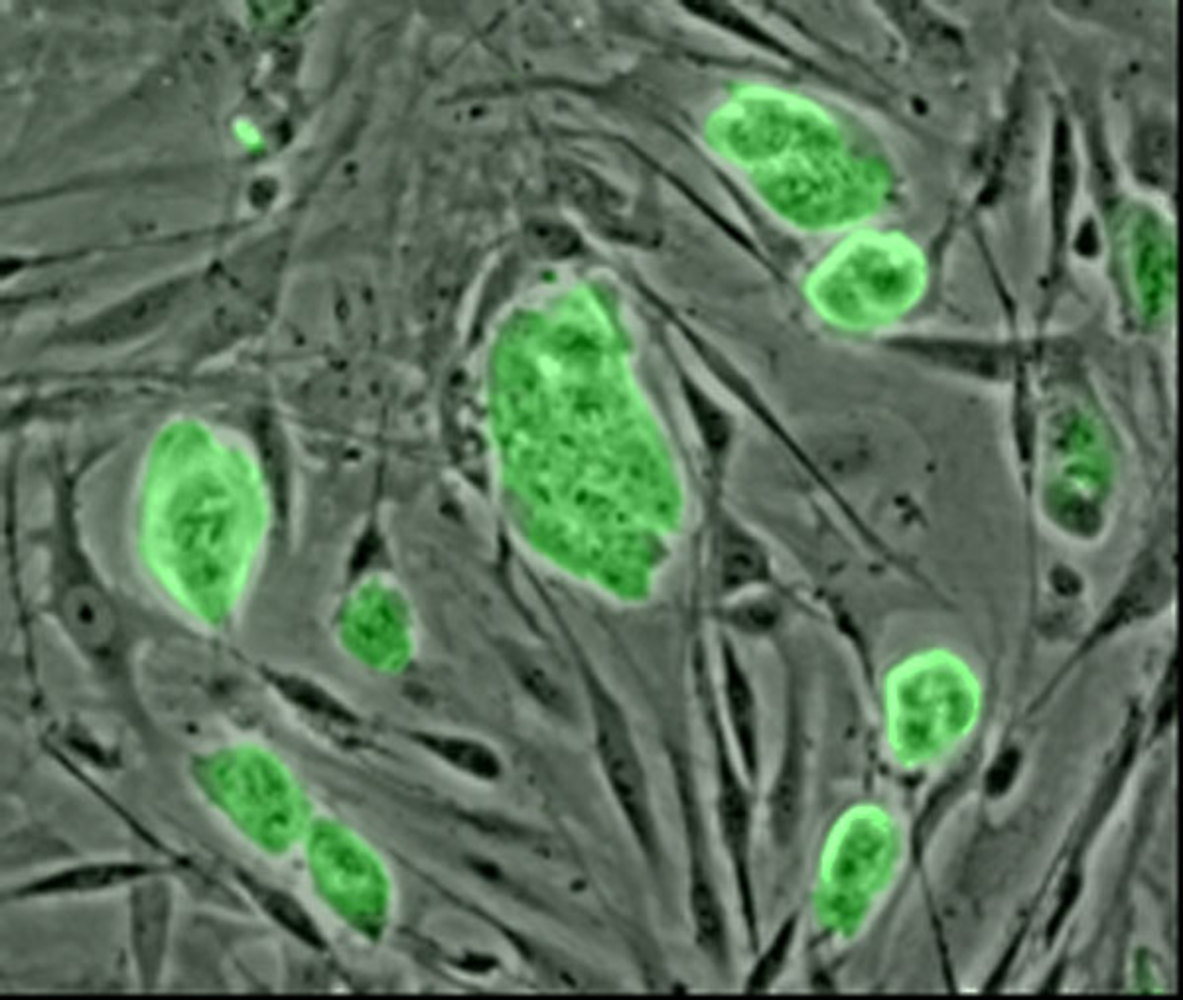 This is work from Konrad Hochedlinger at Harvard and his colleagues. They've been trying to understand the difference between cells known as pluripotent stem cells, which can make a limited number of different types of cells, and embryonic stem cells, which can be converted into all the 220 different types of cells in the body. Although these cells both have the same genes, only certain sets of genes can be used in pluripotent stem cells, limiting their potential. And cloning experiments have shown that while a single embryonic stem cell can give rise to a whole new organism, it's never been done with plurpotent stem cells.
This is work from Konrad Hochedlinger at Harvard and his colleagues. They've been trying to understand the difference between cells known as pluripotent stem cells, which can make a limited number of different types of cells, and embryonic stem cells, which can be converted into all the 220 different types of cells in the body. Although these cells both have the same genes, only certain sets of genes can be used in pluripotent stem cells, limiting their potential. And cloning experiments have shown that while a single embryonic stem cell can give rise to a whole new organism, it's never been done with plurpotent stem cells.
