DNA Decoded: Past, Present and Sausage
This week we delve into DNA and what it can tell us about our past, present and future. And, what happened when we decided to read the DNA sequence of a local sausage? Plus, in the news, the Nobel Prize announcements, the world’s largest HIV survey, and why doing exercise you don’t like makes you more likely to binge on junk food!
In this episode

01:08 - Gene therapy lets blind mice see
Gene therapy lets blind mice see
with Samantha De Silva, University of Oxford
A gene therapy technique that can help to repair the retina and restore lost vision has been pioneered at the University of Oxford. The retina is the light-sensitive sheet of tissue at the back of the eye where rod and cone cells convert light waves into electrical signals that the brain can understand. But in diseases like retinitis pigmentosa, these cells die off, leaving patients unable to see. Now Samantha De Silva and her colleagues have developed a way to use a harmless virus to deliver genetic instructions that can make other, surviving healthy cells in the retina become light sensitive and make blind mice see…
Samantha - In these conditions there’s a gradual loss of light-sensing cells at the back of the eye, so those are the rods and cones, and, over time, the rods and cones eventually die resulting in blindness. Despite the loss of rods and cones, the rest of the cells in the retina actually remain largely structurally intact, along with the connections between those cells and the connections to the brain. So what we aim to do is see if there’s any way of stimulating these remaining retinal cells to try and restore light sensitivity and even vision.
Chris - So essentially, you’re turning cells in the back of the eye that wouldn’t normally function, like rods and cones, into cells that can take up some of the light sensing functions of the rods and cones so you can get signals into the brain again?
Samantha - Exactly. Initial work done in 2005 by Professor Mark Hankins, who’s one of the senior authors on this paper, showed that if you take a light sensing protein, and one of those is called melanopsin which is already present in the human eye, and put it in a cell which is not light sensitive, that cell then becomes light sensing. Based on that work we tried to see if we could express melanopsin, so human protein, in these remaining retinal cells and make them light sensitive in the absence of rods and cones.
Chris - How do you get the instruction to make that light sensing chemical melanopsin into the cells that now need it then?
Samantha - This is where the gene therapy comes in. Essentially, what we do is we take a virus called an adeno-associated virus which is harmless to humans, and we genetically engineer that virus to get rid of all the genes and proteins that it normally expresses and make it express proteins that we want it to express, and that’s the ultimate concept of gene therapy. What we did is we engineered these viruses to express melanopsin and then injected them underneath the retina of mice with retinitis pigmentosa to see whether the virus was taken up, and whether it would be effective.
Chris - Does the virus go into these cells that are surviving and add the genes to them?
Samantha - Yeah. The first finding of our paper really is that this virus was effective. It was able to get the melanopsin into the remaining cells and that expression was initially seen after a few weeks but persisted after 15 months after a single injection. So it essentially lasted the whole lifetime of that mouse.
Chris - Was there any inflammation because, obviously, you’re putting a lot of something foreign into a part of the body where it would never normally be seen - does the immune system react?
Samantha - That's partly the beauty of using the subretinal approach in that the space underneath the retina is what we call immune privileged, so there is not normally an immune response mounted to anything being in that area. The eye lends itself very well to gene therapy for that reason so we didn’t see any immune response in any of the mice that were injected or treated.
Chris - How did you know you had made a difference to the vision in these mice?
Samantha - Once we were able to establish that we had got the virus working and there was melanopsin in the cells, we used electrodes to record from the treated retinas and that showed that when we shone a light on the retinas they generated electrical signals that could be transmitted to the brain.
Chris - And then, how did you prove that the brain was actually interpreting this information because it’s one thing for the retina to be a little bit more electrically active but that doesn’t mean the signals are going to the brain?
Samantha - Exactly. So the next step, we looked at a number of different things - one was the pupil light response. Normally, when you shine light into someone’s eye their pupil constricts and that’s a marker of the appropriate circuits within the brain being activated. In blind mice, that pupil response is severely attenuated because of the lack of detection of light. Whereas in the treated mice, the pupil light reflex was restored both at 2 months and then going on to look at mice at 13 months.
In terms of the light responses and vision responses we found, we found that the mice in our study were able to detect a change in their visual environment and that would probably equate to a completely blind person being able to recognise their environment, so possibly where a door is, where a window is, where an obstacle in the road or something like that is. So we’re not talking about rapidly dynamic vision because I don’t think melanopsin would be able to restore that but, obviously, if you’ve got nothing at all to start with then that’s a significant improvement.
Chris - Is the next step to now do a clinical trial? Are you at the stage where you could safely do that?
Samantha - Essentially, all the groundwork has been done and to take it forward to a clinical trial there are further steps such as to make a virus which is suitable for use in humans rather just in the lab. All that, and the regulatory work takes a couple of years so we’d hope to try and get it to patients within the next few years.

06:44 - Nobel Prize: Tick Tock, Body Clock
Nobel Prize: Tick Tock, Body Clock
with Ned Hoyle, Laboratory of Molecular Biology
The 2017 Nobel Prizes have now been announced. The first to be made public was for Physiology or Medicine, which went to Jeffrey Hall, Michael Rosbash and Michael Young, who sussed out how the “body clock” works. “Clock doc” Ned Hoyle researches this subject himself at the Laboratory of Molecular Biology in Cambridge, and he took the time to take us through it...
