Phenomics: A Medical Revolution
This week, we’re looking at the future of medicine, phenomics! Including the toilet that analyses what you put down it! And in the news, sending wine to space, new insights into the origin of life, and why a lack of sleep gives you food cravings!
In this episode
00:50 - High frequencies key to understanding speech
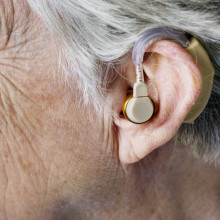
High frequencies key to understanding speech
Lina Motlagh Zadeh & David Moore, Cincinnati Children’s Hospital, Ohio
Do you have trouble understanding what others are saying in noisy places? If so, new research out this week might explain why. Researchers in the US have found that high frequency sounds play a key role in the intelligibility of speech. And if you can’t hear them properly, you struggle in noisy places. But these critical frequencies are usually not routinely checked in hearing tests, and perhaps they should be, as Adam Murphy heard from Cincinnati Children’s Hospital, Ohio’s Lina Motlagh Zadeh and David Moore…
David - So in a normal audiology exam, they generally check out frequencies between about 250 Hertz and over 4000 or 8000 Hertz - where 250 Hertz is low pitched sound and 4-8000 are very high pitched. But actually healthy young people can hear up to twenty thousand hertz (20 kilohertz), but audiologists have tended not to pay attention to that extended high frequency between eight and 20 kilohertz partly because most of the energy of sound is at lower frequencies. We know from everyday experiences of using telephones and so forth we don't actually need to use those extended high frequencies, at least in normal listening conditions.
Adam - Lina now that we've set the groundwork with David, can you explain to me the experiment you've run?
Lina - We recruited young adults, more than 60 percent of them less than 30 years old and we surprisingly found more than half of them have extended high frequency hearing loss, which was related to the self reported difficulty listening to speech in noisy environment. So we first started with routine ideology tests, this involves hearing tests. When you go to visit an audiologist you send some pure tones with different pitches and intensities and you ask them to respond to the sound even if it's soft to obtain the minimum amount of things that they can hear. Then after that we used the 'speech in noise test', we wanted to see if we give access to high frequency hearings, very high frequency hearings, how the speech understanding will be for this participant. So, surprisingly when we put the cutoff of the filter giving access to frequency above 8 kilohertz their score improved significantly. So, it shows that listeners could use these frequencies to understand the speech in noise.
Adam - and why is it that not having these high frequencies means you can't pick noise out of crowds so well?
Lina - Understanding the speech mainly depends on high frequencies, because high frequencies can provide intelligibility or clarity and we need to hear high frequency parts of the speech to follow the conversation in noisy environments. Then you have high frequency hearing loss as a result of age noise or a lot of toxic drugs, this mainly affects high frequency range of hearing - you can't easily follow speech like a person who has normal hearing in these frequencies.
Adam - What do you want people to take away from this research?
David - I think this gives us an opportunity for some sort of personalised medicine, because it's important to realise that some people have this problem while others don't. We can then tailor our intervention, be it protection of hearing, or even some new drugs which are coming online soon to prevent hearing loss to the people who really need them.

04:48 - How the first cells formed in deep sea vents
How the first cells formed in deep sea vents
Sean Jordan, UCL
It's a question no one's tired of asking - where did we come from, and how did life begin? One theory is that deep sea vents, where mineral-rich warm water issues from the planet’s interior, played a key role; here, the conditions could have been just right to allow simpler molecules to link up and form the crucial oil-based membranes that enclose cells. But nice though that theory was, no one had managed to prove that it was possible. Until now, that is. Because, by recreating conditions similar to a subsea vent in his lab, UCL's Sean Jordan has made it work. He told Katie Haylor how, and why these vents are so critical to the process...
Sean - So, they're really unique in that they're formed with an alkaline fluid and four and a half billion years ago when we think first life would have emerged, you have an acidic ocean and alkaline fluid inside a vent and it's almost like a battery - so a positive charge on one side negative on the other. This provides the energy that would allow you to create the first organic molecules and then you can go through stages of more complex chemistry. We can concentrate molecules inside of the vents because of their internal structure and you can form these non-living cells that eventually become living cells.
Katie - We've thought about hydrothermal vents and origin of life for a little while now. So, what was the specific problem you were trying to solve with this study?
Sean - There are many theories for the origin of life and alkaline hydrothermal vents have a lot going for them. In previous research no one has been able to form these cells under alkaline, salty conditions. So that's a big hit for this theory because if all life is cellular, we were able to form them for the first time and we think it's a being kind of boost for this theory.