They compared mouse embryonic stem cells with genetically identical pluripotent stem cells, to look for key differences in the patterns of gene activity between the two. Importantly, they found that a cluster of genes on chromosome 12 were switched off in the pluripotent stem cells, but not in the embryonic stem cells. This region contains a number of genes that are important for fetal development.
The researchers looked at over 60 different pluripotent cell lines, and found the same genes switched off in the majority of them, suggesting that they are genuinely important. And when they tested whether they could generate cloned mice from the pluripotent cells, they could only make mice from the few pluripotent cells lines where the crucial genes weren't inactivated. In fact, this is thought to be the first time that this has been done using pluripotent stem cells.
This allows researchers to tell whether the cells they're dealing with have the potential to generate all different cell types, or just a limited range. This will become increasingly important in the future, as stem cell technology comes closer to medical applications - doctors will need to be able to choose the best quality stem cells for the job. And it also tells us more about how to change the properties of stem cells - depending on whether we want them to make a wide or restricted range of different cells.

10:23 - The Neanderthal in your Genes
The Neanderthal in your Genes
with Professor Jeffrey Long, University of New Mexico
Diana - Also in the news this week, a new genetic analysis of nearly 2,000 people from all over the globe suggests that our ancestors interbred with Neanderthals over at least two different periods. We're joined by Professor Jeffrey Long, leader of the team at the University of New Mexico, who have reported on this finding.
Jeffrey - Good morning. What we looked at were genotypes, genetic typings, from about 600 places throughout the genome from people from about a hundred different populations throughout the world.
Diana - And what did you look at in their DNA?
 Jeffrey - Well we looked at subtle variations that are sometimes called microsatellite loci or short tandem repeat loci. And these are just regions of DNA. Typically, they don't code for any products. They're sometimes called 'junk DNA' where people have small differences in the amount of DNA that they have. These are the kinds of genetic markers that are used in DNA fingerprinting and forensics and parentage testing, and a lot of routine tests these days.
Jeffrey - Well we looked at subtle variations that are sometimes called microsatellite loci or short tandem repeat loci. And these are just regions of DNA. Typically, they don't code for any products. They're sometimes called 'junk DNA' where people have small differences in the amount of DNA that they have. These are the kinds of genetic markers that are used in DNA fingerprinting and forensics and parentage testing, and a lot of routine tests these days.
Diana - So, do we know if the microsatellites actually do anything?
Jeffrey - There are a few circumstances where they do. Most notably, there's a microsatellite type of locus in the gene that is important for Huntington's disease. But there are only a couple of dozen of those where they're actually in genes that do anything. There are thousands of them throughout the genome and most of them - the vast majority of them - don't do anything.
Diana - So when you looked at these microsatellites, what sorts of genetic variations did you see?
Jeffrey - What we saw when we looked at these microsatellites is that people typically, and this is standard for genetics, have different amounts of DNA in each of these locations and they have more or less high mutation rates, by genetic standards at least. And because of that we have all of these variations and you get clustering of patterns throughout the world. And that's what we started looking at in this study. We were interested in a model that's called the serial founder effects model and with serial founder effects, what we've postulated and other groups have postulated, is that you have that original population in Africa and then a small group left Africa and populated Europe and Asia, and eventually Asia and the Oceania as well. We were interested in studying that effect.
Diana - So how did you find these clusters of variation varied geographically as well? Did they tie-in with these models?
Jeffrey - Well for the most part, what we found was that what you find outside of Africa is a subset of what you find in Africa. What you find in any region outside of Africa, say Asia, is a subset of all Africa and the like. But the thing that surprised us was that in the out of Africa group, there was a little bit more variation than the model could account for. So we found variation in Eurasians, people in the Pacific Islands, and the Americas that couldn't be accounted for by the out of Africa migration.
Diana - So you think this genetic information is coming from somewhere else?
Jeffrey - Yes. So we had to look at possibilities of where it could come from and what could account for it. Now, one of the things that our colleagues suggested to us is that it might be just by what we call gene flow which is sort of the 'boy marries girl next door' effect that goes on throughout the world where people in these different regions do occasionally mate with each other. But that couldn't account for the effect, and in retrospect, it really could not have because that sort of gene flow just spreads things around, it doesn't create or destroy variation itself. And then we eventually came to a conclusion of is it would have had to have been from, essentially, hominids that were like people that were existing in this area or at least we think that's the best explanation at this time.
Diana - So one possibility would be Neanderthals. And if it was, what does that mean for human populations?
Jeffrey - Neanderthals are certainly a possibility but there were many different sorts of archaic people around the world before we got the modern Homo sapiens evolving. But the main implication of this is that for the last one or two decades, we really believed that once Homo sapiens evolved, they replaced all of these people around the world and didn't mate with them or incorporate any of their genes. It was a very rigid speciation event. Now what this is telling us is that our closest relatives were pretty much similar to us and it was possible to interbreed and that perhaps the speciation event wasn't quite as rigid as we thought in the past.