Ned - Animals, plants, some fungi, and even bacteria possess a 24 hour timer which is hardwired into their biology. This allows them to pursue a lifestyle which reflects a day/night cycle of our planet. This year’s Nobel Prize for Physiology or Medicine is for pioneering work in fruit flies. The scientists identified the genetic behind how organisms can keep to a daily rhythm. Hall, Rosbash and Young didn’t just identify the genes involved, which was an arduous task at the time taking several years, but they also figured out how they act to measure time.
The genes involved in the biological clock can turn themselves on and off in a process known as negative feedback. The activity of period, the first gene they identified, rises during the night until dawn when it switches itself off. During the day levels of period fall until it switches itself back on again, beginning the cycle again. It’s this on/off switching repeating itself each day which is the fundamental basis of daily biological clocks.
This elegant mechanism has held true not only for flies, but also for mammals, plants, and even fungi. Later research has shown that a large part of human physiology is affected by daily biological timekeeping including hormones, sleep, body temperature, and metabolism. Understanding of biological timekeeping has impacts upon human health for example, shift work where our internal clocks are misaligned with the day/night cycle is a risk factor for a number of diseases including type 2 diabetes and cancer. With greater knowledge of our body clock we might one day mitigate these harmful effects or at least make more informed choices about our lifestyle.
09:53 - World's largest HIV survey to begin in 2018
World's largest HIV survey to begin in 2018
with Sani Aliyu, Nigerian National Agency for the Control of AIDS
Nigeria is Africa’s most populous country: about 190 million people live there, and it’s growing rapidly. But millions of people there are also infected with HIV, and Nigeria has received billions of Dollars in international aid to help them to control the spread of the disease.
No one knows, though, how many people are actually infected, or where, so it’s hard to ensure that these resources are properly used. Last year Sani Aliyu, who’s an infectious diseases consultant from Addenbrooke’s Hospital in Cambridge, was seconded to Nigeria to help, and he’s now setting up the largest HIV survey in the world. He told Chris Smith how they're going to do it...
Sani - What’s happening in Nigeria is the country is one of a handful of countries facing the triple threat of a high HIV burden, a treatment programme that doesn’t really cover the majority of patients with HIV, and also a very slow decline in the number of new infections. So when I started, one of the key problems we had was being able to assess the impact of the programme. We were having targets that were clearly not achievable. We had issues with prevention of mother to child transmission where, on average, one in every three babies born in the world with HIV is a Nigerian child. But the targets set up for the implementing partners was really not achievable.
So we decided that the best way to go about dealing with this is to establish the true prevalence of HIV in the country to find out how many people have we got, and what we are planning to do is do a two stage cluster survey of households. It will be the largest HIV survey ever done in the world. Our sample frame constitutes about 30 million households.
Chris - That’s geographically dotted across Nigeria? So what are you doing, dividing the country into sectors so that you’re taking a representative cross-section of each of these geographies?
Sani - That’s right. We have what we call enumeration areas, so first of all we will take up enumeration areas within every sampling frame. And within each enumeration area you will have 25 households. We have 60 geopolitical zones across the country so that makes it much easier and we’ll be taking states at a time, so national entities at a time. As we go through, we’ll not only be providing HIV counselling and testing, we’ll also be looking at incidence of HIV, and for those who are already HIV positive are they virologically suppressed and do they have any resistance issues, are there issues with quality of services? So a whole range of things but it’s going to be a huge exercise. The US government through the Presidential Emergency Fund for AIDS relief is funding this to the tune of about 85 million US Dollars. It makes it, by far, the largest investment on a survey ever done on HIV.
Chris - But why will this help to get a handle on what’s going on in the country and achieving the decline in the spread of HIV in Nigeria?
Sani - A lot of money has gone into the HIV national programme in Nigeria. We really need clarity on the impact of the programme. Are we doing well in the first instance and, if we’re not doing well, why is that we’re not doing well? My feeling is that a lot of the problems arise from the difficulty in really targeting high prevalence areas, being able to identify parts of the population that have a high prevalence of HIV. This survey will give us an answer to that because we’ll be able to channel those resources effectively.
The US government have spent about 4.3 billion US Dollars since the beginning of the HIV programme in Nigeria alone. Global fund has spent about 1.3 billion US Dollars. And even as I speak, the US government are spending between 350 to 400 million US Dollars every year on more than a million people on treatment. But, as we get more and more people on treatment, it’s going to become more and more difficult to find those positives left.
Chris - The law of vanishing return?
Sani - Exactly. And we need to be more cost effective in terms of how are using their resources to find those positives.
Chris - Some people may say that why are we spending so much money on one country and there are lots of needy situations, lots of deserving cases and causes - what’s the value of investing such prodigious sums in one country? Admittedly, there are a lot of people there, why should we be spending this money in Nigeria?
