Cancer is literally the enemy within us -it starts when our own cells get damaged and go rogue, multiplying out of control and spreading around body. But how can we use new genetic knowledge to beat it? Plus, decoding the wheat genome, finding out where birds came from, and our gene of the month is more art than science.
In this episode

01:02 - Advances in cancer genetics
Advances in cancer genetics
with Dr Judy Garber, Dana Farber Cancer Institute
At its heart, cancer is a disease caused by faulty and damaged genes. It's usually a combination of the damage we pick up over a lifetime, as well as our own unique tapestry of genetic variation and - in some cases - specific gene faults that we inherit. To find out more about how faulty genes are involved in cancer, and how our understanding of them is helping to shape the cancer treatment of the future, I spoke to Dr Judy Garber from the Dana Farber Cancer Institute in Boston.
Judy - Well, it turns out that we have many genes in the body that have as their main job keeping the function of cells accurate and particularly making sure that when cells divide and make new cells that their genetic material is exactly the same as the parent cell, so that no mistakes are passed on. The genes that help make sure that cells can police themselves and make sure that they're correct are genes that must be functional for the cells to be accurate. When one of those genes doesn't do its job, then it's easier for mistakes to occur and for cancers to develop. So, the inherited piece of this is that in normal cells, we usually have two copies of each gene and both of them usually work just fine, but in people who have an inherited risk, one copy of the gene may not work so well. And usually, the cells are fine with just one working copy, but when something happens to knock out the working copy, some carcinogen exposure, radiation or chemicals, whatever it is that allows the normal copy to disappear, now the cell is more defenceless in that way and cancer can occur.
Kat - Now, what are some of the big hitters that we already know about, some of the genes that we know really do increase cancer risk if you inherit a dodgy copy?
Judy - Well, I think we've known for a long time that cancers can cluster in families. So we've known about the BRCA1 and BRCA2 genes that were found in the early '90s to explain a subset of breast cancer families. We know about a lot of genes that give familial risk of colon cancer and we probably don't recognise as often as we should that some of those families also get too much uterine or endometrial cancer, cancer of the womb, and ovarian cancer. And then unfortunately, we know some cancer syndromes that affect children and that's always the hardest, I think.
Kat - Now, what are we starting to discover about some, maybe less common or harder to find gene faults that are running in families and increasing cancer risk? What are the new things that are coming through?
Judy - So, I think it's been easier to find genes to explain rare cancers and very powerful genes that give an enormous amount of risk and would make anyone wonder about a family. But now, we're starting to find more information about genes that are a little less powerful and that are probably a little more common in the population. And we have a lot of work to do to be more exact about their effects on cancer. Some of these is happening because the technology has improved for analysing genes in general and so, we're finding more changes in genes that we know have a role to play in cancers, and therefore, also apparently in cancer susceptibility or cancer risk.
Kat - Now, we hear a lot about how cancer treatment is moving towards personalised medicine, so you take a sample of a tumour, you do a genetic analysis, and you say, "OK, you need this drug and you need this drug." How does that interplay on top of what might already be going on in someone's cells, just the kind of gene profile they've inherited because they're them?
Judy - So, there are many ways for that to work. One way people expect to be an issue is that some of the changes in the tumours started out as changes in some people in their original genes, and the tumours that they get have a particular biology because of their contribution of that original gene, and you find it by looking at the tumour genetics and then saying, "Well, wait. Let's look also at the person's own genes and get rid of the ones that were abnormal to begin with." So, that's one possibility. The other is and that because we're going to be looking at all these genes, we're going to learn about genes that affect very basic things about drugs we've known all along like the metabolism of drugs. So, why is it that two people can take the same dose of a medication and one has side effects and the other does not? So, it's probably nothing about the drug, but it may be the way the person handles that drug in their own body. And of course, that can be affected by lifestyle and other factors, but some of it is determined by genes that affect drug metabolism, so we should learn more about that too. Maybe that can help us avoid some side effects.
Kat - It's seeming to me that cancer is now as unique as we are. An individual's cancer is almost completely unique. That seems to me to present a huge challenge for doctors and scientists. What do you think are the challenges and how are we working to overcome them?
Judy - Well, I think that the challenges certainly are huge. Cancers are to some extent unique and they develop changes that are unique, but if they are using the same set of fundamental genes that drive them then hopefully, we'll be able to group them just differently instead of thinking about breast cancer, lung cancer, prostate cancer we may be looking for a group of cancers that have a particular gene at their base and the fact that they occurred in a different tissue may not matter quite as much as we use to think. But this requires a huge amount of analysis and a lot of thinking and, unfortunately, that's all pretty expensive, so there's some of that to contend with.