Diana - Indeed, well thanks very much, Jeffrey. That's Professor Jeffrey Long on the genetic evidence that most people alive today carry a little bit of Neanderthal or potentially another early human species in their genes. This work was presented at the recent meeting of the American Association of Physical Anthropologists' Annual Meeting in Albuquerque, New Mexico.

15:55 - What is Archaeogenetics?
What is Archaeogenetics?
with Professor Colin Renfrew, University of Cambridge
Kat - Now this week, we're looking at the science of archaeogenetics where modern genetic techniques are used to compliment traditional trowel-in-the-soil archaeology and we're really honoured to have with us Professor Lord Colin Renfrew who's a Cambridge University archaeologist and he's credited with being the person who coined the phrase "archaeogenetics." Hello, Colin.
Colin - Hello.
Kat - So, we tend to think of archaeology as people on their hands and knees digging with trowels in the soil. How does archaeogenetics work? How do we meld modern genetics with these kinds of archaeological studies?
Colin - Well the remarkable thing is that most of what we've learned from archaeogenetics actually comes from specimens, blood specimens or genetic specimens, taken from living populations. The remarkable thing is that because the mitochondrial DNA is much the same from one generation to the next or the Y chromosome DNA much the same from one generation to the next, there's great stability. By comparing living populations, it's possible to construct trees which, it seems, give a very persuasive account of human decent. On the other hand, it is possible to take specimens from long dead humans and look at their DNA, the ancient DNA. But that is a much more difficult technique and the picture we have of the out of Africa migrations of our species around 60,000 years ago essentially comes from mitochondrial DNA taken from living people who are with us today or just of recent decades.
Kat - So let's look at a little bit more of that mitochondrial DNA because obviously, when humans are made by mommy and daddy, you mix mom and dad's chromosomes together, but why is mitochondrial DNA so special for these kind of studies?
Colin - That's right. You've put your finger on it as you say, most DNA - nuclear DNA is a mixture of the DNA from the mother and from the father. But the mitochondrial DNA is passed on in the female line. So your mitochondrial DNA comes from your mother and hers from her mother and so on, and indeed mine comes from my mother and from my maternal grandmother. So it goes down in a single line in that way, a single lineage. There is very little variation and less mutations which happen infrequently. So there's great stability and in this way, one can go back up the lineage as it were, studying those mutations, and getting overall, a picture which has led to the conclusion, and part of the conclusion comes from this work, that our species must have originated in Africa something like 200,000 years ago and all of us humans are the result of migrations out of Africa, something like 60,000 years ago.
Kat - So I've read about things like the original Eves and this is because you can follow the mitochondrial DNA up the female line. How far back can we really go? Could you almost pin-point one person or is it more just groups and populations?
Colin - It's a group certainly. One wouldn't want to pin-point one person although that may be theoretically possible. Well, you can go back to 200,000 years ago and that would be one of the very first humans of our species, Homo sapiens but you can of course go further up the decent line where you get to the point where our species began or the ancestors of our species began to separate out from the Neanderthals, something  like half a million years ago.
like half a million years ago.
Kat - And as we've heard, maybe not quite as separate as we previously thought.
Colin - Yes. Well that's a very interesting new discovery. The mainstream idea hitherto has been that our species, Homo sapiens, emerged out of Africa something like 60,000 years ago. And so, it's very exciting that there may have been Neanderthal survivors who may have made contributions to the Homo sapiens' genetic gene pool, very exciting discovery.
Kat - So we've discussed how we can follow the maternal line, follow the women by looking at mitochondrial DNA. What about the chaps, because obviously, you have a Y chromosome. Is that as useful or is it less useful?
Colin - Yes, it is as useful and it's been more difficult to do so the research has come about more recently but the Y chromosome has been extremely important because the non-combining part of the Y chromosome passes down exclusively in the male line. So in a way, it's rather a mirror image of what you'll find on the female line with the mitochondrial DNA. The only problem is that the chronology isn't secure. Dating is very difficult in genetics and there is less confidence about the precision of the Y chromosome DNA dating. But certainly, it gives a parallel picture which broadly supports the conclusions that have emerged over the past 10 or 15 years from the mitochondrial DNA.
Kat - And the sort of things that you're finding through studying DNA, studying mitochondrial DNA and Y chromosome DNA, how do you actually anchor them to a real life archaeological timeline and do you ever find that it disagrees with the archaeological timeline?
