Cancer is a devastating disease, and one of the largest killers in the Western world. This week, in a special show, Kat Arney investigates how scientists are fighting back, from building tumours in the lab to a "Google Earth" for cancer.
In this episode

00:47 - What is cancer?
What is cancer?
with Dr Emma Smith, Cancer Research UK
Around the world, more than 14 million people are diagnosed with cancer every year, and that figure is expected to rise to more than 21 million by the year 2030 so it’s a growing global problem. Kat Arney spoke to Emma Smith, science communications manager at Cancer Research UK, to get the basics about cancer.
Emma - Normally cells are very highly controlled. They grow only when your body needs them and when there are signals to do so. Now in cancer, some of these signals, some of the genes in the cells have gone haywire. For some reason the cells start dividing and multiplying out of control and then you have too many of a certain kind of cell, and that’s when you get a tumour forming.
Kat - Are we talking about cancer as one disease? There is the ‘big C’ - is it just one kind of thing?
Emma - Absolutely not! We are made up of hundreds of different types of cells and cancer can appear in lots of different tissues: the bowel, the lung, the pancreas, everything and all these different types of cancer have different characteristics. They’re caused by different faulty molecules and different faulty genes and, of course, that will require different treatments. So we absolutely cannot treat cancer as one type of disease.
Kat - What do we know about what causes different types of cancer? People blame things in the environment, people blame stress, some people say well, it’s just in your genes isn’t it?
Emma - It’s a really, really complicated mixture of all of it. Some of it is down to things that we do, ways we live our lives, things in our environment. We call these preventable causes of cancer. These are things in our environment that can cause damage to our DNA, and it’s this damage to our DNA that introduces a fault and the faulty gene can accidently tell the cell to start multiplying and, bingo, you’ve got a cancer.
But, of course, that’s not the end of the story. Some of cancer is down to - I’ve seen it called in headlines “bad luck.” Well, let's not call it bad luck, let's just call in nature instead. All of our cells are capable of dividing and, every time they divide, unfortunately, as much as we would like to believe it, humans aren’t perfect. As a cell copies itself and copies its DNA, it can just accidentally introduce a mistake. It’s not a 100% perfect process and just by doing so, again you can end up with a fault in a key gene and possibly cancer developing. So that’s the kind of nature side of it.
But chemicals and things in our environment can also introduce these DNA mistakes, so it is a mixture of both. Around 4 in 10 cancers are, let's call them “preventable,” so there are lots and lots of things that people can do if they want to reduce their cancer risk.
Kat - You’ve mentioned faulty genes and molecules can drive cancer. So how much cancer is actually in the genes? Can it be inherited - what’s the story there?
Emma - It can be inherited. There are certain examples where people inherit a fault from either of their parents and this particular gene is a really important and, therefore, it increases a risk of cancer. But this is really a minority of cancers. Most cancers are simply a result of nature, or simply getting older. Because damage builds up in our cells over time; all those things that I mentioned in our environment. So there are certains genes, but these form a very small proportion of cancers that we see.
Kat - So, so far we know that cancer is not one disease; it’s many different diseases. There are complex causes. There’s an interplay between the things in our environment, the things we do, and the things in our body itself. But then how do we treat cancer? Presumably, it’s not one size fits all for treatment either?
Emma - Absolutely not, but treatments can be broadly grouped into categories. Drugs that often make the headlines and modern drugs are very expensive. But these drugs aim to specifically home in and target cancer cells; they’re called targeted therapies. Chemotherapy, which is one of the cornerstones of cancer treatment. It’s more general in that it’s not specifically designed just to hit cancer cells but it is, in some cases, very, very effective. The other is radiation treatment, so radiotherapy and, of course, surgery. Put them all together and, usually, patients receive a mixture as well. They don’t just receive one therapy; normally it’s a combination of several different types of therapy.

05:24 - Zooming in on radiotherapy
Zooming in on radiotherapy
with Alison Tree, Royal Marsden Hospital, Uwe Oelfke, The Institute of Cancer Research
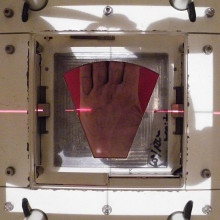
One cancer treatment in the arsenal of doctors is radiotherapy. It may not be as well known as chemotherapy or the expensive cancer drugs that hit the headlines, but radiotherapy is the cornerstone of many successful cures. Kat Arney caught up with Uwe Oelfke, and before him Alison Tree, a consultant clinical oncologist at the Royal Marsden Hospital, who’s a specialist in treating prostate cancer, to find out more about radiotherapy.
Alison - Radiotherapy is high energy X ray treatment that we use to target cancers to kill off the bad cells, the cells that cause cancer. It works because the cancer cells turnover very quickly - that’s part of what makes it cancerous. Whereas your normal tissue turnover very slowly so they’ve got plenty of time to repair any damage caused by the radiotherapy. Whereas the cancer cells get killed off and, hopefully, the cancer is then cured.
Kat - What sort of cancers can be treated with radiotherapy?