Sani - We talk about trying to eliminate HIV in the long run; having epidemic control globally. You cannot have epidemic control of HIV in the world without having epidemic control in Nigeria. It will just not happen because we estimate about 3 million people have HIV. We have the second largest HIV burden in the world. We need to able to get on top of the HIV situation in Nigeria otherwise it will be like throwing good money after bad money. A lot of the East African countries have very done well based on similar surveys that have been done, but because they are smaller countries, it’s cheaper to get these surveys done whereas in Nigeria it’s a completely different issue. The answer we will get from Nigeria will enable us to target where the resources need to go to and we’ll be able to know what additional resources we need to get on top of the 1990 targets set by the United Nations.
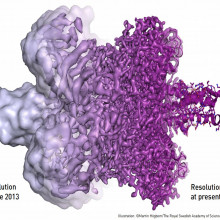
15:10 - Nobel Prize: Mapping Moving Biomolecules
Nobel Prize: Mapping Moving Biomolecules
with Tony Crowther, Laboratory of Molecular Biology
Next on our Nobel Prize list is Chemistry, which was awarded to Jacques Dubochet, Joachim Frank, and Richard Henderson for their work on an imaging technique called cryo-electron microscopy. Tony Crowther uses electron microscopy in his research and has known Richard for many years as a colleague at the Laboratory of Molecular Biology in Cambridge. And many thanks to MRC for providing us with audio from the winner himself, Richard Henderson...
Richard - I’m delighted to be sharing the Nobel Prize. I think…
Tony - To understand the way life works in health and disease, it’s necessary to know the detailed structures of the complex molecules involved. Many processes in the cells of all living things depend on large, dynamic, molecular machines. Previously this required making crystals of the molecule and solving the atomic structure by X ray crystallography. This could be difficult as important molecules or complexes might not crystallise and movements in the molecules key to their function would be inaccessible.
This has all changed with recent developments in cryo-electron microscopy, particularly in electron microscopes, in electron detectors, and in computer software. The molecule of interest can be trapped in a frozen state and its atomic structure determined by computer averaging of multiple images of views in different directions. This may also illuminate different conformations important in understanding the function of a molecule.
The newly developed technique, still open to some improvements, will be useful to all biologists interested in molecular structures and to pharmaceutical companies involved in drug development. For example, already in the LMB the structure of the tau protein filament important in Alzheimer’s disease has recently been solved.
Richard - So we think in a few years, or five years we might know most of the structures certainly in the human biological population and then in pathogenic bacteria. So really it’s quite an exciting time.

17:21 - Gym or junk food: What would you choose?
Gym or junk food: What would you choose?
with Natalya Beer, University of Western Australia
Michael Wheeler’s been working up a sweat down at the gym; but did he decide the exercise regime, or did someone else? It’s an important distinction because it could determine how tempted he is afterwards by the junk food in the vending machine...
Michael - Many of us can relate to those days when we manage to get in a tough exercise session but then all we want to do is eat is junk food. But, on other times, completing and exercise session motivates us to eat healthy… our bodies seem to work in mysterious ways. But now researchers from the University of Western Australia have shed some light on the link between exercise and subsequent eating behaviour…
Natalya - We all know the benefits of exercise, but it’s also important to note that sometimes our behaviour after an exercise session can actually counteract some of these benefits.
Michael - That’s Natalya Beer; she’s an exercise researcher at the University of Western Australia…
Natalya - We wanted to see how manipulating people’s psychological approach to exercise would influence the way they approach food after exercise. And specifically we manipulated people’s sense of choice in a workout to then see how much food they would eat afterwards, and the types of food that they would reach for.
Michael - The experiment that Natalya and her colleagues did was to randomise people into two groups prior to completing an exercise session. One group had lots of choice about things like the kind of exercise they did, the exercise intensity, the exercise duration, as well as what music to listen to. Now, for every person in this choice group, there was a matching pair who did a similar exercise session but with no choices available.
The matching pairs were similar in almost every respect, except one had lots of choice and the other had no choice. They were the same gender, they were similar in height, age, weight, fitness, and they expanded a similar amount of energy in the exercise session. The researchers then secretly measured the amount and the type of food that the participants ate after exercise...
Natalya - At no point were they told that we would be measuring their energy intake or their appetite. And when we presented the food buffet to them we actually said that that was a thank you gesture.
Michael - The researchers used a questionnaire to confirm the participants were not aware of the true nature of the study which was to measure what they ate from both the pile of healthy food and the pile of unhealthy food…
Natalya - We provided obvious options such as low fat milk, whole grain bread, whole grain breakfast cereals, fruits, things that we’re quite familiar with being healthy. And then with the unhealthy foods we had quite obvious choices again, so we had muffins, lollies, chocolate biscuits.
Michael - I certainly know what foods I would reach for, but the researchers found something quite interesting. Despite both groups reporting similar ratings of appetite, the group who weren’t given any choice in their exercise session ate more of the unhealthy food and had an energy intake that was about 30% higher than the group who were given lots of choices. So what is going on here?
Natalya - In the instance of this experiment, perhaps exerting self-control by continuing an exercise they don’t particularly enjoy may lower their capacity to be able to exert self-control after that when they’re exposed to foods that they might want to go for. We looked at their blood glucose levels as there are a number of theories out there that suggest that with this lowered self-control, they might also experience low blood glucose levels, but we found quite similar blood glucose levels between the two conditions.