Kat - We're entering an era now where's just a huge amount of data coming out from genome sequencing, these big cancer genome projects, and things like that. And even people themselves, you can pay maybe around $1,000, 1,000 pounds and have your own genome sequenced. How do we try and get the grips with this kind of information and then what might people do with it if they find they have a risky gene?
Judy - The first question I think is fortunately, the province of people who love big data sets and they're out there and that...
Kat - The computer nerds.
Judy - Those computer nerds, they're finally going to really come in handy. And they can take this information and help sort through all of the noise to try to find the music, and I think that fortunately is what they're good at. The challenge will be to try to make the information available and to annotate it. And by that, I mean, that it's not enough just to know your genes. You have to know about the clinical impact and so, you're going to need information about the tumours and response to treatment, and family history, and things to give you context to interpret all the rest. So that's just more data, but I think that will be important.
Kat - And if someone say, has a test done either by a researcher, or pays for it, and then discovers they have a mutation, how do we know if it's actually important? What does it mean for the person or for their family?
Judy - Yes, well I hope that's where programmes like those that exist in the UK diffusely and elsewhere in the world, where people have tried to think about this 'if' issue. So, if you happen to have an alteration in a gene that's well-studied like the colon genes or breast cancer, then there's information to help you know what that might mean for you and for your children, your siblings, your family members. If it's a gene that hasn't been so well-studied, then perhaps you're someone who can contribute your information so that we can have research now give information to the next group to come along.
Kat - And do a little future looking for me, where do you think we're going to be, let's say, in 5 years' time in this kind of area?
Judy - Well, within 5 years, I think almost everyone will be having their tumour sequenced and that means that they will also have some of their own blood sequenced to try and understand the genetics, and for treatment that will be hopefully allowing us to use novel medications that are more effective and less toxic. And for our blood, it means we're going to learn more about our risk of cancer and other conditions. And we're going to have to think about how best to handle that information.
Kat - That was Judy Garber from the Dana-Farber Cancer Institute.

09:18 - Genetics Society Autumn Meeting
Genetics Society Autumn Meeting
with Professor Dirk-Jan de Koning
Kat - But now it's time to find out what happened at the Genetics Society Autumn Meeting, which was held at the Royal Society in November and celebrated 25 years of the scientific journal Genes and Development. Professor Steve West from the Cancer Research UK London Research Institute was awarded the annual Genetics Society medal, and gave a fascinating lecture looking back on his work on how damage to our DNA gets detected and repaired - a vital part of our cancer defences. We also heard talks from experts from around the globe, discussing the very latest progress in genetics research. To get the low-down and find out a bit more about the meeting, I spoke to Professor Dirk-Jan de Koning from the Swedish University for Agricultural Sciences, who's a member of the Genetics Society committee.
Dirk-Jan - So, the Genetic Society organises two meetings - a spring meeting and an autumn meeting and they are organised or planned by our members and they come with proposals. This one is a specific one because the journal Genes and Development was established 25 years ago with the Society as a co-founder and so this meeting was really about the science that is presented in that journal. So we have now seen two days of very detailed, cutting edge of the DNA and the RNA regulation and the consequences. If you look forward to the spring meeting, we have much more vertical meeting there where we look at how genomics has delivered for healthcare, and then we look much more from the science to the implementation in personalised medicine. So, the meetings vary in content and in topic, as well as in scientific depth.
Kat - And what are some of the themes that have been covered in the meeting over the past two days?
Dirk-Jan - So, if I take my personal highlights, it started yesterday very much with the DNA regulation, the DNA repair, how the whole cell machinery works to make sure all these processes that make us what we are, function correctly. And it moved on later in the day with the thing that brought it all together for me, was the medal lecture by Steve West where he really illustrated how some of the genes that we know as cancer genes like BRCA2 really, in their normal day job act as a chaperone protein that makes sure that DNA repair functions well. That for me was fascinating - to find out what all these suspect genes do in their normal day job.
Kat - I thought it was really interesting because he's a biochemist, but you've given him a Genetics Society award, and I thought his talk was excellent, really engaging.
Dirk-Jan - I fully agree, but again, Genetics Society medals are nominated by our members and then voted for, so it is clear that within the Genetics Society, someone like Steve West is very much appreciated for his contributions. So, we don't check someone's CV before we make sure whether they're eligible for an award. It is really...