Alison - Most sorts of cancers can be treated with radiotherapy and, actually, radiotherapy is involved in about 40% of cancer cures. So it’s a very cost effect and effective treatment for cancer. Most cancers, for example, prostate cancer; the largest proportion of prostate cancer is cured by radiotherapy rather than surgery.
Kat - What sort of research is being done into how best to target this radiotherapy? It seems very powerful, but you’ve got to make this a precision weapon.
Alison. Yeah, absolutely. At the Royal Marsden, we’ve used clinical trials as a way to accelerate progress. Unless you test something you can’t assess the benefit of it, and my colleagues have created some of the trials that have changed the way we treat prostate cancer, for example, over the last ten years. And part of that innovation is in the way we deliver the treatment. We can effectively shape the beams, shape the way the beams deliver their dose around the cancer and spare the normal tissues, which reduces the side effects of treatment.
Kat - One of the problems with cancer is that it’s inside the body, and if you’re directing beams of radiation at a person, surely you have to know where that cancer is inside them. How do you find a cancer inside someone so that you know you’re targeting it?
Alison - Good question. At the moment what we do is we direct our beams based on CT. CT is a fairly standard scan that many people have had. And we have the ability to do CT scans just before the patient has their radiotherapy while they’re on the bed waiting for their treatment. But CT is not really the best way to image cancer. Most cancers are made of soft tissues and bone cancer is very rare.
Soft tissues are better shown by MR scanning - MRI imaging. And that is where the MR Linac, a new machine that we have at the Royal Marsden, will be become very important because we’ll be able to see the cancer much better. Now this is a very technologically difficult thing to do. If you put radiotherapy and a magnet in the same room, they interfere with each other and so it’s taken many years of collaborative developments across the world really to make this a reality.
Kat - One of the other things about human beings is that they move, they breathe, bits of us inside us kind of wriggle around. Does radiotherapy take that into account?
Alison - Yeah. We can do everything we can to set the patient up at the start of their treatment in the right position. But, as we stand here talking, our bladders are filling, our lungs are going up and down. The cancer, if we had any - I hope we don’t - inside them would be moving as well. So, at the moment, the best we can do is put a safety margin around our radiotherapy field to incorporate that movement and make sure we don’t miss the target.
But in the next generation, in the next decade of innovation, we will be able to see the cancer better and track it as it moves during the treatment, and maybe even adapt our radiotherapy beams and our plans during the treatment.
Kat - What are the big questions for the future of research into radiotherapy?
Alison - Our research here, particularly into the MR Linac, which is only one part of our research strategy, we’re working, as well as with collaboration all over the world, with our partners at the Institute of Cancer Research. We’re trying to establish where the MR linac is most able to make things better for patients, where we can change the way we treat cancers, try out new doses of radiation in different way that we hope will be more effective, and cure more cancer with less side effects.
Kat - In terms of reducing the number of hospital visits for radiotherapy, what sort of numbers are we talking about?
Alison - My colleague, David Dearnaley, led a trial called “the trip trial,” which has reduced the standard treatment for prostate cancer down from seven and a half weeks of treatment down to four weeks, and that’s become standard across the NHS already.
We are now conducting the next trial, which is called “the pace trial” led by Nick Vanas, which is comparing the now standard 20 treatment, so four weeks of treatment versus just five daily treatments of radiotherapy for prostate cancer. So a big effect on the patient’s quality of life and their ability to just get on with life and forget they’ve had cancer. But the ultimate goal is to test whether we can got to the ultimate in radiotherapy - can we cure prostate cancer with a single treatment? We don’t know the answer to that yet, but that’s where our research should go over the next decade.
Kat - Just one dose. That’s almost the “nuke it from orbit” approach. Is that going to work?
Alison - As with every innovation you have to test it robustly. You’ve got to start in a very safe and controlled way and test this kind of treatment on patients that you think will benefit from it. Then, if the initial data is promising, then expand it to a bigger number of patients. But this is very much the future vision, the blue sky vision, and we don’t yet know if this will turn out to be the right thing but there are theoretical reasons why it should. Prostate cancer is very sensitive to bigger amounts of radiation in a single dose and so there’s a theoretical reason why giving just one treatment might work in prostate cancer.
Kat - Alison Tree from the Royal Marsden Hospital. As Alison just explained, of the newest developments in radiotherapy is the ability to combine imaging - being able to look at tumours inside the body - with delivering the radiotherapy.
At the Royal Marsden Hospital, together with The Institute of Cancer Research, they’ve just unveiled something known as an MR Linac - a brand new state-of-the-art device that marries a magnetic resonance imaging, or MRI, scanner with a Linac - the machine that produces the radiotherapy beams.
Funded by the UK Medical Research Council it’s one of only two in the country, with the other still under construction up at the Christie Hospital in Manchester. Professor Uwe Oelfke, leader of the Radiotherapy Physics Modelling group, introduced me to this new piece of kit, which sounds more like a steam train than a medical device...
Uwe - This is the MR Linac facility at the Royal Marsden Hospital in Sutton, and we are standing between the operating room, operating this new machine and the facilities for handling the patients.
Kat - What are the benefits then of being able to see exactly where you’re treating someone, where you’re delivering this cancer killing dose of radiotherapy?
