Genes are the instructions that tell our cells what to do, but how do different types of cells know which genes to switch on or off at the right time? The solution lies in epigenetics - the molecular bells and whistles that act on top of our DNA to control gene activity. Plus, a new gene involved in severe obesity, and a mythical gene of the month.
In this episode

01:05 - Robin Allshire -Spools and strings
Robin Allshire -Spools and strings
with Robin Allshire, University of Edinburgh
Kat - This month I'm reporting back from a fascinating meeting I went to up in Edinburgh, a Wellcome trust Waddington Symposium entitled "Epigenetics in dialogue with the genome". We're starting to hear the word epigenetics used more and more frequently, but what does it mean, and what's it all about? A lot of the talks at the conference were centred on so-called epigenetic modifications of DNA - chemical tags that are put directly onto DNA, or onto the spool-like histone proteins that package it up, which are associated with changes in gene activity such as when genes are switched on or off. To find out more about these mysterious marks and how they work, I spoke to one of the organisers of the meeting, Robin Allshire from the University of Edinburgh.
Robin - Our DNA is not just a naked string. The best analogy I can come up with is you wrap it twice around a spool, there's a gap, it gets wrapped around it again, wrapped around it again a bit like beads on a string. You can have these chemical adaptations that cause it to close up or to open up. The debate is whether those adaptations, these additions of chemical groups onto the spools themselves are instructive. In other words, do they force the DNA to shut down? Are they just weak signals of some sort that are helping repression but are not the main driving force? So, in my point of view, what we really want to find out is whether these adaptations or marks, if you want to call them, on the spools actually can carry information through cell division.
Kat - So, let's go into a little bit more detail then. So, we've got their DNA. It's this long string of letters, of bases. It's wrapped around these spools. Tell me a bit more about them and the kind of marks that we can find on them.
Robin - So, these spools are made up of proteins and there are 8 individual proteins that makes up a spool, two of each type, and they're called histones. Those histones form a blob, basically, that the DNA wraps around. But they have these extensions that hang outside.
Kat - You're waving your hands around, like kind of octopus tentacles.
Robin - That's what they're like! That's the way I imagine them in my head. So, you can have adaptations that clamp those tentacles down onto the DNA. They maybe hug neighbouring spools so they're more tightly locked and you're going to have adaptations that cause them to wave around like crazy so that it says, "Come and get me and turn me on."
Kat - So, everything kind of loosens up and that the gene can be read.
Robin - Yeah, exactly.
Kat - So, it's all about kind of combinations of these marks, the tails, how open and close it is, and then all these molecules that are recognising them and locking them down. It all seems complicated.
Robin - So for example, you can generalise to some extent with respect to one of the type of chemical marks which is acetylation. In general, acetylation promotes gene expression. I like to think that it sorts of oils of these tentacles and loosens them off. But again, it can bring in various machinery that opens up then the spools, allowing the DNA to be read.
Kat - What do we know about how these marks can be propagated as cells divide or even as organisms divide and reproduce because it seems to be quite a controversial area? What do we know and what do we not yet know?
Robin - These are chemical marks on the histones are very frequently, referred to as epigenetic modifications. The word 'epigenetic' means many different things to different people. A strict definition is the propagation of a state without the presence of the inducer that enabled that state. So, the question becomes whether these marks can actually be copied during the process of cell division so that both daughter cells have an exact copy of the marks.
Kat - So, if a skin cell divides, it makes 2 cells and they know that they're meant to be skin cells.
Robin - Yeah, exactly. So the question is, can the more actually be recognised, so as the cell goes through division, you have to deposit or assemble, make new spools on the new DNA, and can you copy the marks from the pre-existing spools that were there so that they take on the same state?
Kat - And can you? This is the big question, isn't it?
Robin - So, this is the big question. So recently, what we've been able to do in a model organism, fission yeast, we have tested this in a very simple way where we artificially bring the enzymes - so that's the proteins with an activity that puts a certain chemical mark which is called lysine9 methylation on a specific part of one of the histones in the spool - what we were doing is we artificially bring it somewhere and then we're able to get rid of it. Then when we get rid of it, the question is, can it be copied? In normal cells - so wild type cells - it cannot be copied. But then we ask, "Can we make it be copied?" what we did was we took away an opposing activity that removes that mark. When we take it away, now we see that it can be copied. Not only is it copied for one cell to two daughter cells, we can see it being copied for many cell divisions and we're going to also see it go through the process of meiosis.
