The gym is full, the pubs are empty - it can only be January, as a good proportion of the population resolves to shape up and lose weight. But are your efforts going to help you fit into your jeans (with a J), or are you just fighting against your genes (with a G)? Plus, we discuss how genes might jump between cows and snakes, and we've got gout, goats, giant pandas and a glass bottom boat. This is the Naked Genetics podcast for January 2013 with me, Dr Kat Arney, brought to you in association with The Genetics Society, online at genetics. org. uk.
In this episode

01:04 - Genetics of obesity and diabetes
Genetics of obesity and diabetes
with Dr Robert Scott, MRC Epidemiology Unit , Institute of Metabolic Science, Cambridge
Just looking at the families around us, it’s obvious that at least some aspect of our BMI - that’s body mass index, a handy if imperfect measure of weight - is encoded in our genes. This is borne out by genetic research, as well as the fairly obvious finding that a good chunk of our weight is down to our lifestyle. But what do we really know about how our genes influence our weight and metabolism, and - more importantly - our risk of metabolic diseases including type 2 diabetes. I spoke to Dr Robert Scott at the MRC Epidemiology Unit , based at the Institute of Metabolic Science in Cambridge, to find out how he’s hunting for genes involved in weight and disease, and that the answer may lie within the brain rather than the belly. I started by asking him how new genetic technology is helping the search...
Robert - So, it used to be the case in the past when we were doing these early genetic studies we would look at biological candidates and we would extract DNA from individuals, and we would genotype individual at known variants. So in the past, what we would’ve done was said, “Okay, this gene seems to be important in obesity or diabetes. Let’s look and compare individuals who are obese, compared to individuals who are not obese, and then look and see do the obese group carry more of this genetic variation than the other?” But what we can do now is we use this genome wide arrays where we can look at perhaps 1 or 2 million variants all at one time on this tiny microchip. So we extract DNA from individuals, we put the DNA onto this microchip, and it gives us information on what an individual’s genotype is at perhaps 2 million of these different genetic variants, all from the one experiment. So, we can then compare individuals who are obese versus non-obese, or individuals with and without diabetes, and look and see do the individuals with diabetes carry more particular genetic variants in individuals without diabetes.
The real approach that we’re taking now to identifying these genes for obesity, for metabolic disease, for diabetes is we take thes hypothesis free approaches where we measure genetic variance of millions of sites across the genome in lots and thousands of individuals, and then we performed a statistical test to see what are the genes that come out. We’re not driven by any particular hypothesis to say, “Okay, we’re only going to focus on genes which are involved in laying down fat or fat metabolism for example. We’ll just look at all the genes across the genome and see what comes out.” And what's really interesting is that most of the genes I think it’s fair to say, are not things which we would previously have said, “Okay, biologically, that looks interesting.”
Kat - The usual suspects.
Robert - Yeah, so they're not coming out with as the usual suspects as you see. They're coming out with things and giving us real insight into new biology. So perhaps then you might say, “Well, maybe we don’t know as much about the biology of obesity or diabetes as actually we thought we did” because we’re getting many new candidates, the function of which was previously unknown actually. So using these genetic approaches is giving us novel insight into biology of obesity, diabetes that we didn’t have before.
Kat - So, what sorts of genes are we talking about here that are involved in controlling things like weight and risk of diabetes, and other metabolic disease?
Robert - So, what's quite interesting is, when we do the experiments that we’re doing now, many of the genes which we’re coming up with are genes which have biological functions which we don’t really know actually, so we’re getting lots of novel insights. So, actually, what we found for BMI is that many of the genes that we’re identifying which are variants predisposing you to higher BMI are actually genes which seem to regulate your feeding behaviour, so they maybe make you have a preference for high fat foods or for high calorie foods and make you predisposed to want to eat more for example.
Kat - So, this is literally a gene almost for sweet tooth?
