RNA Vaccines, Privacy, and Penguins
The first group of people in the world have received a 'genetic' vaccine against the coronavirus. What is it, and how does it work? Naked Scientist Chris Smith breaks it down and addresses your concerns. Plus, why some genes have to change rapidly just to stay the same; a new way to keep functional genetic information private; and three new species of penguin arrive on the scene…
In this episode
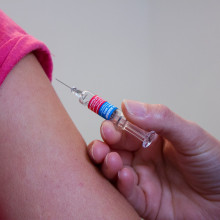
00:32 - COVID vaccine: is it safe?
COVID vaccine: is it safe?
Chris Smith, the Naked Scientists
The first authorised vaccine for the coronavirus has begun its rollout. Produced by companies Pfizer and BioNTech, it’s what’s known as an mRNA vaccine, just like another one that has passed phase 3 clinical trials: Moderna’s. With other countries close behind, we can expect many hundreds of thousands of people to have received it before 2020 is out. But a sizeable number are nervous about getting the jab. So Phil Sansom asked virologist and Naked Scientist Chris Smith to explain what an mRNA vaccine actually is…
Chris - RNA is a chemical relative of DNA; so it's a form of genetic information. And in our bodies, it is used to carry a copy of the gene that you want to use, out to the part of the cell that can decode that gene and turn it into a protein; something the cell can actually do something with. So it's a short-lived message that moves something from a gene to becoming a protein. But some forms of life, and specifically some viruses, rather than using DNA like we do, they use this form of genetic material - RNA - as their main information storage mechanism.
Phil - What RNA message then is this vaccine RNA carrying?
Chris - What they've done is to go to the virus genetic code and find the piece of that code that corresponds to the outer coat of the virus, and specifically a structure called the S protein, or spike - S for spike - protein. They've taken just that short piece of the RNA from the virus, and they've put that into a package, so it's effectively wrapped up in an oily coat. So when injected, it's picked up by cells, unwrapped; and because cells understand genetic material like that piece of RNA, they can decode the recipe on there and make the protein that it would have made if the virus was in the cell for real, and then display that protein to the immune system, showing the immune system what a cell that would be infected with coronavirus would make.
Phil - What happens to the little bits of RNA? Where do they go?
Chris - The thing about RNA is that it is a very transitory thing. It has a really short lifetime in cells, and it very quickly is degraded. There are mechanisms there whose job it is to eat it and break it apart. So when you put the vaccine mRNA into the cell, it too is terminated very quickly, which is why you need to have a big dose of the vaccine; and not just once, but twice.
Phil - Now am I right this is the first time that this tactic's really been used?
Chris - People have made genetic vaccines in the past, but they've never used them in humans. They've been used, tested and deployed successfully in animals - most recently actually for Zika virus - but this had not been done in humans. And so now the know-how from those earlier experiments which we've been learning about for a couple of decades, actually, have now for the first time been successfully deployed in humans to combat coronavirus.
Phil - What are the risks here? Because you'd expect any medical treatment come with inbuilt risks or side effects.
Chris - The risks range from the very trivial to the more serious. At the very trivial end of the spectrum, all medicines have side effects. This vaccine series is no exception. Those side effects usually are things like pain at the injection site; you're sticking a needle into somebody, it will be a bit uncomfortable. Slightly more serious: a handful of people might get some trivial side effects. By 'trivial' we mean things like: you might feel a bit fluey for a day or so. This is actually to be expected; it's because the immune system is responding to what's come into the body - that's the whole point of doing this after all - and when you have an immune response you produce various immune signals, and you feel unwell for a day or so, because that is the process that's happening when the immune system is being stimulated. The more severe end of the spectrum: very, very rarely, with some medicines you get what are called idiosyncratic reactions; and this is where, because every single person on earth, unless you have a twin or you've cloned yourself... everybody's genetically distinct, therefore they're biochemically distinct on the inside. And as a consequence of that, there is a small chance that in a very rare circumstances, some people may just overreact to some medicine or react in an unpredictable way. It's very, very rare; this is what clinical trials are designed to screen out; and so you don't license things that you think there's an appreciable chance of this happening.
Phil - What if you have some sort of autoimmune condition? Does that make a difference?