Kat - That's kind of yeast babies?
Robin - Yes, yeast babies.
Kat - Well, it sounds to me like there's a lot of molecules that are putting these marks on, taking them off. There's this balance between the copying and the removing, all these kind of things. It feels like a very complicated field. Where do you think the really big questions are, the things that really need to be sorted out?
Robin - Well, so one of the things that people are trying to figure out is, whether these modifications are instructive in the sense of responding to environmental differences or pressures for example in a plant temperature or geographical location. In us, in people, you could say whether you've got high fat diet or a low fat diet. That's a very active and hot area at the moment where people are trying to see, is there an underlying contribution of these type of events - for example, there's lots of studies that show that diet can influence traits in children but then the question becomes is that just because they were developing in the womb of a mother who had a bad diet or smoking too much, or whatever, so environmental effects. So, the real test is whether it can be transmitted via the father through his sperm because of his bad habits. What we found at this meeting actually is that there seems to be some interesting things happening in that area and I think that's a big hot topic for the future.
Kat - What's it like to be part of this now?
Robin - It's exciting and there's various controversies that need to be sorted out but that just adds to it - scientists are naturally questioning both in terms of asking how something works but also questioning whether the way we think things work is actually the way they do work. And we're all the time tearing down the models that we create. That makes it exciting because we're still trying to figure out things. It hasn't gone stale. There's a lot of things to find out yet.
Kat - That was Robin Allshire, from the University of Edinburgh.
08:43 - Alex Blakemore - New obesity gene
Alex Blakemore - New obesity gene
with Alex Blakemore - Imperial College London
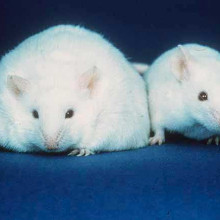
Kat - More than 30 genes are now known to be involved in obesity and, intriguingly, most of them seem to be active in the brain, controlling eating behaviour and hormones. Researchers at Imperial College in London, led by Alex Blakemore, have used DNA sequencing technology to pin down the cause of one young woman's severe obesity to a faulty version of a gene called carboxypeptidase E, or CPE, publishing their findings in the journal PLoS ONE.
Alex - The earliest studies of genes causing human obesity followed on from looking at mouse models of obesity. So, you have certain strains from mice that, just hanging around in their cage with other mice, had a tendency to eat very much more than the others and became very, very fat. Those mice weren't doing that because they were mice with weak personality or rubbish life. They were just mice along with the other mice. And so, by looking at those mice, we found a way into understanding what controls eating behaviour. The first mouse that was investigated in that way was called the 'obese' mouse, and that began to be understood in about the early 1990s. But since 1974, there's been another mouse that became obese which is known as the 'fat' mouse. That mouse had a defect in a gene that processes the hormones that control appetite and the regulators in the brain that control appetite. But there had never been a human person found with problems with that gene.
Kat - Until now.
Alex - Until now. This is the first case of what's called carboxypeptidase E deficiency found in humans although we've had the mouse models for some time. The mouse has eating behaviour problems becomes very obese. It has obesity, it has fertility problems, and it has memory problems. And so, this woman that we found was a woman of 20 years of age who has problem with the same gene. She also have that same constellations of features. So, she has severe obesity since she was a child. At the age of 20, she weighs twice what her maximum recommended weight would be. She has reproductive issues, some intellectual disability and hasn't been able to learn to read despite schooling. She also has type 2 diabetes.
Kat - And how did you actually find out that this woman had this faulty gene?
Alex - She was referred to us by the consultant endocrinologist, Tony Goldstone, who was looking after her. And so, we included this family in a sequencing study looking at the sequence of their genes. And we first looked for problems with some of the already known obesity genes, but we drew a blank there. So then we started to look at other genes that had been implicated in obesity but not found in humans and big among those among those was the fat mouse. So, we looked at that gene and we were surprised to be that this young woman had two broken copies of the gene so she couldn't make carboxypeptidase E at all.
Kat - Given that you found this one woman, this one family where it seems to be this gene fault, do you think that there may be other people out there with this faulty gene? Do you think that it might lie at the heart of many cases of severe obesity?