Kat - Are you really a geneticist?
Dirk-Jan - Exactly. If you make a useful contribution, your colleagues or your peers will nominate you at some stage.
Kat - And finally, we've got the last afternoon of the meeting. What are you looking forward to this afternoon?
Dirk-Jan - I'm very much looking forward to Nick Hastie from Edinburgh who's a former medal winner of the Society, how he puts it all in the context of human diseases that we have sort of already given a glimpse from some of the earlier talks, so I'm really looking forward to that in particular.
Kat - And as well as listening to all the fantastic talks we've heard, it's obvious here that lots and lots of scientists, you're bringing them together from all over the world to talk and collaborate. Do you know that fruitful collaborations come from meetings like this?
Dirk-Jan - I don't have any examples at hand, but I do know there are a lot of interactions over dinner and discussions. You've also seen that we have very lively discussions on the basis of the talks, and I can assure you, those have continued over the dinners that we've had over the last 2 days. So, that's been fruitful from that perspective.
Kat - The next Genetics Society meeting will be in April 2013, and is on the topic of Genomics of Health and Society. If you're interested in coming along, there's more information on the Genetics Society website at genetics.org.uk
14:05 - Hunting for Huntington's genes
Hunting for Huntington's genes
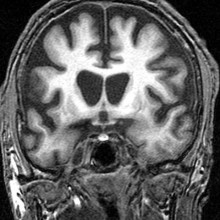
Writing in the journal PLoS Genetics, researchers in the US have developed a genetic "roadmap" that could speed up the search for new treatments for the currently incurable disease. Led by Robert Hughes, the researchers used a technique called RNA interference to systematically switch off nearly 8,000 genes in human cells, searching for genes that play an important role in the disease and affect its severity. This is the largest genome-wide search in human cells so far and has thrown up a number of exciting potential drug targets, including a gene called RRAS which they're now investigating. The researchers have also made all their data publicly available so researchers around the rodl can get stuck in and look at new targets for Huntington's Disease.
14:45 - Brain disorder gene found
Brain disorder gene found
Scientists led by Guntram Borck at the University of Ulm in Germany and David Adams at the Wellcome Trust Sanger Institute in Cambridge have tracked down the faulty gene responsible for a rare disorder that causes problems with brain development. Using a combination of genome sequencing from four patients with the disease and mouse models, the scientists discovered that mistakes in a gene called UBE3B are responsible for Blepharophimosis-Ptosis-Intellectual Disability Syndrome - the first time this gene has been linked to a human disease.
The UBE3B gene is involved in a process called ubiquitinylation, where enzymes inside cells stick little molecular tags on certain proteins, marking them out for destruction. Mice with faults in the gene show the same symptoms as human patients, with smaller bodies and brains, as well as problems with cholesterol metabolism. The research highlights the importance of using animal models in combination with human genetic studies for helping us to understand rare and complex human diseases. The scientists now hope to use their mouse model as a basis for developing therapies for the disease, to help sufferers in the future.
15:49 - Revealing the roots of pancreatic cancer
Revealing the roots of pancreatic cancer
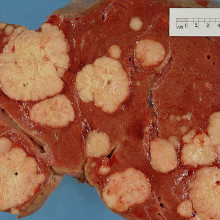
Writing in the journal Cancer Cell, scientists in California led by Maike Sander and Matthias Hebrok have uncovered the events at the very beginning of pancreatic cancer. Survival is currently very poor, and often the disease isn't diagnosed until it's too late to treat effectively. It was thought that pancreatic cancer started from lining, or epithelial, cells in the pancreatic ducts growing out of control, but this new research shows that these cells don't start growing in response to faults in genes that are known to be involved in driving pancreatic cancer. Instead, the new data suggest that it might be the acinar cells in the pancreas that lie at the heart of the disease. Inflammation, a known risk factor for pancreatic cancer, helps convert these cells into duct-like pre-cancerous cells in combination with overactivity of a gene called Sox9. The new discovery helps to shed light on the origins of pancreatic cancer, and significantly increases our understanding of how the disease starts and how we might tackle it.