Uwe - The advantages are twofold. First, if you concentrate the radion just within the tumour, this enhances the probability that a tumour is controlled. And secondly, of course, every exposure of healthy tissues induces, to some level, toxicities, and these toxicities are also reduced.
Kat - So you’re going to reduce side effects and then, hopefully, enhance the treatment?
Uwe - Yes. We can use this; it’s called the therapeutic window between tumour control and normal tissue complications. And we can either use the same dose for the tumour and reduce toxicity, or we can use the reduced toxicity to enhance the dose within the tumour to dose escalate the tumour such that the tumour control will go up, while the toxicity still stay in tolerable limits.
Kat - How will this new treatment be used?
Uwe - This new treatment will be first used to really dynamically adapt the radiation fields to the anatomy we are treating. So this is still purely anatomically based but the next step with this kind of machine, you can also take other images. Not just anatomical images but functional images which basically inform you about how much oxygen is available in a tumour. These kinds of functional images then could guide further how we dose a tumour.
For instance, it is well known that tumours are lacking oxygen, that you need more radiation dose and these kinds of effects, at the moment, we cannot really take into account, or it’s very difficult. With this machine, first step is purely anatomical guidance and the next step would be biological guidance.
Kat - We’re currently in the facility and it’s completely quiet - there’s hardly anyone here. So, obviously, it’s not treating any patient’s right now. When will patients start to be treated here?
Uwe - We are hoping that the first treatments will either start this December or at the beginning or next year.
14:55 - Building tumours in the lab
Building tumours in the lab
with Professor Fran Balkwill, Queen Mary University of London
Cancer causes tumours, but tumours aren't just made out of cancer cells, as Kat Arney found out when she went to visit Fran Balkwill, Professor of Cancer Biology at Barts Cancer Institute, Queen Mary University of London, who is working on ovarian cancer, which spreads inside the body, particularly into the peritoneum - that’s the lining inside the abdomen - and the omentum, the scientific word for tummy fat.
Fran - My interest is the entire cancer. A lot of people think that cancers are born of malignant cells that have gone out of control. And this is true but that’s only half the story. When we think of really, probably all cancers, then they are actually a mix of the malignant cells, the cancerous cells, but a whole host of other cells that the cancer has recruited and, in some ways, corrupted to help the cancer grow and spread. 50% or more of any lump that is a cancer can be these normal cells that are promoting the cancer growth and spread.
Kat - What type of cancer are you looking at?
Fran - We’re looking at a type of ovarian cancer called high grade serous ovarian cancer. What you find when you look is that you find a mixture of fat cells, which are in parts of the peritoneum, and we think the cancer can use that fat to help them grow. Then you find fibroblast builder cells, you find new blood vessels, you find lot’s and lot's of immune cells that really should, and we know they could, fight the tumour but they don’t, and the malignant cells as well. There’s up to 20 or more different cell types actually in a cancer alongside the cancerous cells.
Kat - But certainly from my experience of working in a lab, you just look at cancer cells growing in a plastic dish but, obviously, that doesn’t capture all this diversity you’re talking about?
Fran - No. Absolutely and that’s the problem. Now that’s why we use mouse models of cancer because then you can in a living organism, but sometimes we need to have all human cells. Some of the new treatments actually have antibodies that act against human cells and not mouse and so we really need to compliment the mouse models, we need to have some human models. So how do we set about this?
Kat - The human lab…
Fran - Yeah. The human lab. We are not made of plastic and cancers are more than just cancer cells growing on a plastic plate. But with all the advances in bioengineering and regenerative medicine we think, and many people are trying to do this now, that we can begin to build complex three dimensional models of more than one cell type. More than just a cancer cell and answer some of these basic questions about how tumours form as well as what new treatments we can use.
Kat - How do you go about doing that then? How do you about reconstructing all this three dimensional glory of a tumour in the lab?
Fran - It’s with great difficulty I think I would say. But the approach that we’ve taken is to first deconstruct, so we get from surgery the omentum, which is part of the peritoneum from patients with high grade serous ovarian cancer.
First we found out many things, not all of them, but many things that we thought would be useful so that we would have a kind of template for a model, but also we would be able to validate. We would be able to say how close is any model we build to the real thing. So we did that for quite a long while and that’s proved extremely useful and also rather interesting. We learnt things about the tumours that we didn’t know before when we were just looking at the cancer cells.
Now we’ve taken various approaches; the most promising is to build an artificial omentum because in normal people the omentum is mainly fat cells. So we’ve been able to grow an artificial omentum and we have some idea how stiff it should be from the deconstruction and we have some idea about what other cells are there. Then we are growing the tumour cells, and the fibroblast builder cells, and also some other cells called mesothelial cells and we’re trying to recreate a model where the tumour is first invading that omentum. And also another model where the tumour is already there in situ with all the cell types.
We haven’t got all the cell types yet and the next challenge is the immune cells. But we’ve got some of the major cell types and all from the patient putting together and now we’re just testing to see how close those cell types resemble, in some of their functions and things that they make and things they do, the tumours that we take from the patients.
Kat - So it’s a bit like taking a house and unbuilding it brick by brick, taking everything apart, the beams. the bricks, the insulation, looking at what you’ve got and then trying to reconstruct that again?