Chris - Autoimmune conditions are where the immune system attacks your own tissues. And there are lots of control mechanisms in place to stop the power of the immune system being turned on you, because it can be very destructive. The reason that this may be a problem with vaccination is that when we have a case of an autoimmune disease, it's controlled by suppressing the immune system with various drugs. These are a bit of a blunderbuss thing; they turn down all of the immune system. So when you do give people vaccines, they might not make the same robust, resilient immune response that someone would make were they to have a normally functioning immune system. It's not a given that they won't, but it's a possibility that they won't respond as well as someone with a healthy immune system. So it can be a consideration. Also, if people have an immune system that doesn't quite work properly for a range of reasons - there are inherited reasons, there are some acquired reasons, some diseases that could do this - they may overreact to certain things. But those sorts of situations are very rare.
Phil - Some people are sort of understandably hesitant, because they see a vaccine that's been developed, to anyone's eyes, remarkably quickly. Is it possible that there are unknown unknowns here?
Chris - There are always unknown unknowns in everything that we do. And unfortunately, inherent to everything there is some degree of risk, and that's a fact of life. And there is no way of getting around that. But what you do have to do is to take the greatest steps that you can to minimise that risk, to the greatest extent that you can. And that has been done here. Because what they have done is to take decades of learning about vaccines, and about actually making vaccines the way that these ones have been made; you then do trials on people to test the safety of them; you then put in place post marketing trials, so follow-up trials, so that you follow the people and you have a system in place to look for adverse outcomes. And at the same time, the approvals process has been streamlined for this new vaccine. Now what that means is to have a two way conversation between the drug manufacturers and developers, and the regulators, all the time, so that many of these things have been speeded up enormously so that there are no delays. What they haven't done is to cut the corners in the scrutiny, and they certainly haven't lowered the threshold for what that they'll accept as safe, because number one on the list is 'safe'. What we can never know is what the long term effect is. We don't have a time machine or a crystal ball; you can't see into the future. So the question we'd most like answered is: how long will I remain immune for? And we have no idea. But what we do know is: we're going to find out, and we'll find out by following up the people that get vaccinated.

08:23 - New, evolving genes sometimes have vital roles
New, evolving genes sometimes have vital roles
Harmit Malik, Fred Hutchinson Cancer Research Centre
One loose principle in genetics is that the oldest genes in our bodies, the ones that haven’t changed for millenia, must be like that because they do something critical for our survival. If they changed, we would die, simple as that. Whereas anything that is changing must be non-essential. Harmit Malik’s research overturns this principle, based on genes in fruit flies, and the reason why appears to be that the genes are constantly running on a treadmill; changing to effectively stay the same. Phil Sansom asked Harmit to explain...
Harmit - The dogma in the field has been for a very long time that the more conserved a gene has been in evolution, the more likely it is to encode an essential function. My lab was interested in asking what happens at the other end of the spectrum, where genes that are apparently not very well conserved: how likely is it that they encode essential functions? To address this we actually focused on one category of genes that we already knew showed some diversity. These are called ZAD-ZNF, which are the largest class of transcription factors required to turn on genes or turn off genes in a very regulated fashion in insect genomes. And we were surprised to make two discoveries. Discovery number one was that ZAD-ZNF genes that were not strictly retained were just as likely to encode an essential function. The second, more surprising finding was that ZAD-ZNF genes that were actually quite slow to evolve were actually less likely to encode essential functions.
Phil - So these genes, these ZAD-ZNF genes: a lot of them are pretty new, and despite that they're still coding for really important things; and even some of the ones that are evolving the most quickly are doing the most important stuff.
Harmit - That's exactly right Phil.
Phil - How is that possible? If they're doing such important jobs, how come they're so new?
Harmit - We were exactly puzzled by the same question, Phil. How is it that the genes that were very rapidly evolving were actually more likely to encode these essential functions? So to take a closer look at these genes, we focused on two of the dozen genes called Nicknack and Oddjob. These were genes that were named with a sort of inside joke, because these are both referring to James Bond henchmen.
Phil - Are these like henchman genes?
Harmit - Oddjob was named partly because it is a fairly odd gene, in the sense that typically if you have a transcription factor, you expect it to be localised to where all of the action is, where all of the genes are. Instead, Oddjob appeared to be localising to this essentially unmapped part of the genome which is really devoid of genes, for the most part. We refer to this as 'heterochromatin', which literally stands for 'other chromatin'. And we were really surprised to find that Oddjob and Nicknack do not localise to the gene rich, but instead to the gene poor heterochromatin part of the nucleus.
Phil - What is the gene poor part doing that is affecting the essential jobs, that are presumably part of the gene rich part, really?
Harmit - About 25 years ago, I would have told you that we know almost nothing about the gene poor part of the genome. More and more we are actually recognising that heterochromatin actually plays very important roles within the cell. Regions in the heterochromatin actually help regulate all of the other genes. So you could even imagine that they're sort of master puppeteers of the rest of the genome. It's actually becoming clear that the heterochromatin is just as important, if not a more important part of the cell.