16:45 - Wheat genome decoded
Wheat genome decoded
The genome of your breakfast toast - or at least the wheat that goes into it - is surprisingly complex. But now an international team of scientists from the UK, US and Germany have published the first analysis of the wheat genome, publishing it in the journal Nature. The researchers revealed that wheat has around 96,000 genes, and understanding what they do and how they work and interact will help to improve wheat crops as the world changes. The scientists hope their analysis will help speed up the development of new varieties of wheat that can cope better with disease, drought and other stresses that can cause crop failure. AS food security and climate change become ever more pressing issues, research like this is helping to put food on our plates - and bread in our toasters - in the future.

17:43 - The Cancer Genome Project
The Cancer Genome Project
with Dr Peter Campbell, Wellcome Trust Sanger Institute
Kat - But now it's time to delve a bit deeper into cancer genomes. At the Wellcome Trust Sanger Institute, Dr Peter Campbell is joint head of the Cancer Genome Project - an ambitious research programme aimed at analysing the genetic code in thousands of different tumours from patients around the world. I started by asking him what kind of gene changes were involved in cancer and where they come from.
Peter - Most of the gene changes that we're talking about that drive cancer accumulate during your life as you go through life basically. As we're walking around, we're exposed to things that are damaging our DNA. Cigarette smoke is a classic example. Sunlight is another example, and there are also processes that can happen within cells that mean that they can accumulate these genetic changes.
Kat - I remember hearing that just being alive and breathing oxygen is probably more damaging to your DNA than anything else.
Peter - I don't know about [more] than anything else, but it is true that these mutations occur pretty much throughout our life in some or other cells in our body, and if we lived long enough and didn't die of other things, then it would be statistically inevitable if we would eventually develop cancer, every one of us.
Kat - So, what are you trying to do with the cancer genome project to understand some of these gene faults that are involved in cancer?
Peter - The really exciting thing is that these genetic faults, these mutations that can cause cancer, they tend to be recurrent. So, if you sequence for example, some leukaemias, you will find that the same genetic changes will be found in multiple people, and those genetic changes are the ones that are driving the leukaemia. And that then acts as a natural target for diagnosis. We can use those genetic changes to diagnose someone's cancer accurately and also for therapy so that we can design treatments that specifically block the action of those drugs. What we're trying to do here at the cancer genome project is to - to do this is to characterise all of the genes involved in cancer in a systematic way. We take hundreds and thousands of patients with cancer and we sequence the entire genome of their cancers. And then by looking across different patients, we can look at those genes which are most frequently involved in a given cancer type, and then that's how we set up the targets for future diagnostics and treatment.
Kat - When you look at a cancer cell, often, its DNA is pretty messed up. There's a lot of things that are wrong with it, bits of DNA are stuck to other bits of DNA. There's all sorts of gene faults. How do you tell what's an important gene fault in a cancer cell and what's just there for the ride?
Peter - It's absolutely true. Cancers can have tens of thousands of genetic changes and mutations, and probably, actually only 10, 20, maybe a few more are actually responsible for turning that cell into a cancer cell. And the way that we identify those 10 or 20 amongst this morass of other things is to look across different patients. So, if we sequence enough different patients, the same genes keep cropping up as recurrently mutated, and that tells us that they're important for those particular cancers.
Kat - You talk about thousands of patients. This is only something that's been possible, thanks to recent advances in technology. How long now does it take to sequence a tumour genome?
Peter - When we started in the original human genome project which completed about 10 years ago, it took a number of institutes around the world about 10 years and many millions of pounds to sequence the original human genome. Now, we can do an entire cancer genome, plus the normal genome from that same person in about a week for £10,000. In the future, it's likely that we'll be doing this for less than a thousand pounds and in even shorter time than a week. So, the change has been absolutely remarkable. The really exciting thing is, that from a scientist's point of view, this is leading to all sorts of new and exciting insights into the way that cancers develop, and it's extraordinary time to be doing this kind of research.
Kat - Looking at some of the results that you have already for the cancer genome project, how many cancer genomes have you already looked at and has there been anything intriguing, surprising? What sort of things have you found so far?
Peter - I guess overall, with the latest sequencing technology, we have looked at, I guess several hundred patients with cancer. The kinds of things that we're finding are remarkable. We're finding all sorts of new cancer genes, so these are genes that are clearly driving some cancers. Some of them, we had no clue before that they were even remotely involved in cancer, and yet suddenly, these mutations crop up and then we've identified a whole new pathway that we can begin to think about - developing drugs to target. We've also found some remarkable processes that are actually causing these mutations in the cancers, so not just the genes which are being targeted by the mutations, but actually, what's causing the mutations in the first place. These are really remarkable. I mean, I think rather naively, going into this kind of study expected it to be rather consistent across patients, but it in fact is completely complex. There are all sorts of different processes going on in a number of different cancers. You can tell what a breast cancer will look like and it can look quite different to say a pancreatic cancer and have completely different patterns of mutations. And that's telling us something about how these genetic changes are occurring and that's just as remarkable as the discovery of new cancer genes in terms of the sort of scientific value.