Michael - It’s a clever experiment and the researchers believe there is a lesson to be learned…
Natalya - Well, from what we’ve seen in our study it seems that simple acts of providing choice in an exercise session, if they can influence the way we behave in a positive way after that exercise session, then why not incorporate those aspects. It’s an important consideration for anyone that’s involved in exercising themselves and wants to get the most out of their exercise. But also for people motivating others to exercise to ensure that they’re giving those people they are motivating some choice about what they’re doing and making sure that they enjoy it, and doing it because they would like to as opposed to being forced to do it.
Thank you to DjDust for the apple-crunch sound effect

21:60 - Nobel Prize: Gravitational Waves
Nobel Prize: Gravitational Waves
with Martin Hendry, Univeristy of Glasgow
The 2017 Nobel Prize for Physics has been awarded to Rainer Weiss, Kip Thorne and Barry Barish for their work leading to the first detection of gravitational waves in 2015. Glasgow-based astrophysicist Martin Hendry explains...
Martin - Gravitational waves are ripples in spacetime predicted a century ago by Albert Einstein’s general theory of relativity, and produced by some of the most violent events in the cosmos. By the time they reach the Earth, however, gravitational waves disturb our patch of spacetime by a mind bogglingly small amount - a million millionth the width of a human hair.
The 2017 Nobel Prize for physics was awarded to three key figures in the story of LIGO, who helped to develop the incredible technology required to measure those tiny ripples, and to isolate them from all the local disturbances that otherwise could completely drowned them out. The two LIGO facilities, the most sensitive scientific instruments ever built, first detected gravitational waves in 2015 from the merger of two black holes more than a billion light years away.
Katie - LIGO is the instrument used to detect gravitational waves. Using the interference of laser light to detect the squeezing and stretching of spacetime as gravitational waves pass by…
Martin - The LIGO discovery was a huge team effort that involved thousands of scientists across the world, working together over decades to turn what many had considered an impossible dream into the hottest topic in astronomy. But LIGOs first detection of gravitational waves was only the beginning. Since that discovery was announced in February last year, three more detections of merging black holes have been confirmed and we’re seeing the first hints of how this population of black holes fits into the big picture of how the universe evolved.

24:23 - DIY: Extracting Strawberry DNA
DIY: Extracting Strawberry DNA
with Jonathan Lawson, University of Cambridge
What exactly is DNA and what does it look like? Jonathan Lawson from the Babraham Institute near Cambridge, has an experiment you can try at home...
Jonathan - DNA is the instruction manual for every living thing. In every human cell there’s about two metres of DNA; what it is series of chemical units called bases which are joined together in a long chain which is what makes the helix structure. There’s four different chemical units A, C, G, and T, and the order of those is how information is stored.
So now we’re going to extract some DNA from some strawberries and we’ve come to the Old Cavendish Laboratory in Cambridge, which is where DNA was first worked out by Watson and Crick in 1953.
We’re going to take some strawberries, put the strawberries in a plastic sandwich bag, and then to that you want to add a little bit of salt, some water and, finally, some washing up liquid. Then basically with you hands you’re going to squish the strawberries as much as you can so that all of the DNA gets released from the cells, and there you have a nice, fairly clear red liquid, but you can’t see the DNA in this form.
The way you visualise it, the way you get to have a look at it, is to make lots and lots of DNA stick together so that they’re then thick enough, in clumps, to be able to see them. And the way you do that is by adding some alcohol. Now when you look you’ll see very thin, furry white strands and that’s the DNA all clumped together. And all the genetic information that was needed to make strawberries is in those clumps of white DNA.
Although what we’ve done just here is quite a simple experiment that you can easily do in the kitchen, exactly the same procedure - a detergent to break open the cells, salt to protect the DNA, and alcohol to separate the DNA from the rest of the cells is exactly the same thing that happens in labs all over the world right now when they are sequencing DNA and looking to understand genes and the genome

26:23 - The Experiment: What's in a sausage?
The Experiment: What's in a sausage?
with Ed Farnell, University of Cambridge
So we can get DNA out of a strawberry in under a minute. But why stop at a strawberry? We wanted to know if we could get DNA out of a sausage, and whether we could read the sequence of genetic letters - or bases - in that DNA to work out what meat was in there. We put geneticist Stevie Bain on the case...
Stevie - I’m looking to buy some sausages. I’ll go for a couple of the pork ones please.
So the butcher claims that these sausages are 100% pork, but can we test this using DNA. I head to the Pathology Lab at the University of Cambridge where Dr Ed Farnell will help me figure out exactly what’s in these sausages. The first thing we have to do is extract the DNA from the sausage and to do this we use essentially the same techniques described in the strawberry experiment… albeit with slightly more expensive chemicals and fancier equipment…
Ed - We’ve got our DNA extracted now and we’re going to start preparing that DNA for sequencing. But instead of sequencing the entire genome of the pig and whatever else might be in the sausages, we’re going to focus on a small region which acts as a barcode for each animal. Now the way we’re going to target that region is using primers and these are short stretches of DNA which target our enzymes, and then the enzymes are going to make lots and lots of copies of it. Then that will allow us to make a library which we can put on the sequencer and get the sequence of those regions.