Fran - Yeah. I think that’s not a bad analogy. It’s very complicated. We’re never going to be able to completely recreate the cancer obviously. The important thing now is to find questions that we cannot answer in any other way, or we need to complement our studies in the mouse models and then work out what combination of cells we think can answer those questions.
Kat - So now you’re managing to grow these little tumours, these little systems in the lab, what’s the ultimate goal? What are you going to do with them?
Fran - First we need to check that they do enough to resemble the human tumour and then, from our point of view, we want to find new treatments that disrupt the interaction between the malignant cells and all those cells that they corrupt and recruit because, if we can do that, we can break of some of the supply lines of the tumour. We also want to use them to understand how the tumour cells and the other cells interact to form tumours in the first place. So it’s understanding some basic cancer biology, but also a complementary way to the other ways we have of testing new treatments.
Kat - I look at some of the stories that are coming out of world of tissue engineering, and researchers using things like 3D printers to think about making new organs and all this kind of stuff. What would be your dream for making these kind of tumours in the lab?
Fran - Well, I think that you’re absolutely right, and we’ve just got some funding to get a 3D bioprinter. It’s going to take us a while; it’s not going to happen overnight but I think it’s a nice idea. I was talking about this to a load of clinicians the other day, oncologists. One came up to me afterwards and said “do you know when you were talking about that, I had a vision of a bioprinter printing little tiny living people.”
Kat - That would be the ultimate model, wouldn’t it?
Fran - Yes. Unfortunately there’s something about the size of cells that make us the size we are and, unfortunately, you can’t make tiny people but it’s just a wonderful idea.
23:12 - Is cancer in your genes?
Is cancer in your genes?
with Nazneen Rahman, Anguraj Sadanandam and Richard Houlston, The Institute of Cancer Research.
Over a century ago, the German biologist Theodor Boveri put forward the idea that something inside cells was responsible for causing cancer. Today, we know that the disease is fundamentally driven by faults in the genes that normally act to keep our cells and bodies healthy, only dividing when they should, and dying if they’re damaged. Kat Arnet went to visit The Institute of Cancer Research in Surrey to meet some of the UK’s leading experts on cancer genetics and find out where we’ve come from in terms of understanding the faulty genes that drive the disease. First, she spoke with cancer genetics expert Professor Nazneen Rahman, to find out how studying genetics is useful in cancer.
Nazneen - There are different ways in which we use genetics in cancer. The first fundamental difference is whether we’re actually testing the cancer itself or whether we’re testing the whole person.
If you test the cancer itself, there are all sorts of genetic changes that have made that cell turn into a cancer cell, and by doing that kind of genetic testing we can work out what type of cancer it is and what kind of treatment it might have and that’s one side in which genetics is incredibly useful.
We can also look at the DNA of the whole person to see whether there was some potential reason why they developed cancer in the first place. And also whether or not there might be information there relevant to other members of the family due to inherited predisposition to cancer. So that’s the other way in which genetic information is incredibly helpful in cancer.
Kat - This genetic connection raises the question of whether cancer runs in families. But, as I found out from Professor Richard Houlston, who has spent his career tracking down genetic variations that increase cancer risk, just having several cancer cases in a family doesn’t automatically mean that there’s an inherited gene fault at work.
Richard - As many people know, cancer is actually a relatively common disease so, effectively, if everyone lives to a ripe old age, then you’re going to get lots of cancers in the family. Because, essentially, in most Western countries it’s that and coronary heart disease and then stroke are the major causes of death.
So the next point is some cancers are highly environmental or lifestyle induced. Examples are: if you had a large number of people who are elderly who developed lung cancer and they’re all smokers, you get a conglomeration of cancers in the family simply because of that.
What really alerts you to the fact that there might be an inherited predisposition in families is if cancer is diagnosed at a young age, and secondly when you have more than one cancer diagnosed in a young age of the same type of cancer. So an example is early onset colorectal cancer in a family under the age of 45, so that’s quite unusual. You’ve got, essentially, defined consolations of specific cancers or site-specific cancers that are generally diagnosed at an early age.
Kat - We hear a lot about cancer in adults. Cancer in children is mercifully relatively rare. But is it the same kind of gene faults that are involved in childhood cancers compared to cancers in older people?
Richard - Yes, it is. They’re critically dependent on the early phase of development. So an example is in retinoblastoma, it’s the RB gene that’s important in the early phase of the development of the eye and then it doesn’t have a role. In fact, we know that people with retinoblastoma gene defects that are inherited actually have an increased risk of osteosarcoma in later life. So, effectively, the gene defect is having an effect in different contexts at different ages of the person’s life.
Kat - So, there are relatively rare ‘strong’ hereditary gene faults that significantly increase the risk of one or a few types of cancer in a family. Then there’s a much wider spectrum of genetic variations across the whole population, that have a less severe impact on cancer risk.
On top of this, our DNA is under assault every day - from things in the world around us, such as tobacco smoke, and even the fundamental processes of life at work within our cells. And it’s this damage, combined with our underlying genetic makeup, that can result in cancer developing.
Thanks to advances in DNA-reading technology, known as sequencing, scientists can now delve into the detailed genetic makeup of an individual patient’s cancer, and even use that information to shape treatment.