Robert - Well, yeah. One of the main genes for BMI or for risk of obesity seems to in the biological functions are being unravelled at the minute, but seems to give you a preference for high fat foods and what makes you want to eat these high fat foods.
Kat - It’s more of a fatty tooth rather than a sweet one.
Robert - Well perhaps, yeah.
Kat - And are you looking across a very wide section of society, because obviously, there's lots and lots of different people living in the UK with the different backgrounds, different genetic makeups. Do you find any interesting kinds of correlations between things like, say the risk of diabetes and other factors as well?
Robert - Well, that’s something we have to control for actually. So, for example, you're alluding to ethnic differences then one of the things you have to control for is this – what we term population stratification. So, exactly what you say. So, individuals for example, particular ethnic groups are predisposed to diabetes, but what you have to really be careful of is how you design your experiment because if of course you take individuals with diabetes, compare them to a control population without diabetes, but all of your individuals with diabetes are, for example, of south Asian descent (a group predisposed to developing diabetes) and all of your controls are of European descent, then what you're likely to find is not necessarily genes which are associated with diabetes, but genes or gene variants which are associated with your ethnic background.
What's ongoing at the moment is lots of multi-ethnic studies. We’re really looking in European individuals and individuals of south Asian descent, of African ancestry. What's really nice is that if you see a signal, a gene variant which is associated with diabetes in Europeans, if you then take that and see the same signal in individuals of African ancestry or South Asian ancestry, then it gives you real insight that these signals caused by something which is real and causal, rather than something which is a chance association. So, replicating things across multiple populations is really important actually.
Kat - How do you just marshal all this information and actually extract meaningful patterns from it? It’s a huge amount of data.
Robert - It really is a huge amount of data. So for example, in the standard experiments, we have for every individual, genotype information at 2 and a half million different mutations or polymorphisms as we term them (changes in the DNA sequence across individuals). Some of the more recent studies we’ve been doing, we’ve had up to quarter of a million individuals, so you could imagine 250,000 people with two and a half million different genetic variants, that's a whole lot of data that we have to handle. And that means when we're mining through all this information and trying to find out what's really important, we have to be statistically very rigorous.
So in a regular experiment what you say is, you perform your statistical test, you find your result, and then you compare that result to the chance of finding that result simply by chance. And normally, what one would see is, you say, “Okay, we’ll accept a 1 in 20 probability that this result is due to chance.” And we say, “Okay, there's a 95% likelihood it's not due to chance.” If that’s the case then we think this might be a real result. But if you're performing two and half million statistical tests then a whole lot of them, 100,000 or more, are going to meet that threshold based on chance alone, so these are not real. So of course, we can't then say, “Okay, 100,000 things look real.” What we have to do is be very rigorous and say, “Okay, we’re not going to accept this 1 in 20 threshold any more. We’re going to go to let’s say, 1 in a million chance.” So, things have to have a very defined, not necessarily a large effect, but a very precise and defined association with the disease, a very highly, statistically significant threshold for us to consider it real.
Kat - You're talking about a lot of people here. Where do you find them from? Where do you recruit all these people?
Robert - We’re analysing together lots of individual studies, so this is not one particular study of obesity in quarter of a million individuals, but what we do is we go across the world and we say, “Okay…” So for example, here and Cambridge, we have a number of studies, looking at the risk factors for obesity and diabetes, and we have studied perhaps 10,000 individuals. But what we can do is we collaborate with centres across the UK, across Europe, across the US. We bring together all these different studies, so each individual study will perform their analysis, and then we perform this method of data amalgamation to bring everything together, and tease out what's real and what's interesting.
Kat - You've looked through all these information, you found correlations with variations in genes that are important for the risk of obesity, for the risk of metabolic diseases. What are you going to do with this information now? How do we make this useful?