Phil - That doesn't quite explain to me how they're in this fast evolving state, where you've got new genes coming up and the genes are changing so much.
Harmit - Yeah, so one of the really cool things about heterochromatin is because it's actually made up of repetitive elements, these elements are extremely different as you compare them between species or even between different members of the same species. So the paradox is really that they're able to carry on this master puppeteer function, but they're not actually doing so with exactly the same conserved DNA sequence. And therein I think lies the resolution of the paradox of Oddjob and Nicknack. In a way they're sort of acting as this buffer or mediator, to take all of this churn at the DNA level, and yet ensure that the conserved functions of heterochromatin are conserved over billions of years. So you have simultaneously this rapidly evolving, almost a competitor part of the genome, and yet you're dependent on this competitor for your essential function.

13:11 - Data sanitisation tool plugs privacy gap
Data sanitisation tool plugs privacy gap
Mark Gerstein, Yale University
A big issue in functional genomics studies - looking at gene functions and interactions - is privacy, because these often require many thousands of genomes to make a really statistically strong case. The best way to do this is collaboration, but there are valid privacy concerns around sharing your participants’ genetic information. A new paper has validated those concerns; researchers combined genetic databases with DNA samples taken off of used coffee cups to prove that even innocuous-seeming data can identify someone if you combine it with a second source of information. They’ve come up with a solution to this, though. Phil Sansom heard from author Mark Gerstein...
Mark - We've created a method for sanitising functional genomics data so it can be shared without impinging on patient privacy.
Phil - What situation are we talking about here? Is this if I've sent my DNA off to 23andMe, or is it something else?
Mark - This is different from 23andMe. This is a subset of that type of genomic information called functional genomics information, that has to do with the activity of genes. Now when someone does a functional genomics experiment, the useful bits aren't the variants associated with you, usually; they're levels of activity of genes, the degree to which various things bind on the chromosome.
Phil - I'm surprised this kind of data wasn't secure already.
Mark - The leakage of information can be very subtle. And one of the ways of ascertaining this is to do a very subtle attack on the data called a linking attack, where you look at their ability to link to another data source of information. And so we've used these linking attacks to measure the very small amount of leakage from the experiments, and to calibrate our cleaning methods to be sure we really scrubbed all the variants away.
Phil - Okay, say I'm a genetics researcher then. How would I take your method and use it to sanitise, as you put it, my data? So that there's a balance between... I've not got rid of the crucial stuff, but I've got rid of enough so it doesn't identify someone or link to someone's identifying data.
Mark - This is less targeted at the individual researcher and more targeted at data repositories, where people would upload the data to be shared with the community. But an individual researcher could do this. It's a piece of software that you would run on the reads - these little bits of sequence that you get from the sequencing machine, and these things - and what it will basically do is go through the reads and find where you have variants in the reads, and remove a small number of those variants, a very measurable number, so as to disable any sort of linking attack or privacy breach from those variants. And then once you do that, you can essentially share the reads from the experiment publicly and without worrying about privacy.
Phil - Of course the other thing is that, if someone gets ahold of information like my street address, they can come find me or rob me or whatever. If someone gets a hold of my genome, what have they got? Well, if I haven't got a condition that I'm worried about... kind of bupkes, right?
Mark - We might not know what to do with genomic information now, but 50 years in the future from now we might know a lot more. And so there you have a potential avenue for people unearthing a lot of private information about one's offspring and future generations. Another scenario is that even with the information today, potentially people can find subtle statistical hints that you might have a predilection to one condition or another, which... a political candidate for office or someone in the public space, imagine if their genome got out and people could easily mine it, they might say, "oh geez, this person has a predilection to alcoholism". Or, "the person has a predilection to this," and whether it's exactly true or not is not really that important. The fact is it still can be very damaging information. There's a final scenario too, which is even more farfetched, but I'll just say it because it gives you a sense of the bizarre things that people think about in this. If someone got a bit of your genomic information, they could go to a crime scene and put a little bit down, right? And then they could implicate you in a crime. And again, this isn't something that the normal person thinks about, but it's something you could imagine, say, a state actor might be thinking about in relation to some foreign adversary or something.

17:43 - Gentoo penguins: three hidden species discovered
Gentoo penguins: three hidden species discovered
Josh Tyler, University of Bath
The gentoo penguin has been traditionally known as a species of penguin with colonies across the Southern Ocean, first described in the 18th Century by the naturalist Johann Forster. Hundreds of years on, new evidence suggests that his species is, in fact, a multitude. Josh Tyler from the University of Bath explained to Phil Sansom...