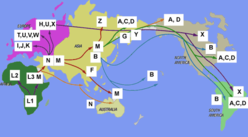 Colin - Well somehow, the research works to try and bring them into relationship. For instance, the mitochondrial DNA dating seems to correlate fairly well with the fossil record. For instance, we have the first humans in Australia, something like 50,000 years ago, and that ties in quite well with the out of Africa migration along the southern coasts of Asia so that humans reached Australia something about 50,000 years ago and the pattern works well. It works also in the Americas where it seems that our species reached North America something like 15,000 years ago or maybe a little more. So that the archaeology and the DNA do harmonise together to some extent, but as I say, it's only more recently that ancient DNA from actual fossil humans has been brought into play.
Colin - Well somehow, the research works to try and bring them into relationship. For instance, the mitochondrial DNA dating seems to correlate fairly well with the fossil record. For instance, we have the first humans in Australia, something like 50,000 years ago, and that ties in quite well with the out of Africa migration along the southern coasts of Asia so that humans reached Australia something about 50,000 years ago and the pattern works well. It works also in the Americas where it seems that our species reached North America something like 15,000 years ago or maybe a little more. So that the archaeology and the DNA do harmonise together to some extent, but as I say, it's only more recently that ancient DNA from actual fossil humans has been brought into play.
Kat - And do you find that really ties everything together and helps you to anchor points of this timeline?
Colin - Well it's certainly beginning to if we're looking for instance at the population history of Europe. When we look at Neolithic remains, and there one can get skeletons and look at the DNA, usually, the mitochondrial DNA which is usually better preserved, one can look at the haplogroups and begin to put together a story there. But there are quite a lot of problems and I don't think everything is yet clear. This discipline is obviously in its infancy and it has enormous potential because the data are abundantly rich or will be abundantly rich. But there are still quite a lot of problems of interpretation. For instance, if we're looking at the origins of language families, there's a lot of argument about that at the moment which it was hoped DNA might solve very rapidly. But the solution hasn't quite yet come, though I think it's on the way.
Kat - Just very briefly, what really intrigues you? What's the mystery that you really want to get your teeth into?
Colin - Well I myself would like to get clear on what are the origins of the Indo-European language family and that's difficult to determine probably because we're talking about languages there. And there are two main theories. One is that this language family might have emerged out of Anatolia with the coming of the first farmers to Europe. And there's another theory that it may have been to do with the domestication of the horse in the European steppes which have put it at a much later date. But to bring the DNA and the archaeology and the linguistics together successfully is quite complicated and I don't think we've quite got there yet.
25:21 - Understanding the Past with Ancient Pathogens
Understanding the Past with Ancient Pathogens
with Dr Helen Donoghue, University College London
Diana - Now it's time to join Meera Senthilingam. Meera's been out in London this week, finding out how the DNA of ancient pathogens, that's things like bacteria found in ancient human bones and tissues, can be extracted and sequenced to help us understand more about the evolution of diseases such as tuberculosis...
Meera - This week, I've come along to University College London to meet Dr. Helen Donoghue who's a senior lecturer in medical microbiology. Helen, you focus on the occurrence of particular pathogens in our past. So, what's the importance of understanding more about their occurrence in our past?
Helen - Well I happen to be in a department that has a long standing interest in a group of bacteria called mycobacteria that includes the bacteria that cause tuberculosis and leprosy. But it also so happens that these are particularly good subjects to look at in the past. This is because they are surrounded by very resistant cell walls which protects the DNA and the DNA itself, its chemical composition is more stable than that of some other bacteria.
Meera - What kind of samples do you work with and how old are they?
Helen - The age of the samples varies greatly. I think the oldest one I've ever looked at was from a Pleistocene Bison 17,500 years ago and the oldest human samples I've looked at were 9,000 years old. Generally, they're not quite so old as that. I've got large number of samples, for instance, from a collection from the 18th century in Hungary. So that would be the 17- or 1800s. And really, all the ages between then and ancient Egypt, everything in between. 
Meera - Now Helen, we are here in your lab in the department of infection. You've got some samples here in front of us today. One particular one I can see looks like a rib. So tell me more about this sample.
Helen - Okay. Well, I'll take it out so we can see it a bit more clearly. It's from a child. It's a lower rib from the Hungarians from the 1800s. It's curved but it's about 15 centimetres long. We had quite a few children in this group of individuals. Children are not quite so susceptible as adults for tuberculosis but the results I've got so far do show that many of them were infected with TB. The reason why we're interested in looking at a rib is because tuberculosis is an infection we normally catch by breathing an infected aerosol, it goes into our lungs and the lungs are the main site of infection. And when somebody dies, the evidence of the disease can be left on the outer surface of the ribs that were next to the lung.
Meera - Now as well as the rib, what other parts of the body can you target when obtaining your samples?
Helen - Well once tuberculosis breaks out of the lung and gets into the bloodstream, it can go all around the body. Also, you can catch tuberculosis by swallowing infected sputum or by eating or drinking infected milk or meat, perhaps from an infected animal. But actually, that's very rate in the archaeological record. But I've certainly found evidence of tuberculosis in the abdomen and in bones that don't show any sign of any lesion, and that's because they were infected by the bloodstream.