One example already in the clinic is the breast cancer drug Herceptin, designed to target an overactive protein molecule resulting from a genetic mistake in some types of breast cancer. But it doesn’t work for every patient - because it turns out that not all breast cancers are the same.
To find out how the genetic revolution is changing our understanding of cancer, I spoke to Dr Anguraj Sadanandam, who’s combining lab research with computer modelling to target tumours more effectively.
Anguraj - Conventionally, people were thinking that breast cancer, or colon, or pancreatic cancers were one disease. And that has been tried, and therapies have been given so far and it was not successful because different people respond differently to these drugs. But genomics is improving so much that we can untangle that and try to identify the sub-disease within each cancer type and try to match them to the therapies.
Kat - So, basically, all breast cancers are not the same even though they might all start in the same part of the body?
Anguraj - Yes. So though they start in the same part of the body, the question is are they starting from the same type of the cell - that is one interesting question. If they are not starting from the same type of cell then they will have a different prognosis and therapy responses.
Kat - So by finding the key gene faults that are driving these different groups of cancer cells, how then do you target those with treatments?
Anguraj - These key genes are later translated into proteins, and the proteins are the function units of the cells. These proteins have a particular structure and that structure defines how that cell would move, how that cell would change their shape, and all those things. So we can try to by, again, computational modelling and make certain chemical molecules which would go and bind at that particular structure. So that way the cells cannot get the signal to move or change shape so you know that you can control the cancer cells.
Kat - So these are very, very targeted therapies, targeting the faulty products of these genes in the cancer cells - wipe em out!
Anguraj - Yeah, so that’s the basic idea so that these cells don’t behave that way.
Kat - What do you see as the really big questions that still need to be answered when it comes to using genetics to understand tumours and how to treat them? What are still the big mysteries?
Anguraj - One of the major complications now - we are thinking that one patient do differ from one other patient but the much more complex thing is within a patient there are lot of subtypes and subgroups existing. Because, as I mentioned, if they are starting from the same cell within the tumour they can become into multiple groups. If you take one patient’s tumour and cut them into say 20 or 30 pieces, there are possibilities if they’re not going to twenty different ways but at least four ways, or five ways, or ten ways. That is a more complicated thing and we still are in a really early stage to understand how this change is happening. So if we go and start treating them thinking it’s one tumour, it’s not going to work.
Kat - I guess it’s like looking at large families and saying well, you’ve all got a lot of things in common but, when we really drill down to it, each member of that family is unique and individual.
Anguraj - Between two individuals within a family there may be a significant change to make them behave differently and also do things differently. Similarly the cancer cells can have very tiny changes, but those tiny changes can make them interact differently. At the same time, there are not only cancer cells within a tumour but there are also other cells, immune cells, all those interact and make them even much more complicated, and they behave differently and that’s a major challenge to untangle right now.
Kat - What do you want to find out next - what are you looking at now?
Anguraj - The major challenge right now is to define the key set of genes which are called biomarkers, because these biomarkers would define these subgroups. That’s the challenge and I’ll be thinking bowel cancer, for example, we have defined those genes.
The second thing is which type of technology that would take that to the clinic. The question now how to handle a lot of data given a patient coming into the clinic in a single day or two is very, very challenging. New methods and technologies are coming which will have to deal with thousands and thousands of genes, tens and hundreds of genes, so that we can give the answer to the patient within a day or two.
Kat - Advances in gene sequencing have the potential to target cancer in a more precise and personalised way than ever before, but this new wealth of data brings its own problems, as Nazneen Rahman explains.
Nazneen - So we’re at an extremely exciting time in genetics at the moment. We’re able to generate a vast amount of information about the genetics for an individual but our problem really is about curating that information, understanding its meaning, and there are lots of different ways in which we’re not able to use that information in the best way possible.
So one of the things that we’re currently doing, together with a lot of different people across the UK and indeed the world, is an initiative called the Transforming Genetic Medicine Initiative (the TGMI). Really what that’s trying to do is harness all of this information and use it to bring knowledge and, hopefully, wisdom so that we can make sure that we’re using the information appropriately to bring the maximum benefits. But also to make sure the information is not used inappropriately as well. So I think that’s the key challenge for us in genetic medicine currently.
Kat - How do you go about doing that; how are you bringing all this information together?
Naazneen - At many levels it’s really quite prosaic and a lot of it is logistically trying to pull together lots of different information and then curating it. There are 20,000 genes. At the moment it’s not entirely clear how many of those genes can be associated with human diseases, and how many are not. And, historically, there’s a lot of unclear information that was unclear before because we were having more and more knowledge, but you have back now and then look at the previous things and think oh, now we’ve got this new knowledge, how do we have to reinterpret previous information?
So, at the very highest level, we’d really like to go through all of the genes and say OK, these genes can be associated with human disease and these ones are not and be able to separate those out. For the ones that are associated with human disease: how are they, what diseases, what types of mutations? And then once we’ve sorted that out how does that then translate into how those are going to be tested and what are the best ways of doing that.
So at the heart they are really quite simple questions which, until now, we haven’t had enough data to be able to answer in a comprehensive and careful way.