Katie - How on earth do you make what came before the cell. What is that?
Sean - Modern cells are formed with phospholipids. So these are quite large molecules, they have tails that are hydrophobic which means they don't like water and heads that are hydrophilic, which do like water. When you put them in solution the hydrophobic tails connect, the hydrophilic head groups point outwards and then they form this sphere - looks like a soap bubble. We can see them on a microscope they look like a fatty sphere!
Katie - Are these just really, really, simple cells. How would these compare to a normal animal or plant cell, that we might see today?
Sean - Yeah. So they're super simple. So we've composed these using 14 lipids, so fatty acids, alcohols, and isoprenoids. A modern cell would be composed of many different types of phospholipids, so again larger molecules but you would also have proteins in there as well. So, these are all different molecules that would make up a cell membrane and cell membranes in modern cells are completely active in working with metabolism - allowing things in and out. What these simple cells are like, they're actually quite leaky, so that's good because when you don't have an active way of passing things across the membrane. If the cell itself is leaky, things can get across without needing to do any work essentially. But if we can get simple enzymes like protein, into this membrane then it can start to play a role in metabolism.
Katie - Okay. That might take us a bit closer to the cells that we know and love today.
Sean - Exactly.
Katie - How did you manage then to make these cellular precursors in the conditions that you have, that people haven't been able to do before.
Sean - The reason that we think that people haven't been able to do it before, is because the approach that's been taken is a kind of standard chemistry approach where everything is kept really clean. They were using one, two, and three molecules to make cells putting them under really harsh environmental conditions of high temperatures, varying P.H and different salts. These simple cells they don't like it, but the good thing is that at the origin of life you would have had close to a hundred of these types of molecules. So, we thought let's make things a bit more messy a bit more realistic! So we just used 14 different lipids and we mixed them together and they were able to be stable under these conditions.
Katie - Ok, so you've got a bigger variety of starting components as it were. What about the actual conditions? Because these vents are hot, alkaliney and salty - right?
Sean - A lot of people confuse them with these black smoker vents, with violent black smoke coming out. They're around three hundred and fifty degrees, it's much more difficult to try and form any sort of life in those but what alkaline vents have, you're talking around 50 to 100 degrees - we used 70 degrees. Seawater concentrations of salts because that's the best approximation we can have for what they see the ocean would have been like back then and then the alkalinity of around P.H. 11 page 12, we've used that's representative again of the fluids in the inside.
Katie - Now without getting too philosophical, these precursors aren't actually alive, right?
Sean - they're absolutely not alive.
Katie - What does being able to do this process tell us about how life may have started in the first plac?
Sean - All we can do is think about what life requires, even that in itself is a controversial topic! So, most people don't agree on what's alive and what's not, but we can take simple things, like DNA, cell membranes, metabolism and we can try and replicate each of those by themselves in the lab - then we can start to think about putting these together. If you can get all of those components to work under conditions that are representative of your theory, then that lends more weight to the idea that could have been where life emerged.

11:17 - A lack of sleep could cause the munchies
A lack of sleep could cause the munchies
Surabhi Bhutani, San Diego State University
It sounds counter-intuitive, but an extra hour or two asleep in bed can help to reduce the risk of becoming obese. Less sleep, on the other hand, seems to be a potent stimulus to over-eat, and especially to binge on high-calorie, fatty and sugary treats. But why is this? A new study by Surabhi Bhutani, from San Diego State University, suggests that sleep deprivation leads to a surge in the body’s own cannabis-like chemicals. These, she’s found, cause a region of the brain called the insula, which controls food intake, to slacken its inhibitory grip on the brain’s smell areas, making the aroma of delicious treats just too tempting to resist, as Chris Smith discovered…
Surabhi - There is a huge body of research that suggests that chronic lack of sleep is associated with overall poor health and there is a bunch of data showing that when you do not get enough sleep you increase your food intake, and people become more reactive to unhealthy foods and foods in particular that are high in sugar and fat, that we call junk food. What we really wanted to understand was why people crave these high fat foods after a sleepless night.
Chris - Back in the past, when people first began to flush out this association between not getting enough sleep and then rebound overeating, one speculation was that the hunger hormone “ghrelin” - which is produced by the stomach and is suppressed by sleep - that goes up. So there's just a rebound overeating to compensate. So is it as simple as that?