Meera - How careful do you have to be with your samples, I imagine - first of all they must be rare, and second of all you must not want to destroy very much of it?
Helen - Well luckily, we don't need very much. So normally, when I do an extraction, I use between 20 and 60 milligrams of sample, that's if it's bony sample. You need rather more than that if it's mummified. You can have that much without destroying the bone.
Meera - So I guess having identified if particular individuals from the past had tuberculosis, how does this all come together? What does it tell us about our past?
Helen - Well let me just say something about one of the collections I've got which is 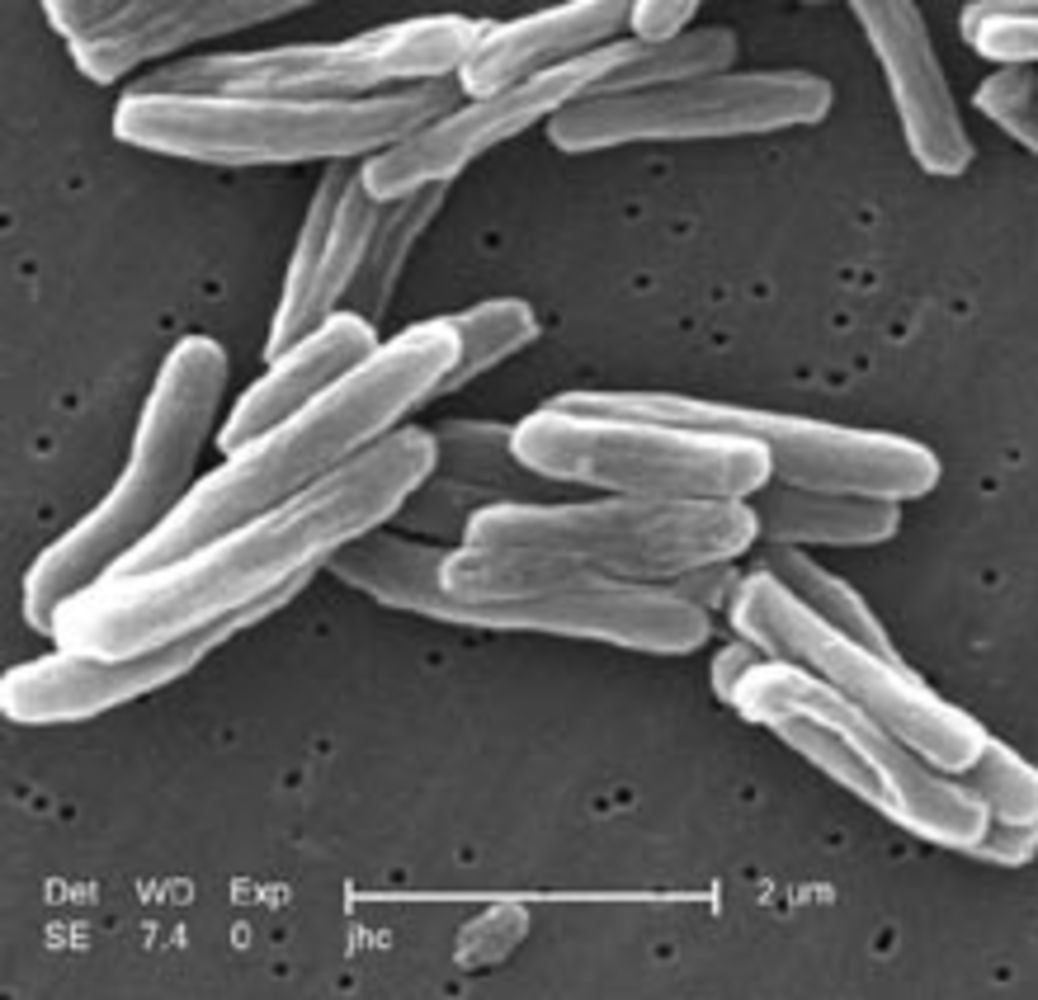 from the 18th century Hungarians. They've got a very good archive so we know the age and family relationships of many of the individuals, and there are over 200 of them. That means we can do some epidemiology. We can look at family groups. We can look and see, because the samples were very well preserved, we can actually look at different lineages of the MTB (Mycobacterium tuberculosis) DNA.
from the 18th century Hungarians. They've got a very good archive so we know the age and family relationships of many of the individuals, and there are over 200 of them. That means we can do some epidemiology. We can look at family groups. We can look and see, because the samples were very well preserved, we can actually look at different lineages of the MTB (Mycobacterium tuberculosis) DNA.