Alex - It's very difficult to tell right now. While we were writing the paper about this, we saw that in an online database of genome sequences from around the world, two other people have been seen who carry exactly the same mutation. So, it could be that it's more common than we anticipate but has been missed.
Kat - So, is it fair to say that obesity is really in the genes, but then what can we do about it?
Alex - I think we have to stop thinking of obesity as a single disorder. Everybody is obese for a different reason. For some people, it might be almost totally environmental and for others, it's almost totally genetic. One of the useful things we can do is try to find ways of seeing which is which, looking at people to say, "Okay, is this a genetic condition for you or is it an environmental condition for you" because the way we manage people might be different, depending on the different causes.
Kat - So, it's not as simple as saying, "It's just all in my genes. There's nothing I can do. Pass me the cake."
Alex - No, absolutely not. That's a very funny idea that people have got that because you might tell somebody that they've got a genetic condition then they won't be motivated to do something about it. In our experience, the opposite might be true. It's very, very helpful for people to have a diagnosis and be told, "You have an actual disorder which we can describe. It's not your fault. It's not a character flaw, but it is a battle you're going to have to fight for the rest of your life." In our experience, it doesn't make people throw in the towel. It gives them renewed vigour because what really saps people's energy is the sense of guilt and shame about their obesity and the feeling that there's nothing they could do about it because of that. So, to say something genetic, it doesn't mean it's cast in stone or that you shouldn't fight against it. It just lets you know that you're likely to have a harder fight than the general person in the street.
Kat - That was Alex Blakemore, from Imperial College.

14:35 - Peter Joshi - The same or different?
Peter Joshi - The same or different?
with Peter Joshi, University of Edinburgh
Kat - Every human on the planet is distantly related in some way, but people who live in the same geographical area tend to be more genetically similar than those living a long distance apart. Now scientists have discovered that people who inherit the same versions of certain genes from mum and dad, are likely to be shorter and do less well at school than those who get different versions from each parent. To find out more, I spoke to University of Edinburgh researcher Peter Joshi, one of the hundreds who contributed to the study.
Peter - We inherit one copy of each gene from our mother and one copy of each gene from our father. Sometimes we inherit the same copy from both parents. What we look at is, in the context of the genome, relatively short stretches of about 1.5 million letters at a time and see whether or not those 1.5 million letters had been inherited the same from your mother and your father. Somebody in that situation doesn't have genetic diversity in that part of the genome. And vice versa, somebody that has inherited different genetics over those 1.5 million letters is genetically diverse.
Kat - And we're just talking not necessarily about really bad gene faults. We're just talking about you know, sometimes the variation that makes us unique and different.
Peter - Yes, absolutely. So the traits that I study are what geneticists called complex traits and they're exactly about what makes us all unique and different, and a little bit different from each other. What we see is that there's lots and lots of genes that have small effects and it's the combined effects of those and the environment that all make us individual people.
Kat - Tell me about the study. Who were you looking at? What were you actually analysing?
Peter - What we're doing is using modern genomic techniques to measure genetic diversity. The way that we're doing that is that we're using 350,000 people in total, and to make sure that the study is very robust, we used a hundred different populations across the world with all of that information and then collating it centrally in Edinburgh. We basically looked and measured the genetic diversity of each of those individuals.
Kat - So, you've got hundreds of thousands of people from populations across the world, how are you analysing their genomes? What are you looking at?
Peter - What we do is we look at the genetic diversity of individual people within that population and compare them with other people within the population. At the same time, look at their cognitive ability, height or blood pressure. Within that population, see whether or not there's an association between genetic diversity and say, educational attainment. Having done that, we then combine all the results of the different studies to see whether or not the effect is robust and reliable across lots of populations.
Kat - So, when you started looking at the levels of diversity about whether people had two copies of a gene or two different copies of a gene, what did you start to find?
Peter - What we were doing is we were looking at the whole of the genetic code of each individual and we're scanning along that genetic code and checking what proportion of that genetic code is identical from the mother and the father. What we typically find is that about 0.1 per cent of the genetic code is an inherited identical from the mother and the father. But that number varies from individual to individual with some people that might be none at all and in some other people that might be 3 or 4 times that. What we find is, that if you've inherited 2 or 3, 3 or 4 times the norm in terms of lack of diversity, that reduces educational attainment, cognitive ability, and height.