Josh - We found that the gentoo penguin, which is currently one species, is in fact four species. And we use DNA evidence, and we use morphological evidence to support our findings.
Phil - Really? Four species were hiding as one?
Josh - Yeah. So this is actually quite a common thing across birds, but also a lot of other species. It's called cryptic species, which are where the organisms are actually separate species but they look identical.
Phil - So how does that bump up your penguin species numbers, then?
Josh - Before this piece of research there were 18, and if we split the gentoo from one to four, that will take the total number up to 21; which is really exciting, that's an over 10% increase in species number.
Phil - Now what exactly are gentoo penguins?
Josh - Sort of medium-sized penguins. You can think of the king and emperor penguin being much larger; the gentoo penguin is the next largest penguin. And they have these really charismatic red tone bills, they have black heads, and they have these two contrasting white patches on their face, above the eyes.
Phil - Do these four species... is it, one lives on one island, one lives on another one, one lives in Antarctica, something like that?
Josh - You're absolutely correct. They have a range that covers the Southern Ocean; they're on a number of different islands there, and also on the actual Antarctic peninsula itself.
Phil - Were you down with the penguins, measuring how tall they are, and getting a bit of their genes, something like that?
Josh - Biology is sometimes not as glamorous as people might think. I was in charge of collating all of the data, but we had teams that went down and collected the genetic data directly from live penguins. And for that we take blood from the penguins, so you would lift the flipper up and take a small amount of blood; or alternatively sometimes you can get DNA from plucked feathers. And for the morphology we use museum specimens, which are much easier to get measurements from; because you imagine holding a penguin squirming, it's quite difficult. So we used museum collections in London and in New York, and you can take your trusty tape measure and your calipers, and you can take all of these really interesting measurements from across the penguin body. And we use those measurements for the physical characters.
Phil - What did you find?
Josh - When we looked at the genetic data, we found that members of each of these different populations were really closely related to each other, to the exclusion of members of other populations on other islands. So you can imagine that the penguins on South Georgia are very closely related to each other, but then they're far more distantly related to the penguins, say, on the Falkland islands. And in terms of morphology, we find there's actually a really interesting statistically significant body size trend. So the penguins in the Falkland islands are fractionally larger than their counterparts, say, in South Georgia, and with the smallest penguins being in the Western Antarctic and on the South Shetland islands,
Phil - How big a genetic difference is that?
Josh - From their genetic signal, we can see that they aren't mixing. And when we did the analysis, there were no individual penguins that might be jumping islands or jumping populations. And that's really exciting, and that is not always the case in genetic analysis, so it really reinforced this idea that they're each their own species.
Phil - Do you know how this dividing into species happened? Is it just the fact that they were separated from each other for a while?
Josh - I think that's absolutely correct. So if you were to imagine the Southern Ocean, the distances between these islands are very large. There's thousands and thousands of kilometres between some of these populations. And also there's a body of water called the polar front, which is where this current that goes around Antarctica sort of envelops Antarctica and the Southern ocean. And actually it's really difficult for penguins to cross that line. And so despite the fact that they can swim a very large distance, these gentoo penguins don't seem to be traveling too far away from their breeding grounds.
Phil - It is definitely interesting, but is it actually important for... I don't know, for the penguins themselves, for their conservation? Because in some ways it seems like it's quite a fine line between a species and just groups living apart, and in some ways it's a line that we humans have created.
Josh - You're absolutely right that there is a very subtle difference maybe between populations and species, depending on what avenue of science you're looking at. The most important thing from this study that comes out is: in terms of conservation, we're trying to protect and conserve diversity of organisms and life on the planet. And when organisations such as the IUCN Red List look at extinction risk, they operate at a species level. And so it's really important, when we find that there are these differences, that they are raised up to the species level so they can be accounted for accordingly. Whilst gentoo penguins across the board, if we took them as one species, are doing fairly well in the fight against climate change, they're not consistently doing well across their range. Some of these island populations are actually seeing marked decreases in population over the past couple of decades. So in order to protect them correctly and introduce measures that will protect the whole diversity of gentoo penguins, it's important to recognise that they're in fact four different species.
Phil - You better introduce me to those four species then!
Josh - Currently they are just named geographically. So we'll have the Falklands gentoo, the Western Antarctic gentoo, the Kerguelen gentoo, and then the South Georgia gentoo. I think that makes it easiest for us.
Related Content
- Previous Magic thinking
- Next Universal Flu Vaccine










Comments
Add a comment