We know there were at least two distinct lineages of the MTB DNA at that time and this is from an era before there were any antibiotics. We know there were many people in this population who lived to a great age even though they were infected with TB. We know there were other people who died early in life or in their early 20's of TB. This gives us the chance to look at the evolution of the bacterial infecting organism and its human host over time from an age before there was any distortion in how these bacteria evolve, because there was no antibiotic therapy. That enables us to compare that with the modern day trend and we can see that there is now a logarithmic, an exponential, increase in the rate of change of the organism.
Meera - And how important would you say it is then to understand more about tuberculosis?
Helen - Tuberculosis is probably the greatest cause of death from any single infectious disease in the world today. According to the World Health Organization, we think one third of the world's population is infected and in 2007, there were just under 2 million deaths. This is from a disease where we can treat it with antibiotics, even though drug resistance is becoming an increasing problem. So we really do need to understand our relationship between ourselves and this particular species.
Meera - How can this research help us understand more about our past and our ancestors?
Helen - Well, both tuberculosis and leprosy are examples of clonal diseases where the infecting bacterium has sometimes a lifelong relationship with its human host. And so, as humans migrate, these bacteria migrate with them. So by studying these organisms or the DNA from these organisms, from samples from different geographical sites around the world, we can understand how humans have migrated around the world.
Diana - That was Dr. Helen Donahue from University College London, talking to Meera Senthilingam about how reading an ancient pathogen's DNA can help us understand how the bacteria that cause diseases like tuberculosis have evolved alongside us.
Kat - And if you want to check out Helen's work, there's an exhibition showcasing her work on ancient tuberculosis called Body Matters which will be in the Leventis Gallery in the UCL Institute of Archaeology from the 17th of May.

32:40 - DNA Profiling an Ancient Person
DNA Profiling an Ancient Person
with Professor Eske Willerslev, University of Copenhagen
Diana - As well as following the way populations have changed and migrated, we can use modern genetic techniques to really get to know an individual body as long as it's preserved well enough. Professor Eske Willerslev is from the University of Copenhagen. Where he's the leader of an ancient DNA and evolution group. So Eske, can you tell us what are the problems about getting ancient DNA samples?
Eske - Well, I think it varies a lot. The problems, or the magnitude of the problems, actually varies a lot from one specimen or one type of specimen to another. But obviously, one of the biggest problems you have when you're looking at ancient human remains is due to contamination with modern human DNA, because many of these specimens have been handled by humans without protective gloves or anything like that. And also the reagents that you're buying have all been produced by humans so we can actually have traces of modern human DNA in there.
 Diana - So how do you get around that? How can you avoid that contamination?
Diana - So how do you get around that? How can you avoid that contamination?
Eske - I think that in the last few years we've actually seen a revolution within ancient human genetics in the sense that some new technological breakthroughs have happened that makes it possible for one thing, getting very, very short pieces of DNA out of the specimens. And the thing is that the copy number of a certain piece of DNA in indigenous DNA actually increases tremendously if you go shorter and shorter in terms of the DNA you try to retrieve. So that makes a huge, huge difference. Basically, if you go short enough, you can almost swamp the contaminant DNA by the indigenous DNA. Additionally, you can say the knowledge we have gained about how DNA gets damaged through time can also provide evidence to what extent you're working with indigenous DNA or are suffering from contamination. So some of these breakthroughs really means a lot. Another thing is that if you're working with specimens from another ethnic group for example, let's say specimens that had been in Britain, stored in Britain but comes from non-British people. Well then there's actually ways to, you can say, distinguish the DNA you have from being European to some other ethnic groups. So there is ways around it.
Diana - So you're using these shorter pieces. But can you then sequence it in the same way that you sequence fresh DNA?
Eske - Yeah. In fact, you can. I mean, with these second generation high throughput sequencing platforms that have appeared over the last few years, there you can actually sequence ancient DNA in the same way as you can with modern DNA. Of course, there are some issues you don't have with modern DNA. For example, if you have a bone specimen you know, it's full of bacteria for example, additional to any modern human contamination that must have penetrated the bone. And therefore, a lot of what you get out when you do what we call shotgun sequencing will in fact be microbial DNA from such a specimen while from a modern sample, you know, 99 point something percent of the DNA would be human. So it's more demanding to deal with ancient DNA certainly.
Diana - And probably a bit more expensive. But tell us about Inuk, the guy you studied. What did you have to work on there?