Robert - So, one of the things when the human genome project was kicking off was the belief that you could take DNA from everyone, measure your risk of disease and then you could say, “Okay, you're likely to develop obesity, diabetes and suchlike.” Actually, things have really turned around and really – and that was probably never likely to happen I think. What really we’re getting and the main benefit for doing this genetic studies is not necessarily to predict to become obese or diabetic, but like I've just mentioned to you, it gives us real insight into the biology of these diseases where we maybe had a prior belief that what’s important in obesity is how your body metabolises the food that you eat.
These new genes that we’re finding are telling us, “Well, okay that may well be important.” We have hypotheses that that’s important, but actually, people’s preference for particular foods are important, so it gives you novel insight into the biology of obesity. Similarly for diabetes, many of the genes we’re finding for diabetes that are involved in particular processes in the aetiology of diabetes and that perhaps not now maybe, but in 10 years and 20 years gives you novel insight into therapeutic targets.
Kat - Have there been any findings that have been very intriguing or things that you found particularly fascinating?
Robert - I think it’s quite easy for individuals like myself over Christmas will indulge and eat too many mince pies and we say “Well, I have genes which are making me eat too much.” Then one has to be careful not to ascribe everything to genes and I have responsibility to look after myself. So, one of the interesting things which has been found in studies previously is that while carrying particular genetic variants increases your BMI, what's been found previously is that if you compare the effect of these genetic variants amongst individuals who are physically active, who get a lot of exercise, who walk a lot, to individuals who are inactive, who don’t get as much exercise or physical activity as perhaps they should, then the effect of these genetic variants actually appears to be bigger in individuals who are physically inactive.
So while having, carrying particular genetic variants might increase your level of BMI, the really important public health message is that, if you're physically active you can perhaps offset some of that genetic risk. So being physically active and being careful about your diet is actually very important, despite perhaps you carrying an increased genetic risk.
Kat - We’re just moving so far away from the idea that, well, if you're overweight, it’s because you’ve eaten too many pies and you don’t do enough sport to a much more sophisticated and subtle idea of how our bodies work.
Robert - At it's very simplest level, if one is to eat less then exercise more, then of course, you can reduce your risk of obesity and your BMI will be lower. But I think it’s very fair to say that it’s very complex. Actually, it’s quite easy for people to ascribe being overweight to simply not doing enough exercise and eating way too much, but what are the causes of eating too much? Why are some individuals driven to eat more than others? And some of the differences, the inter-individual differences in how we do these things are likely to be done to genetic variation. So I think it’s a very complex story.
Kat - That was Robert Scott, from the MRC Epidemiology Unit in Cambridge.

12:42 - Genes jump from snakes to cows
Genes jump from snakes to cows
with Nell Barrie
Kat - But first, it's time to take a look at the top stories from this month with science writer Nell Barrie. What have you got for us this month, Nell?
Nell - So, we've got one study that is very interesting and it's looking at how a quarter of the cow genome actually came from snakes or is this really true? That's the question. This is in the Proceedings of the National Academy of Sciences by Ali Morton Walsh and her team at the University of Adelaide.
Kat - Now, this is a bit crazy because cows and snakes aren't really related, so what is this? What is this bit of their genome?
Nell - Exactly, so you kind of immediately look at this and go, "What? How on earth could that be possible? That's ridiculous!" And it's looking at a piece of DNA which is called Bovine B or BovB sequences. And what they found is that about a quarter of the cow DNA sequence is made up of this BovB. The really interesting thing is it's not just in cows. It's in lots of other animals as well and the question is, how did it get there?
Kat - Because they thought this was something really distinct for cows. They have this repetitive thing that's for cows.
Nell - Exactly, and the idea would be that if you've got this repetitive sequence in cows, if you look in cows' relatives, closer relatives of the cows should have similar levels of this repetitive sequence, but that wasn't what the researchers found.
Kat - What other sorts of animals have got this thing?
Nell - So, there actually loads of different animals and if you look at the kind of family tree of this gene sequence, you find all these really odd things where it seems like cows are actually closer to snakes than they are to elephants, and there's a gecko that seems to be very closely linked to horses with the amount of this DNA sequence, much more closely than it is to other lizards. So, you're looking at this thinking, this just can't be possible. These results must be wrong. It's really weird.