Kat - It feels like there’s a lot of philosophical changes that need to happen. Instead of saying “you have breast cancer” to saying “you have this particular genetic subtype of breast cancer.” And then there needs to be a philosophical change in the way we develop drugs, the way we test drugs, the way we licence drugs. How can we make that happen?
Nazneen - Well, I think recognising the problem is a key step; debating it. I think the scale of technological change that allowed us to go from taking ten years to do one genome to being able to do ten genomes in one day, is such a massive change that it makes the whole system require a sort of ground up, just let’s think about it again and how we’re going to make it work in this modern world.
So I think the key thing is really recognising the issues and debating them. There are philosophical debates to be had, not least about the sort of predictive information as well. Do we want them, as individuals, in society? Do we want to have this window into the future. When do we want it; how do we want it; how are we going to act upon it? These are not things that we’ve had to think about very much so I think that’s the first thing that has to happen.
But then we have to then overlay that with some really quite pragmatic solutions. And I think we might have to do it in an almost experimental way: try things, get the information iteratively, then improve it. I don’t think we can solve it in isolation and create the perfect system ahead of time, not least because it is still changing very fast. So I would like to see some ways in which we can develop the system and improve it as we’re working through it otherwise we’ll always be chasing our tail, and we may never get there.

36:38 - Zooming in on micro-RNA
Zooming in on micro-RNA
with Peter Leedman, The University of Western Australia
As well as studying the genes involved in driving cancer and using this information to drive the development of new cancer drugs that target faulty proteins, researchers are also taking a more direct approach, investigating how to turn down the activity of genes themselves. One of them is Professor Peter Leedman from the Harry Perkins Institute of Medical Research at the University of Western Australia in Perth, who explained his ideas to Chris Smith...
Peter - We’re fascinated with the concept of using part of the genome. That is the DNA blueprint as a new therapy in cancer. And, in particular, some of the solid tumours like liver cancer, pancreatic cancer, head and neck cancer, and brain cancer, for which there are very few treatments, and the prognosis and the outlook may be very poor.
Chris - We know cancer is a genetic disease. It’s when genes go wrong and, therefore, cells begin to obey the wrong sets of instructions or disobey the normal sets of instructions. So when you’re say you’re trying to use the genome to come up with a new way to treat cancer, what do you mean?
Peter - Some of these solid tumours, they are driven by factors such as receptors that sit on the outside of the cell, and they drive the cells to proliferate or grow in an unrestrained way. We’re looking at that as the set of targets. The second part of the story is saying what can we discover inside the genome or the DNA blueprint that might give us an answer for slowing down that receptor which drives the growth. And the dogma for 50 years was that a lot of the DNA was said to contain junk, so called “junk DNA.” What’s been discovered over the last 15 years is that about 2% of the DNA is really important and made into protein, but that 98% of the non protein-making DNA is really important.
Chris - Just to clarify that a bit then. So you're saying that 2% of the DNA code actually are genes, and those genes turn into proteins in cells; they’re like recipes that make proteins. The 98% is really important but how? What is the 98% making then, or doing that makes it so important?
Peter - The 98% tends to make what we call non-coding RNA. So DNA is made into RNA - this copy - and it’s that copy that’s not turned into protein in the 98% of the genome, and it’s that that contains this rich tapestry of little molecules called RNA. And that is an area that’s of great interest globally because we believe that if we can untap some of the secrets in that 98% of the DNA, so called junk, that some of them might be useful for clinical treatments.
Chris - Right, so the dogma of for what’s useful about DNA is that it contains recipes that make proteins. That’s going out the window and what we now realise is that it might not be making proteins but this DNA sure as hell is making something important. It’s making these RNA molecules and what, do they stay in the cell and do something?
Peter - What we know: some of them are really small, only 22 or 23 bases long. What we found is that one of them called microRNA-7 is particularly powerful at slowing down and inhibiting the growth receptor that drives a number of these cancers, and it does it in a very sophisticated way. It basically takes out the key receptor itself and multiple friends of the receptor that help drive the growth and promote the tumour, and what we find when we add microRNA-7 to a number of these cancers, we can stop the growth of the cancer in the test tube. We can also stop it in mice and that’s particularly exciting.
Chris - What do they do physically inside the cell to achieve that effect? Do we understand chemically what they are doing?
Peter - What we know is that the microRNAs work inside the cell and they target the message and basically blow it apart. That protein being a key growth receptor that drives the cancer, that now is taken out of the cell and so the cancer undergoes what we call cell death.
Chris - So it’s a way of specifically, and in a highly targeted way, taking down a discrete genetic message so it robs the cell of the ability to have that genetic message that it was making too much of to make it at all? The only problem is that that would happen in every cell in your body, if you just deluged a person in this microRNA-7, they would be robbed of the ability to make that gene product everywhere. And that would be bad, wouldn’t it?
Peter - That’s a great question. What we know about these tumours, these nasty tumours such as liver cancer and head and neck cancer, for example, is they’re characterised by having lots, and I mean sometimes millions, of these growth promoting receptors on the surface of the cell. In contrast, normal cells which might have a thousand or much, much lower amounts and the principle here is to replace back something that the cancer cell used to make, and that’s a really important point.