Stevie - To generate many copies of DNA fragments in this way, we require a technique called the Polymerase Chain Reaction or PCR which uses an enzyme called DNA Polymerase…
Ed - To amplify DNA by Polymerase Chain Reaction or PCR, you need to slip the two DNA strands. You do that be heating them up to 98 degrees celsius and that splits them in two. And then we cool it down to what we call our annealing temperature, which here is going to be 63 degrees celsius; that allows the primer to specifically stick to the bit of the chain. Then once it’s stuck we heat it back up to 72 degrees; that then allows the polymerase to come in and make a copy of the little region that we’re interested in.
Stevie - The PCR process takes around one hour to generate millions of copies of our DNA fragment. Once that’s done, a liquid containing special tags is then added to the DNA. These tags are important because they mark each of the DNA bases A, T, C, and G with a different colour which allows them to be read by a machine called a DNA Sequencer.
Ed - I’m stood in front of the machine right now, and it’s got a screen on the front and that’s where we do our interaction from. It’s touch screen so it’s really, really simple so I’m going to hit the nice big friendly button that says “sequence”. At that point, it’s going to grab our library that we’ve made which is basically our little copied bit of copied DNA with these tags on. Then you have a computer and a piece of software at the back that takes a picture every cycle and then reassembles that into a string of DNA letters. Off we go…

29:35 - What's the oldest DNA sequence we can read?
What's the oldest DNA sequence we can read?
with Eske Willerslev, University of Cambridge
Now our sausage DNA is “fresh”, but we can also use similar techniques to extract and read the genetic sequence of DNA samples that are far older. Eske Willerslev from the University of Cambridge is a pioneer in this area and explained to Chris Smith how far back in time DNA technology can take us...
Eske - I think my team still have the record for the oldest genome and that would be from a 700 thousand-year-old horse. But my prediction will be that we can even go beyond that, beyond 1 million years in some cases.
Chris - What sort of condition is the DNA in in things that are more than half a million to a million years old then?
Eske - It’s extremely fragmented so when you retrieve DNA from these ancient samples it’s very often that it’s no longer than a hundred base pairs. In comparison, if you take DNA from you and me for example, we would see stretches that would be millions of base pairs long. So it’s getting highly fragmented and there’s also other modifications to the DNA that can result in incorporation of the wrong bases during amplification that you just described.
Chris - Can one think of it a bit like a jigsaw, I suppose? And in me the jigsaw is intact, you can see the whole picture, but in a 700 thousand-year-old bone remnant or something, it’s a bit like someone’s shaken the jigsaw box and broken up the jigsaw into all the little pieces, so in assembling the genome back together you’d have to try and work out where each of the individual pieces go to make that picture?
Eske - Exactly. It’s like a major puzzle right. So you have these very short fragments of DNA that you then have to assemble into genome and to do this you normally use a reference genome. If you’re sequencing DNA from an ancient human, for example, using one of the human reference genomes and then putting piece by piece these small fragments that you have sequenced in place.
Chris - And you’re doing that painstaking working out where each of the fragments go, you’re doing that with a computer because it can basically look at millions of possible arrangements and combinations at a time and do what would take us millions of years to do?
Eske - Exactly, it’s through computer programming. But it’s quite surprising that with fragments down to 35 base pairs, I think it’s in the range of 60% of the human genome you can map uniquely with these small fragments.
Chris - How do you actually go about getting DNA out of these ancient specimens and why have we only gone back a million years? We’ve got lots and lots of things which are much older than that sitting in museums, haven’t we?
Eske - Yeah we do. One thing we discovered more recently is that there are certain parts of the skeleton, if it’s a skeleton that you’re looking at, where the DNA is better preserved. For example, the petrous bone which is part of the inner ear is the most dense bone in the body, and the DNA preservation seems to be very good there compared to other places. Also in teeth, you also find good DNA preservation compared to other places. But, of course, with time and also very dependent on what type of environment you’re dealing with, the DNA would finally get so fragmented that it’s actually impossible to retrieve any information out of it.
Chris - Are we at a stage now where with people like you able to get DNA from smaller and smaller specimens that are older and older, that sometimes we find the genetic image of a fossil, something that existed historically before we actually find the thing itself? So we know we’ve got a genetic signature for it but now we’ve got to go and find the thing that we’ve found the genetics for?
Eske - It’s correct - that’s not uncommon. You can say that we find genetic sequence that doesn’t match any existing sequence and, therefore, it derives most likely from some kind of extinct species that maybe hasn’t been described yet, or at least hasn’t been sequenced yet. But in some cases you also have material that has been described morphologically but hasn’t been sequenced yet and, therefore, you get a sequence that doesn’t match anything. It’s hard to say whether it’s one or the other.
Chris - What can we learn thinking about humans and our origins? What can we now learn and what’s emerging by using the sorts of techniques that you’re developing and applying them to human evolution?
Eske - You can study. You can say the biological history of humans. There’s a number of things that have emerged over the last few years but one thing that has become increasingly clear is that modern humans have migrated over long distances, even from the very early times. Spread all over the landscape and have met each other, mixing with each other at various timepoints. It also makes, for example, the whole idea of races kind of ridiculous because groups have been meeting, separating, meeting again since as far back in time as we can measure.