Kat - So, either Darwin was completely wrong, evolution isn't true, or there's something else. What do they think is going on here? This is nuts!
Nell - Well, we think Darwin wasn't wrong, so that's a relief at least, but what it appears to be is that these are what are called jumping genes. So they're genes that can literally take themselves into the genome, copy themselves all around. They're really, really good at doing this but the question is, how have they got between these different species and the answer could be, something as simple as a tick. So parasitic insects that can actually go between say, a lizard and a cow, and transfer these genes in that way, so it's pretty crazy stuff.
Kat - This sounds pretty controversial to me.
Nell - Well, yeah that's exactly right and I mean certainly, it's not what you'd assume from these results. I mean, I guess the logical thing you'd say is something has gone wrong with these results somewhere and of course, it's always very difficult to do this kind of sequencing.
So, if you're thinking of people who work in a lab, all these different DNA sequences, there's always that possibility of contamination. But the researchers were very, very careful about this because obviously, they recognise that was a huge possible problem. So, it seems that that wasn't the case, but we are going to need some more work to find out really what's going on, and how exactly these genes manage to transfer themselves and spread so quickly. So, it seems like that would be a really rare event for this kind of tissue to sneak through somehow, but clearly, something weird is going on, so it's very interesting.
Kat - Or maybe it's really common and we just haven't found any other ones yet.
Nell - That could also be the case and we know that this does happen a lot, so there's this thing called horizontal gene transfer which happens a lot in bacteria. But, it seems like it could happen in much larger animals as well, so that is really fascinating.

16:03 - Panda Genome
Panda Genome
with Nell Barrie
Kat - Now for some other fascinating animals - pandas. I saw in the latest issue of Nature Genetics from Shancen Zhao that they have sequenced the panda genome. What did they find in it?
Nell - So, this was really looking at how different populations of pandas are different in their genetics because we know that there's many different populations of pandas in China that are quite isolated from one another because of habitat changes, all that kind of thing. And what they found is that these different populations are actually evolving differently which is quite cool, so it's showing kind of real-life evolution happening now.
Kat - I thought one of the interesting things was they've identified evolutionary bottle necks, and all sorts of things, but they managed to show that it looks like human activities have had an effect on the pandas in over the past 3,000 years or so.
Nell - Exactly, yes. So it's looking at kind of how the habitats have changed, where the pandas are living and there's one nice example where some of the panda's taste receptors are actually evolving because of the type of food that it's eating in the environment that it's living in, so that's quite nice. And they even get in the nice phrase, 'The panda eats shoots and leaves' which I enjoyed for grammar pedants out there. I liked that one.
Kat - The other bit I really liked in the report is that they sequenced the genomes of 34 pandas and that's actually 1 in 50 wild pandas which is a recruitment rate to their study which is absolutely phenomenal.
Nell - Exactly, yeah so they say that's something that human geneticists would be really, really amazed and then they could only dream of recruitment rates like that. But, pandas obviously weren't consenting to this in the same way that humans would be, so probably a little bit easier.
Kat - And they're very cute as well. Thanks very much, Nell. That's Nell Barrie, science writer.
17:39 - Cancer-killing cells created
Cancer-killing cells created
Researchers in Japan have created killer immune cells that can be grown in the lab and recognise melanoma skin cancer.
Writing in the journal Cell Stem Cell, Hiroshi  Kawamoto and his team used so-called 'Yamanaka factors', named after the recent Nobel prizes-winner who discovered them, to turn killer T cells into immortal immune stem cells.
Kawamoto and his team used so-called 'Yamanaka factors', named after the recent Nobel prizes-winner who discovered them, to turn killer T cells into immortal immune stem cells.
These stem cells were then grown in the lab to create a large number of cells, then turned back into T cells. Importantly, the original T cells had previously been trained to recognise melanoma cells, and they retained this memory even after the stem cell process.