The cancer cells we’re looking at used to make microRNA-7 and they’ve lost it and it provides this incredible break on growth promotion, and when you lose it you provide the opportunity for the cell to grow in an unrestrained way. Now, if we add back microRNA-7 it targets the growth receptor promoting pathway and all it’s friends in that, but it typically will target those cells in the body that have lots, and lots of the receptor. Much, much more than the normal cells, and so we believe that’s why it would be better tolerated and, of course, fight the cancer very effectively.
Chris - And you can get this message into the cell sufficiently, can you?
Peter - We can. There is, when we give the microRNA intravenously in model systems, it’s very well taken up into the cancer. So we believe this will be a wonderful way of treating human cancer.
Chris - One of the things that cancer is notorious for is it’s genetic diversity. And one of the reasons why cancers respond poorly in the end to chemotherapy is because, eventually, you get cancer cells that have evolved and been selected out, and they have an ability to rid the cell of whatever the agent is you're putting into it to try and kill it. So, is there a possibility that all you’re doing is kicking the can down the road and we’re going to select using your agent cancer cells that can just make even more of the gene for the receptor and you'll be back where you started?
Peter - Great question. One of the key things in cancers is they often develop resistance to the first therapy or therapies that they might be given. One of the things we’ve found with microRNA-7 is that when we give it to cells that have become resistant to a chemotherapy agent such as cisplatin, which is often used for head and neck cancer, we find that we can re-sensitize the cells to the chemotherapy agent by giving microRNA-7.
That’s really exciting because that suggests that we can rescue patients from the terrible traumas of resistance. And in real terms, when cancer kills people, it kills people because it comes back and it comes back in dreadful areas and spreads all over the body. You don’t die from the first time you get a cancer, you’d actually die from the complications of the cancer occurring - so called metastasis or metastatic disease. And we’re very excited about this molecule, microRNA-7, because of it’s ability to work with other agents and make the tumours resensitized to those agents and therefore, hopefully, improve the outcome.

44:27 - "Google Earth" for cancer
"Google Earth" for cancer
with Dr Josephine Bunch, The National Physical Laboratory
The charity Cancer Research UK recently announced the winners of four of its new Grand Challenge awards, which are multi-million-pound grants aimed at tackling some of the biggest questions in cancer today. One of the lucky recipients is Dr Josephine Bunch from the National Physical Laboratory in London, who’s leading an international consortium aiming to create a kind of ‘Google Earth’ for cancer, mapping the multitude of molecules that make up a tumour and eventually enabling researchers to literally zoom right into the heart of cancer. Kat Arney spoke to her.
Josephine - The analogy of a Google Earth view of cancer came originally from Cancer Research UK and one of their early blogs describing the problem. We’ve followed this through really and it works very well as a description of how you might overlay information and so if we liken this to the true Google Earth satellite imaging, this is a series of photographs taken at a range of magnifications that allows us, in the end, to zoom down into a particular location. We’re going to do exactly the same thing using our mass spectrometry imaging data and we also hope to go as far as being able to annotate these maps and make that information available for other people.
Kat - So people could, effectively, browse and go, “oh, I wonder what’s in here?”, and dig around in the same way you can on Google Earth?
Josephine - Yeah, that’s exactly right. At this stage it’s a little bit too early for us to know how much of that information will actually make browsable in that way. We don’t necessarily want to flood a database with lots of things which may not be useful but also, we don’t want to decide, at this stage, what could be useful. So one of the challenges for this is how do we store and make useful that kind of volume of data, and it’s not an easy problem.
Kat - Let’s zoom in a little bit into the kind of techniques that you’re using. How does this actually work?
Josephine - All of our imaging techniques are based on mass spectrometry. Mass spectrometers measure the molecular weight of something so really this is just a way of identifying the molecules in a sample. We move to a piece of tissue and we entice as many molecules out of that location as tissue as we can into our mass spectrometer where we’re able to identify them and with a move to the next location on tissue and do it again. So what we’re doing is building up an image piece by piece of as many molecules as we can extract from tissue at every location.
Kat - How do you weigh a molecule?
Josephine - We weight a molecule by creating an ion. So we start off with an uncharged molecule and we place upon it a charge. We do this by either giving it a proton usually, or maybe by losing an electron. Once it’s charged, we can then manipulate it and sort it according to its mass using a whole variety of different methods. This enables us to build up an understanding of which molecules were present at a different location.
If you imagine this is a little bit like when you go to the supermarket with an enormous amount of mixed change in your pockets and you throw it into a machine that helps sort it into its individual coins, the different types that you have, and we’re really doing that in our mass analyser. We’re creating charged ions, with all the molecules that are present, and then we’re sorting them very fast according to the mass to charge ratio to produce a mass spectrum, which is a kind of graph that gives us information about the relative amounts of different ions of different mass.
Kat - So you’re really mapping, in very fine detail, exactly where the molecules are in any sample of tissue, of cancer, of normal tissue?