Chris - Can you infer anything about the social structure of humans because we’re a very social species aren’t we? Can the association of certain clusters of genes or gene sequences, which might be in certain populations, tell us something about who was mating with whom back in evolutionary time?
Eske - Exactly. We actually had a paper in Science this week on this issue here where we tried to understand did early modern humans actually understand the concept of inbreeding and outbreeding? Of course, if you get inbred you get all kinds of diseases and it’s very bad for the population. In order to do this we sequenced a number of individuals from the same locality 34,000 years back in time and, to our great surprise, these individuals which are coming from a very small group of people doesn’t seem to be inbred. They don’t seem to be as closely related as most people would have suggested. This really tells that at least in the upper palaeolithic, 34,000 years ago, people had an understanding of the need of getting mates from outside the group itself, in that sense avoiding inbreeding...
36:09 - Genetic fingerprints from viruses
Genetic fingerprints from viruses
with Andrea McCollum, US Centres for Disease Control and Prevention
The infections we succumb to, caused by agents like viruses leave genetic fingerprints in their victims. And even though the virus itself may be dead, the sequences of its genetic material can still be obtained from various tissues and specimens and can reveal crucial insights into where those infections came from, and how they evolved. This, in turn, tells us how infections we might encounter in the future may behave. Andrea McCollum works with the US CDC - the Centres for Disease Control, and Izzie Clarke spoke to him about his work...
Andrea - We have a couple of interests: one would be for the smallpox virus itself and how that evolved over time. Then we are also really interested in viruses that were used to vaccinate people against smallpox.
If we can begin to take these old specimens and then fill in gaps in our knowledge, we might better understand what was used, for example, in the United States during the 1800s. And then also for modern-day, it would be very nice if we could determine how these viruses have changed. Have they changed in a way that have affected their ability to infect human cells and to cause disease in humans? Have their infectious properties actually changed to be less infectious over time?
So there’s a lot of really interesting questions that we have that these types of specimens help us begin to fill in those holes and answer the questions.
Izzie - Smallpox has been eradicated, so how exactly can you investigate this?
Andrea - We’re very dependent on looking at older specimens. Some of these older specimens allow us to look at poxviruses, including smallpox, that were present in the 19th century, in the 18th century, and perhaps even further back in time from then. For example, I was initially involved in 2011, we received a call from someone who had visited a museum and had seen what was labelled as a smallpox scab. This actually was a scab that was sent in an envelope with a letter, and it was sent from father to son across the state in Virginia, and it was just after the civil war, so it was around the 1870s. It became clear from reading the letter this was a vaccination scab. The idea is that this scab containing live virus that had been used for vaccination was collected from somebody was being sent to this family member to then be able to vaccinate others.
Izzie - You’re looking at the genetic information of these viruses, does the virus have to be live to take that genetic information and have a look at it?
Andrea - No, it does not. The nice thing about these viruses, they are DNA viruses. Even if the virus itself is dead in the sense that it cannot cause an infection in humans or in other animals, you may still be able to pull out some of that genetic material and then look at it in the laboratory, obtain the DNA sequence and then find out what virus it was. That is one thing we’ve been able to do with the variety of these specimens like the scab that I mentioned in Virginia. Or there have been specimens recently obtained from corpses that have come out of permafrost in Siberia. You can learn a lot about that virus that you’ve uncovered without the virus being alive.
Izzie - What can this genetic information tell us about a virus?
Andrea - One thing that we would immediately be interested in if we found any of these poxviruses is 1) what is the species? Or is it a combination of species of viruses? Then we want to look gene by gene to see are the genes the same as what we know exist in the viruses that exist today. Then within the genes, you can look at the specific sequence information that’s there to determine has there been a lot of evolution over time? What specific sequences have changed? And maybe come up with hypotheses for why those changes occurred.
Izzie - Can this help us in our understanding of modern-day viruses - we’ve seen a lot with Zika or Ebola?
Andrea - Definitely. You definitely need to do that and so I think it can help us in a variety of ways. It can, again, help us understand how viruses have changed over time which may have implications with the pathology or the ability for these viruses to infect human cells, or not. And I think what’s really nice about some of these older relics is that if you can combine it with documented information. For example, in Virginia with the scab, it was found in a letter, so we had documented information about how it was obtained and then it was used for vaccination. If the specimen came from an actual patient and you have some information about the clinical presentation of that patient, and what the patient experienced as a result of that infection, I think it can help us understand a lot about the ability for these viruses to cause disease. How these viruses are interacting with humans today. And, of course, there’s a lot of really interesting historical information as well.
Izzie - Should we be concerned about these live viruses and their genetic information entering modern society?
Andrea - I think first we definitely need to determine when we find one of these relics is there live virus or not? To date, we have not been able to find live virus. But certainly, that is a concern and that’s something I think we want to determine very quickly and very efficiently. One, for the safety again of the individuals who find these specimens and work with them. And then also we want to make sure that this is not spread further.

42:12 - The Results: What's in a sausage?