Kat - So, people who've got more similar genes from their parents, they don't do as well at school and they're shorter?
Peter - Yes, but it's also important to recognise that these effects are small. But on an individual level, it wouldn't be measurable. That's why we needed 350,000 people to robustly demonstrate that it was a real effect.
Kat - So, that height and broad measure of intelligence, but what about something like your risk of illness, because we know that the risk of diseases is also encoded in our genes?
Peter - It has been known for a long time that rare particularly strong genetic diseases like cystic fibrosis are subject to this effect. But what we hadn't shown was that these more complicated traits like educational attainment, which has got a lot of environmental factors in it as well obviously, like height, showed these effects. In fact, what we found for the risk of heart disease that we found no evidence in fact that genetic diversity affected your risk of heart disease. The mechanism that we think might be involved is that there will be things associated with development that might underpin growth and therefore your height. And so, it's not height itself in a way that is being controlled in this way, but it's the sort of underlying biological systems and two bad copies might affect height in that way and cognitive ability. If that happens, if these traits are favoured by evolution then we see this overall effect whereas for traits that are not subject to evolutionary pressures, we don't see that in quite the same way. It's plausible that heart disease hasn't been under the amount of selection pressure from evolution as you might first think, simply because people tend to get heart disease after they've had their families and its reproductive success that counts in Darwinian evolution.
Kat - Peter Joshi from the University of Edinburgh, and that study was published in the journal Nature this week.
20:28 - Rob Martienssen - Small RNAs, big news
Rob Martienssen - Small RNAs, big news
with Rob Martienssen, Cold Spring Harbor Laboratory
Kat:: Now it's time to take a look at some of the most interesting molecular players in the world of epigenetics: small RNAs. Many of the talks at the symposium focused on RNA - a kind of molecular 'cousin' of DNA, which is made when DNA is read, or transcribed. Researchers are now finding more and more roles for these little fragments in controlling how genes are turned on and off - and their effects may potentially even travel across generations. To get the low-down I caught up with Rob Martienssen, from Cold Spring Harbor Laboratory in New York.
Rob - So, small RNAs were discovered in probably the late '90s but they turn out to be everywhere pretty much. So, we work on looking at these short RNA sequences, specifically the ones that regulate transposable elements, repetitive, junk DNA, things that were thought not to be important for a long time. But actually, we think are hugely important parts of genomes that control epigenetic inheritance.
Kat - That's a big word. What do you mean by epigenetic inheritance? How would you describe that?
Rob - Usually, we think of inheritance as being the inheritance of changes to our DNA sequence, to our genome sequence. But epigenetic inheritance are modifications of that sequence that can be reversed or changed and are not fixed in the same way that DNA sequence changes are. But they can be just as important. Importantly, you have to think about how they could be guided to specific DNA sequences that they would modify in some way that would allow that to be inherited. And so, we think that small RNAs are very important in how that inheritance happens.
Kat - So, this is basically how our cells or how any organism's cells manage to kind of be a bit more responsive or responds to the changes in the environment around them and have different types of cells.
Rob - That's right and importantly, how they control their transposable elements which can occur over generations, actually first shown year ago by Barbara McClintock, who described something called cycling where transposons would go on and off over, not just a few cell divisions but over actual generations like multiple generations.
Kat - So, tell me a little bit more about these small RNAs. What do they look like, how are they made, and what do we think they're doing?
Rob - So, the small RNAs are short nucleotide sequences, about 20 to 30 nucleotides long.
Kat - That's kind of letters of RNA.
Rob - Right, letters of RNA that correspond very closely to the letters of DNA and that's how they can recognise particular genes or transposons in the genome. They're actually almost the perfect length to do that within a genome of the size of a human genome or of the maize genome. What they seem to do is they bind to specific proteins called argonaute proteins. There are many, many different argonaute proteins that bind the small RNA and allow them to match a corresponding sequence in a target RNA. The target RNA in many cases gets cleaved by the argonaute protein which is actually an enzyme that can break RNA.
Kat - So, they basically chops it all up and gets rid of it.
Rob - More or less. It actually does more than that because it seems like that process of chopping it up and getting rid of it also guides, targets other enzymes that modify either the DNA by DNA methylation or the proteins that DNA surrounds - the histones, chromatin, we call the ensemble of all of that. These also get modified in response to that processing of RNA. It's one of the things we work on. We still haven't exactly figured out how that happens, but a lot of work has been done especially in plants and in fission yeast.