Josephine - That’s exactly right and, importantly, we’re not deciding what we’re looking for first. The techniques that we’re using may not currently compete with the kind of image information spatially that you can derive from more standard optical techniques. So basic microscopy is really good at giving us a really well resolved image of the distribution of something. But it’s typically allowing us to look down a microscope at a stained tissue and that stain is showing us where particular molecules of interest are, but we have to have stained them. So, in our approach, we don’t know what we’re looking for. That’s the whole point, if we knew what we’re looking for we probably know an awful lot more about tumours and tumour biology.
So we’re taking a different approach and a much more unbiased, or unsupervised approach which is we don’t know which molecules we want to look for. So we’re going to use a variety of different mass spectrometers, useful for measuring all different kinds of molecules, many, many at once, and that will help us then to produce images of more things than we would be able to using a microscope.
Kat - I guess it’s moving from saying here’s a town, show me where the pubs and the bookshops are to here’s a town - what kind of establishments are here?
Josephine - Yeah, that’s exactly right. Afterwards, when we have that information, we can put back together a map that shows just where the pubs are, or a map that shows where the pubs are and the car parks are, or a map that just shows all of the buildings and we can start to work out patterns amongst them.
Kat - Ultimately, once you’ve built that people can zoom in and you can see all the different molecules at all these different levels, what then do you do with that information? How is this going to help us beat cancer.
Josephine - I think there’s a few different things that we hope this information will enable. Firstly, we hope that it will provide unprecedented information at the molecular level about tumour biology. So experts in this area will hopefully enormous wealths of information in our data and be able to better understand pathways or things that might be awrie in cancerous samples compared to more normal tissues. If we can understand some of these differences, then we might start to determine signatures, or even the presence of individual molecules, which could alert us to the presence of a tumour earlier than we currently can.
Kat - So that would be for earlier diagnosis?
Josephine - Exactly. This could be for earlier diagnostics and, hopefully, more accurate diagnostics as well. Then another layer of the ambition or the hope is that by better understanding of these changes at molecular level and better understanding of tumour biology, we can help develop more appropriate, more targeted medicines. Because I think part of the challenge of this project is it’s recognising the heterogeneity of tumours. Tumours are very different, different people’s tumours are very different and I think the more we can understand that, then the better and more personalised therapies that we can develop. If we can pair that with better and more personalised diagnostics, then we we might be able to treat this disease more effectively.
Kat - It does seem to me that it’s really only in recent years that technology, techniques that computing has accelerated to a point where we can do this. It must be incredibly exciting to think we can actually do this now?
Josephine - I know it’s fantastically exciting. It’s such an exciting time for this project and I think we’ve got to a stage where the capabilities of these instruments paired with, as you say, the advances in computational science have meant that we can really start mining this amount of information. I think that’s not to say that there aren't challenges. We still want to produce even more detailed images and we still wish we had better sensitivity and could measure even fewer molecules in a sample. And we’re still going to come up with real challenges as we try and mine this enormous amount of data.
To put it in perspective, one of our images might be 100 gigabytes, and we’re going to collect several of those a day across several sites for five years. So we’re talking about an enormous amount of new information.

The future of cancer treatment
Will we ever beat cancer, and what will the future of treatment be like? Kat Arney heard from Emma Smith from Cancer Research UK, about what the future of cancer and diagnosis might look like.
Emma - Imagine a kind of scenario where people were alerted to having cancer without even noticing any symptoms. Maybe they wear some kind of monitoring device like one of the watches that track various elements of your blood or your sleep patterns. What if there was some way that people could be alerted to a potential warning sign of cancer at a really, really early stage. Then go into the doctor and maybe the doctor could do a blood test and find out if it is cancer.
And, more importantly, if it is is it necessary to treat it? We automatically think cancers are a terrible thing and we should throw everything at it, but there are a lot of situations now where actually those cancers are never going to cause any harm so we shouldn't be treating people. It’s a very complex area but understanding those actual cases where we don’t need to treat is going to be very important as well.
Then once we’ve got a diagnosis, the doctor might be able to prescribe a really targeted treatment that doesn’t have many side effects, effectively wipes the cancer out. Maybe some immunotherapy using your body’s own immune system to go and find and attack cancer cells and then we have patients surviving for many, many happy years.
Kat - It all seems pretty exciting, but, as Emma also told me, there are still some big challenges ahead.
Emma - I think it’s fully understanding the causes of cancer. We still don’t have all the pieces of the puzzle. It’s finding ways to diagnose cancers earlier. We know if cancer is diagnosed at an early stage patients are much, much more likely to survive. We could save thousands of lives just by diagnosing the disease where it’s more treatable.
We need more effective treatments. We also need to learn how to use the treatments that we’ve got better; smarter. There are lots of potential combinations of treatments out there that simply have not’ been explored. Often these are cheap drugs that are already in use. We know that they’re safe, they’re easily available. How do we combine those with other modes of therapy like radiotherapy to make a massive difference?
Finally, of course, with so many people now surviving cancer, we need to not only be thinking about how effective a treatment us but also potential long term side effects. We’ve got now people surviving 10, 20 years post diagnosis. Even a whole life if it’s with children’s cancers. So what do we do to make sure that patients suffer the fewest side effects and have the best quality of life, not only during their treatment but in the many years that they hopefully have afterwards?










Comments
Add a comment