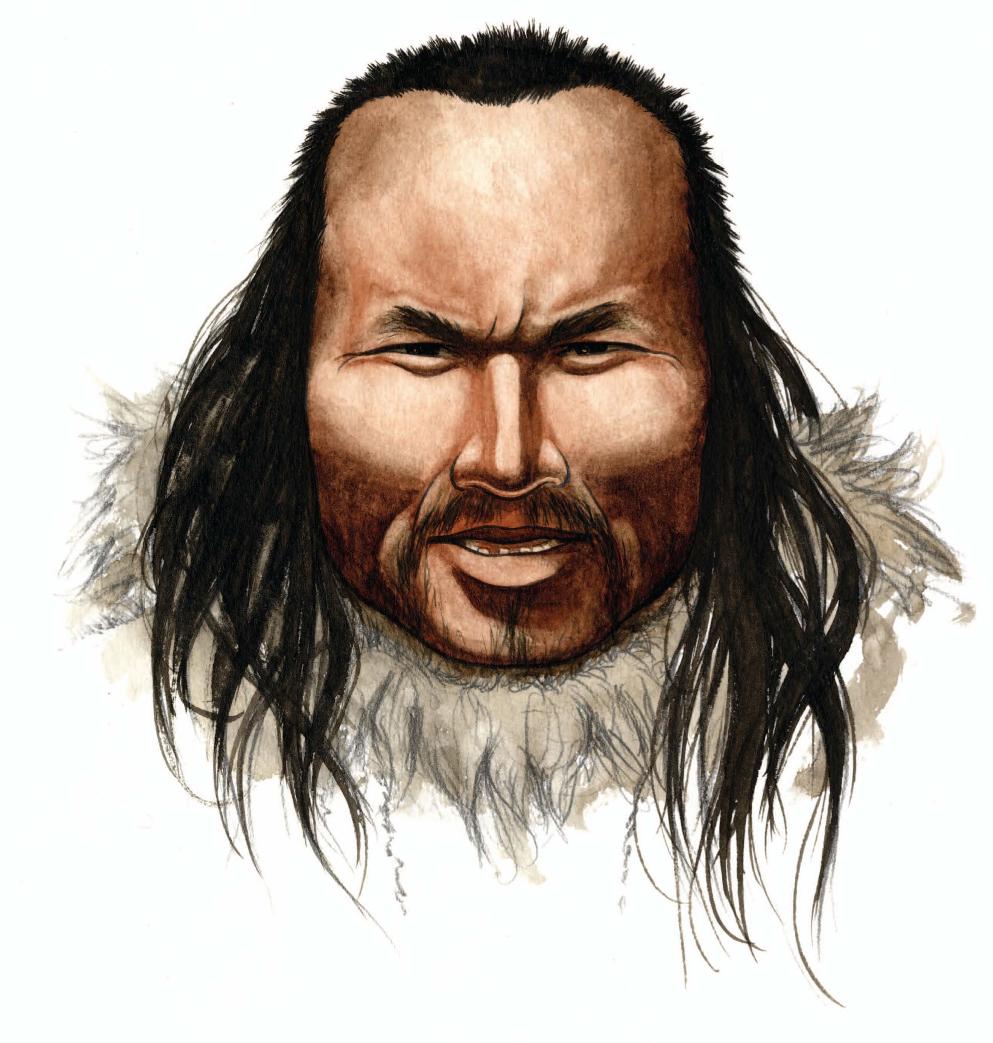
Eske - Yeah. There we worked on ancient hair. So it's hair from a 4,000-year-old individual. And hair is actually a really good source of ancient DNA in the sense that it's not porous like bone and therefore, neither modern human contamination or microbial contamination over the years penetrates very far into the hair, and all the contamination is on the surface. So it's quite easy to clean the hair samples from contamination, [compared to] others like bones for example that are porous and where the contamination penetrates very deep. So, from that point of view, it was reasonably easy to actually sequence the majority of the genome, the complete genome of this individual.
Diana - And once you've done that, what did you find out about him?
 Eske - Well that was actually quite interesting and I think it shows some of the power of using ancient DNA. First of all, the only thing that people have actually found from this culture, which was the first people who settled in the new world arctic - Alaska, Canada and Greenland, is in fact a few pieces of hair and four small pieces of human bone. And that's all we know biologically about these individuals. By sequencing the genome of this person we could show he is a male. We could also show he had brown skin and dark eyes. We could see he had a tendency to baldness for example, so he probably died quite young because it was a fair amount of hair we found from this guy. We could also see dry ear wax. We could see that he was genetically adapted to living in cold temperatures although his people hadn't been there for a very long time. We could see what blood type this individual had. We could say something about his body's stature, that he had relatively low surface area compared to body mass and stuff like that. So you know, there was a quite a number of phenotypic traits that we can actually retrieve from the genome but additionally to that, we could actually see that he came from a migration that had been independent of those migrations that resulted in present day Inuit - people in the new world Arctic. And also independent of that migration or those migrations, giving way to contemporary native Americans.