The researchers suggest this could be a promising way to treat cancer - taking immune cells from a patient that can recognise their cancer, turning them into stem cells and growing them up in the lab to create millions of killer cells that can be transplanted back into the patient to destroy their disease.
Another team in Japan used the same technique to generate T cells that could destroy HIV. Despite the headlines in some papers, at the moment the scientists haven't yet shown that these reprogrammed T cells can actually destroy tumour cells or combat HIV in mice, let alone humans. But it's certainly an exciting new approach using an important new technology.
18:56 - Gout risk genes found
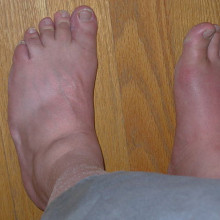
Gout risk genes found
Writing in the journal Nature Genetics, an international team of researchers have found 18 new genetic variations that increase the levels of uric acid in the blood - the main cause of gout. The disease is caused by tiny crystals of uric acid forming in the joints, causing pain and swelling, and the number of cases of gout is on the rise thanks to increasing age and obesity, affecting around one in 70 UK adults.
The scientists studied DNA from more than 140,000 people, combining more than 70 studies from around the world, looking for genetic variations that were linked to higher uric acid levels. The team hopes that their findings could translate into better understanding of the risks and causes of gout, helping to spot people at risk and prevent the disease as well as leading to more effective treatments with fewer side effects.

19:44 - Goat genome mapped
Goat genome mapped
Goats are an important animal around the world, especially in countries such as China and India. They're bred for meat, milk and hair - cashmere, the prized downy hair from certain species, has been collected for more than 2,500 years. Now a Chinese team have used the latest sequencing techniques to complete the first goat genome, mapping the entire genetic code of a female Yunnan black goat - a common domestic species - publishing their work in the journal Nature Biotechnology.
In particular, the scientists focused on the differences in gene activity between the two types of hair follicle in the goat's skin - the primary follicles that make coarse hair, and the secondary follicles that make luxurious cashmere. They found differences in activity between 51 genes, including in genes making keratin proteins - the main component of hair.
The new goat genome is an important asset for researchers and livestock breeders around the world, and the scientists hope it will help lead to more effective breeding and better quality cashmere.

20:50 - Meet the Rinkidinks
Meet the Rinkidinks
with Dr Tiffany Taylor, University of Reading
Tiffany - I wrote this book because I thought that currently the way that evolution is taught, as a starting point is it's taught as quite a static process. Organisms have differences because they live in different environments, but it doesn't really describe how organisms from one environment can move into another environment, and then adapt to this new environment to take on the changes which make them a better fit. What I wanted to do is write a book which is very simple aimed at young children which told the story of these little characters called the Rinkidinks, involved in a natural disaster which splits them between two very different environments which over time they have to adapt to. And then one day, they bump into each other in the forest and they don't recognise but little did they know that they actually come from exactly the same background. So, I wanted to get the idea of how it's a dynamic process that takes a lot of time rather than it being very static.
Kat - And so, you've managed to get quite a lot of what we understand about evolution, how it works into it. How did you find writing it in such simple terms?
Tiffany - It was difficult, but actually, evolutionary processes follow quite simple laws, and really, all you need is reproduction, variation and selection, and without using those terms actually, I think it's something that children can visualise quite easily. I mean, we're all different, we're all unique, that's the variation. Terrible things, natural disasters, you can see that that can cause changes in the environment, that if they want to reach their food then they've got to have certain changes which are going to help them do that. So, I think the concepts themselves are something that children can relate to and you don't need this complex terminology in order to explain it.
Kat - And what are your hopes for the book?