Kat - Finding out all these things about cancer genomes, about how complex they are, about all the different faults and mutations, and different ways they can get messed up, does it make you feel optimistic or slightly pessimistic about whether we will beat cancer in the future?
Peter - I think beating cancer is an unattainable goal for all cancers in everybody. I don't think cancer will ever be cured for every patient that develops it. What I think we will find is that we'll continue to make iterative progress, so we will develop drugs that have more effectiveness. There'll be a fraction of patients who benefit tremendously from those patients. There will be more cures, but cancer will always be a devastating diagnosis to receive. It will always be quite challenging to treat, but when I hope is that in more patients, as we go forward, we understand their cancers better, and we can get treatments that are both better tolerated and/or effective, and we build momentum towards improving the lives of people with cancer.
Kat - That was Peter Campbell from the Wellcome Trust Sanger Institute.

How did birds evolve from dinosaurs?
Louise:: Hello and this month, I'm talking to Dr. David Norman from the Department of Earth Sciences at the University of Cambridge. Benoit in Bristol wanted to know how exactly did birds evolve from dinosaurs? Is there thought to have been a breakthrough mutation or did various kinds of birds evolve and intermixed bird-like characteristics in a more parallel process? Dr. Norman.
David:: That's a very complicated sort of question, but birds evolved from dinosaurs and if you look back at the history of the fossils that seem to go through the transition from being genuinely dinosaur and to genuinely bird-like you see a very, very gradual transition, changes in the anatomy of these creatures. In essence, what you find is that there's a trend within carnivorous dinosaurs, things known as theropods, which become progressively smaller, at least one branch of them. And that smaller branch changed its anatomy. One of the most important changes is the reduction of the tail. The tail becomes thin and very whippy, and that completely changes the balance of the animal and the way its legs work. As a result, they become very nimble, very delicate creatures that are capable of moving very rapidly. And if you link that with how they must've worked as biological organisms then if you build a little nippy creature then it has to be fuelled by a very powerful little engine, and that's the beginning of a change in the way in which the metabolism or the speed at which these animals work changes.
So the smaller you are, the more active you are, the more in a sense, dynamic, your general makeup. And once you get to that point, not only do you become more nimble and more agile, but you also need to be to some extent more intelligent. And you also suffer a problem which is that when you're very small, your surface area becomes very large. As a result, you can lose body heat to the environment very quickly, you need insulation. So, you end up with small, very nimble, very intelligent insulated - that is to say, feathered or filament-covered creatures that look awfully like birds. And honestly, the transition from those small, nimble carnivorous dinosaurs that probably ate small animals, maybe insects through to genuine birds which have an insulated covering and are capable of flapping flight is almost imperceptible. So, you go from one conventional dinosaur, through to a feathered avian or bird-like creature in a very, very smooth transition. It's actually quite a marvellous example of evolution.
Kat:: If you've got any questions about genes, DNA and genetics you'd like us to answer, just email them to genetics@thenakedscientists.com, tweet us @nakedgenetics or post on our Facebook page and we'll do our best to answer them for you.

28:10 - Gene of the Month - Van Gogh
Gene of the Month - Van Gogh
with Kat Arney
And finally, our gene of the month is all arty - it's Van Gogh. A gene found in fruit flies, Van Gogh was first discovered in 1998 by researchers at the University of Virginia. It helps cells to know which way is up - a type of gene known as a tissue polarity gene - and works together with the charmingly-named frizzled and prickle genes. Flies with mutations in Van Gogh have unusual swirly hair and bristle patterns, reminiscent of their namesake's brushwork if you squint a bit and use your imagination.
Van Gogh is also known as Strabismus in flies, and vertebrates including humans have two versions - VANGL1 and VANGL2. Faults in the human version of Van Gogh are linked to problems with developing a structure called the neural tube as a baby grows in the womb, leading to Spina Bifida, as well as being linked to certain cancers including liver cancer. More recent studies have found vertebrate Van Gogh popping up in many different roles in development, so it's definitely a gene that's more than just a pretty picture.










Comments
Add a comment