Eske - Well that was actually quite interesting and I think it shows some of the power of using ancient DNA. First of all, the only thing that people have actually found from this culture, which was the first people who settled in the new world arctic - Alaska, Canada and Greenland, is in fact a few pieces of hair and four small pieces of human bone. And that's all we know biologically about these individuals. By sequencing the genome of this person we could show he is a male. We could also show he had brown skin and dark eyes. We could see he had a tendency to baldness for example, so he probably died quite young because it was a fair amount of hair we found from this guy. We could also see dry ear wax. We could see that he was genetically adapted to living in cold temperatures although his people hadn't been there for a very long time. We could see what blood type this individual had. We could say something about his body's stature, that he had relatively low surface area compared to body mass and stuff like that. So you know, there was a quite a number of phenotypic traits that we can actually retrieve from the genome but additionally to that, we could actually see that he came from a migration that had been independent of those migrations that resulted in present day Inuit - people in the new world Arctic. And also independent of that migration or those migrations, giving way to contemporary native Americans.
We actually found evidence of a previously unrecognized migration from the old world into the new world happening some times around 5,500 years ago.
Diana - That seems a bit more recent than previous estimates have been put. So you know that's definitely independent because of the date?
Eske - Well, it's definitely independent and it's earlier than you can see in that migration that they raise to present Inuits or Eskimos, if you want. Culturally they're only around a thousand years or a little bit more than thousand years old. But the thing is, we could see that this guy is actually more closely related to people in north-eastern Siberia than he is to either Inuits or to native Americans.
Diana - All that from a few pieces of hair. Thanks, Eske. That's Eske Willerslev on what we can learn from seemingly insignificant remains provided there are some DNA.

Where and when did all the different human ancestor species originate?
We put this question to Professor Lord Colin Renfrew:
Colin - Well the fascinating thing is that the fossil remains give us a very clear picture and most of them originated in Africa. There was a theory that to maybe the Homo erectus, form that was about 2 million years ago, might have been ancestral to our species, Homo sapiens. But the DNA helps confirm that you have to go back to Africa to find the ancestors of Homo sapiens. Back to Homo erectus, before that 3 million years ago, Australopithecus... So, Africa is the answer.
Diana - I see and how long ago are we talking? I mean, you mentioned 3 million years.
Colin - Yes. Well, the African ancestors go back before that but things start to happen sort of 3 or 4 million years ago. You've got Australopithecus and then you get the first tool makers, which is very important, about 3 million years ago with Homo habilis. So, once you get people making tools - stone tools, that's when you really feel that you're dealing with our real ancestors.

Does dyeing your hair make it turn white faster?
Kat - No, I think that this isn't true because hair dye works on your hair which is kind of dead. The bit that produces the colour of your hair is the pigment, the melanin pigment that's in the hair follicle and this isn't affected by hair dye. I think probably if you're dyeing your hair, you might notice that you're going grey, the white hairs coming through more or maybe when you stop dyeing it, suddenly you're like, "Crikey! I'm really grey." But I don't think that dying your hair is actually going to affect your hair follicles. So no, I don't think it makes you go white faster.

48:55 - How Old is My Grandmother?
How Old is My Grandmother?
We put this to Kirsty Spalding, assistant Professor in the department of cell and molecular biology at the Karolinska Institute, Stockholm, Sweden: Kirsty - There's a couple of ways you could do this, both involving a newly developed strategy which is radiocarbon dating. Essentially, because of the Cold War, there was a whole lot of above ground nuclear bomb testing and during this procedure, there was a lot of detonations which cause neutron emissions into the atmosphere. And essentially, to make the long story short, this ultimately resulted in increased levels of carbon-14 in our atmosphere. This actually occurred for some years during the period of the Cold War then they put this Test Ban Treaty out, banning all above ground nuclear bomb testing. And then the C14 levels in the atmosphere changed in a very predictable way with time so that for any given time point, the proportion of C14 to C12 represents a particular year in time. So, one way we've developed to use this strategy to look at cell turnover in the human body is to look at the DNA of cells. And by determining the proportion of the radioactive carbon-14 to the stable isotope carbon-12, we're able to say when this cell was born. And we've been applying this to different parts of the human brain and body, and actually found that there are selected regions of neurons, that's the nerve cells of the brain, that are as old as we are. And so, by taking some brain cells from this region of the brain; cortex, cerebellum for example, we're able to take out the DNA from these cells, carbon date them and they will tell us the year of birth of the individual.
Diana - But what if you don't particularly want to take a sample from a living person's brain?
Kirsty - This is perhaps a more cumbersome way to determine age. Another way that I've developed with colleagues and this actually uses the C14 to C12 ratio in tooth enamel. So in this methodology, you can take a tooth from an individual so in this case from your 100-year-old granny and depending on which tooth it is, you determine how old the enamel is and each tooth lays down the enamel at a different time point which we've decoded in a way, so we know how long it takes for each tooth to make enamel. We can figure out then from this information with carbon dating of the enamel, when the person's born and we can combine this with other methodologies to then find out when they died and how old they were when they died. So this is without a doubt the most precise way to determine the age of an individual.










Comments
Add a comment