Kat - I find this whole process really fascinating that you have these tiny little fragments of RNA, they seek out things that they match and then almost like - the magic happens! This is how genes can get switched off, can get silenced. What do we know about how important this is in regular life for cells or for plants? We know that it is maybe important for shutting off these jumping genes, these transposons, but what do these small RNAs do in kind of regular biochemistry?
Rob - They do a lot. They also target genes. When they target a gene of course, they can turn the gene on and off. And so, it's a very powerful control mechanism both in normal development in humans, in plants, also in animals. And importantly, in cancer and a lot of diseases, some of these small RNAs are very, very important and prevalent in regulating genes in that way. They can also influence everything in plant development. For example, one of my favourites is plants have a juvenile and an adult phase, just as - think of a teenager, adult...
Kat - A teenage plant!
Rob - This difference is actually completely controlled by small RNAs. There's one that's specific to juvenile phase, one specific to the adult and they regulate each other and it's a fantastic story.
Kat - Given how widespread these tiny little RNAs seem to be and how people are finding them in more and more organisms and doing more and more things, is it safe to say that they're pretty much involved in everything when it comes to controlling genes? Is there anything they can't do?
Rob - That's actually not far off the case. Literally, half the transposons in the model plant we like to use, Arabidopsis, are targeted in this way - something in the order of 500 genes. In humans, estimates vary but certainly, thousands and thousands of genes are regulated by small RNAs. It could be that every gene is regulated by small RNA.
Kat - We just haven't found it yet. They're so small!
Rob - Exactly.
Kat - Where next do you think for this field? Lots and lots of people are identifying these small RNAs, trying to figure out how they work, how they regulate genes, how they control them. Where do you think things are going?
Rob - There's a lot of interest in connecting the epigenetics - that's the heritable things that happen to DNA - to the small RNAs. That's field has been really exploding in the last few years and I think in the next few years, it will come to a conclusion which will be very exciting. I think inheritance of the small RNAs themselves, or at least of their effects. In plants, it's probably more well-established that this could happen. In humans, are beginning to look into that.
Kat - So, that's from generation to generation rather than just cells divide.
Rob - That's right. So I mean, there are some people who are really beginning to say that maybe germ cells, sperms and eggs actually contain a lot. They contain a huge amount of RNA. It could be that this is actually transmitted in some way. There's also new types of small RNAs, so they're continually being discovered. One of my personal interest is recently, genome editing has been shown to be controlled by a type of non-coding RNA that's found in bacteria. It's work brilliantly well in ourselves as well. I'm wondering if the mechanism might be somehow related. That's going to be a fun area because we know recently that small RNAs are involved in DNA repair. I think that's going to be an important area as well.
Kat - Given how important these small RNAs seem to be and how easy it is maybe to make them and to manipulate them, are there any therapeutic implications? Maybe we could develop medicines or ways of gene therapy using these things.
Rob - There is a lot of interest in that. Several start-up companies have done very well on that specific idea. Using the RNA exactly as it is, it probably needs some modifications to make it useful as a drug, but it's definitely a big area. In plants, some seed companies are thinking of spraying plants with small RNA. It's something that I was very sceptical about but apparently it works. I think that's going to be a big area, absolutely.
Kat

28:19 - Gene of the Month - Ariadne
Gene of the Month - Ariadne
with Kat Arney
Kat - And finally, it's time for our mythical Gene of the Month - and this time it's Ariadne. The daughter of King Minos of Crete, the ancient Greek story goes that Ariadne fell in love with Theseus, who was aiming to kill the terrible bull-headed Minotaur hiding in its Cretan labyrinth. Ariadne sneakily gave Theseus a sword to stab the beast, and a ball of string to help him find his way back out again. Away from ancient Greece, the Ariadne gene was first discovered in fruit flies, and is involved in helping nerve cell axons - the long wires of our nervous system - find their targets, just like that helpful ball of string. Flies with a faulty version of Ariadne don't usually survive, and those that do have problems with their nerves and muscles. There are also versions of Ariadne in mammals such as mice and humans, and there are hints that it might be involved in the development of the neurodegenerative disease Parkinson's.
- Previous Make it Digital!
- Next Climate change is bad news for bees









Comments
Add a comment