The Results: What's in a sausage?
with Ed Farnell, University of Cambridge
Remember the sausage we were genetically sequencing earlier? Let’s head back to the Cambridge Pathology lab where, joined by the butcher who made the sausage, Stevie Bain and Ed Farnell reveal what they’ve found...
Ed - What we’re looking at is a pie chart which shows the amount of reads from each animal as a percentage. Each individual read is a bit of the conserved region that the sequence sequenced.
Stevie - The percentage of reads we say okay, we know that this specific sequence looks like this in the cull. How many kinds is that present?
Ed - Exactly. I guess the good new is your sausage is 98.61% - according to this - pig, which is pretty good mostly. It’s pretty much all pig and then we’ve got a few trace amounts of cow or beef. A little bit of sheep, a little bit of chicken, and a trace amount of human as well.
Stevie - Hold on a second! You’ve found human DNA?
Ed - Yes.
Stevie - In some sausages.
Ed - We mix them up by hand so it could be just a tiny amount of a trace that wa, and also handling them. The way that we’ve prepared it in the lab as well, there's DNa in dust, there’s DNA on the bench surfaces.
Stevie - Going back to some of the other things that we found in there. A little bit of cow in there. We have 0.26% sheep, so that’s the percentage of read. How many times, as a percentage does the sheep genetic code show up in the sausage?
Ed - In our sequencing data, exactly.
Stevie - But that doesn’t necessarily mean that 0.26% of the sausage is sheep?
Ed - Exactly. And there’s a couple of reasons why that might be. One is that we do amplify the region and we might get some biases where some sequences are more amplified than others just because of the type of sequence they are. The other issue we might have as well is that the mitochondria, which we’re actually sequencing, they’re in the muscles of these animals which is what goes into our sausages. And one of the interesting things is across all the different species, the pig, the cow, and the sheep, there are actually lots of different amounts of mitochondria. Say, for example, in a pig, in a single pig cell you might have a hundred mitochondria. Whereas, in a single cow cell you might have a thousand mitochondria or visa versa. So that can also bias how accurate our classification is.
Stevie - And that’s why this type of test it tells us what DNA is present in there but then the Food Standards Agency would use a more accurate test to quantify exactly how much of each animal is in the meat?
Ed - Exactly, yeah.
Stevie - So certainly high quality sausages then and no need to be concerned about those traces of human?
Ed - No, definitely not.
Stevie - Because I guess that’s the amazing thing about DNA, we’re leaving a tracing of ourselves touching the sample or chopping up the sausage it’s just going to show up?
Ed - The knife used to cut it - anything. Because we have sequencing data and we can actually see the variation in the different bits of sequence we get. We did find something quite interesting actually and what I wanted to ask you is were these sausages all made from one pig?
Butcher - No, they’re made from probably I would imagine two or three.
Ed - So that’s really cool because when we looked at the sequence data what we could see was probably three or four different pigs in there, just from the genetic variations. That’s really cool, so it matches up with what you guys do when you make the sausages.
Stevie - How do you feel about the results?
Butcher - I’m relieved, to be honest. Yes, my boss will be pleased, put it that way.
Stevie - Well, now that that’s sorted, the only thing left to do is cook them up and tuck in.
46:06 - Improving medicine with 100,000 genomes
Improving medicine with 100,000 genomes
with David Bentley, Illumina
David Bentley, who’s the Chief Scientist at the company Illumina, joined Chris Smith in the studio. Illumina is based in Cambridgeshire, they developed the system that we used to read our sausage DNA, and they’re the official partners of the NHS 100,000 genome project. This is an ambitious study unveiled in 2012 by then prime minister David Cameron to read the genetic sequences of thousands of National Health Service patients. David told Chris about the project and how Illumina is involved in delivering it.
David - This project is the first of it’s kind to really examine a large population of patients - 100,000 genomes from around 70,000 patients. And to look in particular at how best to utilise an entire genome sequence in clinical practice; what we can learn from it; what challenges we face; how to handle the information, and how, ultimately, to really make a difference to what is being called precision medicine, or genomic medicine. To give a better answer for each patient because we know about the patient through their genome, and we know more about the disease they may be suffering.
Chris - We did a bit of a sausage in a matter of hours. For those who may be not so familiar with your technology and the technique, how much more of a problem is it to read the entire genetic code of a human in the way you’re proposing to do?
David - The whole human genome of a person is much larger than some of the samples you’ve been talking about. Some 3 billion letters or bases of the genome in every person. And the first human genome, the reference human genome that was talked about earlier that took some 7 years to sequence and an international team to do it.
The technology revolution which spun out of Cambridge here actually allows us to do a genome in about two days. In fact, it allows us to do several genomes in two or three days and that, therefore, really makes it possible to access and provide this information for the benefit of patients in timeframes that are clinically relevant. The doctor can really ask the question and make use of the information, and we return it back in the time that’s really needed.
It’s an ongoing process; it’s accelerating as we go forward and it’s important to say that doing 100,000 genomes is a really big undertaking. It involves a partnership, not just with the NHS but, very importantly, with Genomics England who are set up specifically to manage the whole project, and they partner with us. And they have had to deal with the challenges, not just of how fast can Illumina sequence the genomes? It’s also how can they recruit patients, provide consent, collect samples, return the data, analyse, interpret them, and really get them back to the doctor? That whole process is being tackled straight away at the level of 100,000 over the 3-4 year duration of the project.