Tiffany - Well, I really hope that primary school teachers can use it as a resource, in order to help perhaps them understand evolutionary processes a bit better because I mean, a lot of them haven't had any sort of training, any sort of scientific training, and give them also classroom activities which will help solidify these new concepts which are going to be introduced to the children and perhaps, even open up some forums for which they can ask questions, or children can ask questions about evolution, the things that perhaps the teachers feel that they can't necessarily answer that comfortably. So, I just really want it to be a resource for teachers to explain the concepts of evolution, giving children a better grounding so that when it's re-introduced at a later stage and key stages, that they are coming from a more solid foundation.
Kat - That was Tiffany Taylor, from the University of Reading. And you can find out more about Little Changes, including downloadable activities for kids and resources for teachers, at
http://www.rinkidinks.co.uk/

How can we find "cancer genes"?
Answered by Dr Julie Sharp from Cancer Research UK
Julie - When scientists are trying to crack the code of a particular cancer, they look at all the different genes from an individual's healthy cells and then their cancer cells, and they make comparisons between them. This gives them a huge map of all the different genes that are faulty in that particular cancer. They then have to sift through all of these information because there are many, many different faulty genes, but some of them will be the really important genes, the ones that are really causing and driving the cancer, helping it to grow and spread. Some of the faults are just picked up because the cell's in disarray, there are things going wrong, and some of them aren't really, really important.
So, the thing they have to do is then to, from this great amount of information they've discovered, actually work out which are the key driving faults, which are the you know, really important ones. And there isn't just one combination of gene faults and bingo! You've got cancer. There are lots of different possible combinations of gene faults. You can have a number of faults before your cells develop cancer. But there are genes that are commonly faulty in a particular cancer, so for example, the gene BRAF is quite often faulty in melanoma skin cancer, but then it will be faulty along with different other genes and those range of genes will differ between different people.
The genes that lead to cancer when they're faulty, it's stopping them from doing their normal function and some of these genes will normally be involved in things like helping cells to grow and develop. That means when they're faulty, cells will start growing and developing out of control. And in other cases, genes are linked to processes that would normally protect us from cancer. So they're the checks and balances in the cell, when they're faulty, that just means those safeguards aren't there and things can develop out of control.
And finally, there's a real spectrum of different types of faults that lead to cancer. So, we know that a minority of cases, people will inherit this really high risk genes that are passed on through families, and that means you're at greater risk of developing cancer - that's the genes like the BRCA genes. They don't mean you will get cancer, but you have a much greater chance of developing the disease. Most cases of cancer are caused by the faults that we develop through our lifetime, just through the ageing process, through wear and tear in the cell, and through the effect of lifestyle.
So, whether we've smoked, whether we've laid out in the sun and got sunburn, all of those things play a part. What we're gradually finding out more about is that there are also some genes that are passed on through families that have a smaller effect on our risk and so, it's really a spectrum. In most cases, it's what we've done during our lifetime, but there will be things that have been passed on through our families that will have had an effect.

27:51 - Gene of the month - Glass bottom boat
Gene of the month - Glass bottom boat
with Kat Arney
It's time to take a voyage in our gene of the month - it's glass bottom boat, or gbb. The name comes from fruit flies - fruit fly larvae with a faulty glass bottom boat gene are transparent, compared to cloudier normal maggots. There's another tie-in too - when researchers looked at the activity pattern of glass bottom boat, or gbb, in developing flies, they found it was a mirror image of the activity pattern of another gene, decapentaplegic, or dpp. And the letters gbb are a mirror image of dpp - clever, huh? Also known as the less-catchy TGFbeta at 60A, glass bottom boat has a number of roles in fruit flies, including helping to organise the developing fly embryo, forming reproductive cells in the testis, patterning the fly's wings, and regulating the junctions (synapses) between nerve cells, which it does by interacting with a protein produced by the wonderfully-named wishful thinking gene.
There's no exact copy of glass bottom boat in mammals, including humans, but we do have a couple of genes that are related, which are both involved in growing bones. In fact, putting the fly glass bottom boat gene into rats can create new bones, which is pretty amazing when you think that rats and fruit flies are separated by millions of years of evolution.










Comments
Add a comment