Chris - We’re calling this the 100,000 genome project, but there is a subtlety in there because you then said “well it’s from 70,000 people.” So why is there that distinction; why have you got 70,000 people and 100,000 genomes?
David - The project is really addressing two of the forerunner challenges that the genome really hopes to make a big impact on. One is a rare genetic disease which often hits newborns and paediatric conditions. In that case, we really look to try to sequence a child that’s affected and also their parents - that’s three genomes for one disease.
The other important area, of course, is cancer. Cancer is also a disease of the genome but, in this case, it’s the DNA in the cancer which has changed over a period of time. Either through exposure to toxins or radiation, or through other events which have really given rise to changes since the person’s genome was given to them by their parents. Here we’re sequencing that tumour DNA to look at the changes that happened since the germline, since birth. But we’re also sequencing the normal from the same patients so that we can distinguish the differences and really pinpoint what it is that’s happening in the cancer that is not happening in the cells in the rest of the body. That really gives us the clues for what’s caused this cancer; what the person’s been exposed to and, hopefully how we can manage the condition or provide treatment.
Chris - But scientists all over the world have been sequencing genes, and tumours, and patients for a long time, so what does doing this en-mass, in the way you’re doing it, add that other people can’t do and replicate so easily?
David - It’s important to recognise the importance of all those other studies because all that information folds into the present studies. But the key about sequencing the whole genome in one shot, in two days is the possibility gather all the information that might be required. The answer is in there somewhere if you have the whole genome, you just have to know how to find it. You can find the needle in the haystack, but you do at least have the whole haystack to look at quickly and efficiently. We can develop methods, we are developing methods to analyse ever more quickly and precisely, searching through that haystack in a very automated way to look for causes of disease for these particular individuals and provide an answer quickly.
Chris - And how are you getting on; are you on schedule?
David - The programme is going very well. It’s an enormous project as I’ve described. The path from the patient in the clinic right through to the sequencing at Illumina, and back out to Genomic England, and back to the doctor takes some time. We’ve actually done 35,000 genomes so far. The majority of them in the last 12 to 18 months, and we are accelerating the programme all the time as we learn how better to do it. And the teams all over the country from Genomics England to the genomic medical centres, to the research centres, to the data centre are all working together to really decide how best to handle the information quickly and efficiently.
Chris - Just very briefly, David, people often worry when we talk about data, personal data, about data security, so how do you keep a person’s genetic fingerprint safe in your hands or in the projects' hands?
David - It’s important to realise a great deal of effort has gone into the security of individual’s data and their DNA. The chain of custody goes all the way from the doctor and the patient who has consented to be part of the process and has it explained to them. And even by the time the DNA gets to us, it’s been anonymised, so the identity of the person is protected really close to the source and Genomics England take care of that through a high-security data facility. They also store the clinical information downstream to enable people to be secure as they can possibly feel knowing that Genomics England really has the custody of that important privacy and confidentiality of the project.

52:28 - What's the best technique to swat a fly?
What's the best technique to swat a fly?
Michael Wheeler’s been buzzing about this question from John. So without winging an answer to this one ourselves, we put it to our listeners and Ian from Melbourne, Australia has discovered that confusing the fly with a clap of the hands makes the job easier. But what does our expert think - here is animal visual specialist, Kate Feller, from the University of Cambridge…
Kate - After discussing this question with several researchers at an animal vision conference in Finland, literally as naked scientists in the sauna, we all agree. Yes, this is possible.
Because fly motion vision is processed very fast, you can theoretically trick the fly by just moving very slowly. How fast the edges of your hand expand relative to the fly’s vision is what trigger is to flee, so a slow hand could confuse the fly. However, because the fly motion vision is so sensitive you would have to move so slowly that either you get bored and give up, or the fly just takes off. Because, you know, flies don’t stay in once place for very long.
Some alternative methods are to hold perfectly still and watch the fly until it starts washing itself, then strike quickly while it’s distracted, just like jumping him in the shower. You can also approach the fly at a normal speed and, instead of slapping it, clap your hand just above the fly to intercept it as it takes off.
Michael - The old “trap and clap” method. This one can even “be done on the fly”…
Kate - One other solution that a colleague recommended, and I have personally yet to try, is to spread your fingers wide on the same surface as the fly and pull your middle finger back like a slingshot while you slowly slide your hand towards the fly and quickly release the cocked finger once the bug is in range. Since the fly is being approached from multiple directions it should be confused enough to not take off as it normally to an approaching hand, so you can effectively squish it when you release your middle finger. I’ll definitely be experimenting with this method.
Michael - So, to any flies out there, beware the naked scientists.
Next week, We’ll be beaming out an answer to this question from Jayson:
Jayson - Our new house is 140 metres from a cellphone tower. As a family, the three of us feel like we have been affected to different degrees in terms of sleep, motivation, and anxiety, which are commonly reported symptoms of exposure to microwave radiation. It’s a controversial topic, but are there any major health risks with living close to a phone tower?
Thank you to Nebulousflynn for the fly sound effects.










Comments
Add a comment