Surabhi - It's more complicated than just hunger hormones increasing, because there are a lot of studies showing that people may not really physically feel hungry, but they still go for all those foods that are high in calories. So there has to be a different mechanism where, basically, it connects your sleep loss with consumption of very high calorie foods; so your brain, or your body, saying that I really want a doughnut, or I really want potato chips!
Chris - So you're saying that there's a switch in terms of food choices but it's not necessarily just driven by overall increase in hunger?
Surabhi - Exactly.
Chris - And what do you think underpins that then?
Surabhi - We definitely think that there are some brain signals that may be playing a role in overeating of not-so-healthy foods, and past research primarily has shown that sleep deprivation increases certain endocannabinoids. So these endocannabinoids are basically these naturally produced neurotransmitters that bind to some of the receptors in the brain and affect feeding behaviour. So they're very similar to cannabis-like compounds that can cause cannabis-related munchies. On the other hand, we also kind of know that sense of smell is also really tightly related to how we choose food items and, in particular, animal studies have shown that these endocannabinoids enhance food intake by increasing the activity of brain areas that process odours. So what we thought was that, maybe we can put all of this together and ask if what people choose to eat when they are sleep deprived is related to how the brain responds to food smells.
But what we found in our study was that, when people were sleep deprived, so they only slept for four hours, the following day when we scanned their brains and made them smell these delicious food odours and also some of the non-food odours, the piriform cortex - the region of the brain where smells are processed - in that particular region the patterns of food versus non-food odours were significantly different. So what this means in simple terms is the smell processing region in the brain goes into this “hyperdrive” - it sharpens the food odours for the brain so it can better differentiate between food and non-food odours.
Chris - And how do you tie that to changes in the endocannabinoid system, these natural brain chemicals that mimic cannabis?
Surabhi - The piriform cortex also sends signals or information out to other brain regions, in particular insula cortex. So insula receives signals that are important for food intake, and when a person is sleep-deprived, signaling between the piriform cortex - the smell processing region - and the insula, that connection was not as strong. So the signaling actually reduced. And we also found that, because of this reduction in communication, people ended up eating more energy-dense food. Now, how is it connected to the endocannabinoids, or the neurotransmitters? When we did the blood analysis, we saw that people had certain components of this endocannabinoid system very high in the blood. And those people also consumed very high energy-density food. So, putting all this together, our results suggest that the sleep deprivation really influences this endocannabinoid system, which in turn alters this connection between piriform cortex and insula cortex and, ultimately, leads to a shift towards foods which are high in calories.

16:41 - Bottles of Bordeaux bound for space
Bottles of Bordeaux bound for space
Claire Bryant, University of Cambridge
From dinner to drinks now, and the announcement that wine buffs are sending a dozen bottles - not of "rie-slinghot", or even "star-donnay" - but Bordeaux into space! Regrettably for them, the astronauts won’t be allowed to drink it! To find out why on Earth, or more accurately not on Earth, you’d want to do that, Chris Smith spoke with scientist and wine expert Clare Bryant...
Clare - So, the space agency NASA are actually sending the wine into space! There's a guy from the Bordeaux region who has a close association with them so they've decided to send a case of Bordeaux into space.
Chris - Why?
Clare - That's a very good question and I'm not sure I have an answer to it. The spiel they're giving us is that it's an experiment, where they're retaining one case of the Bordeaux in Bordeaux, in a perfectly temperature controlled cellar and they're sending the other case into space to be kept for a year on the space station.
Chris - As a case control study?
Clare - A case control study, where we stored it at 18 degrees in space then they'll bring it back and presumably analyse it.
Chris - What might happen?
Clare - This is all based around the ability to age wine. So wine ageing is a really interesting concept. Where a very good quality wine and it's only really good quality wines that will age well, will change in texture, taste, aromas and these factors happen over time. So, a long maturation of wine will eventually end up w ith a complex interesting product. So taking into space because you're changing the environment, potentially could I guess, age wine faster because everybody would love to know way of speeding up the wine aging process. You can sell a mature wine faster, because one of the things that costs money with wine that age is actually having to store it until it's ready to drink.
Chris - One aspect of this is that the wine ends up on the shelf it's it's laid down to bottle age. Could there be some impact therefore of not ending up with? There must be a gradient in the wine, where some of the heavier molecules will end up at the bottom, some of the sediment will end up on the bottom, people say it's a mark of a good wine if you get that nice sediment falling in the bottom of the wine. So it sounds like it's just gratuitous marketing doesn't it, but do you think there could be some sensible aspect to this? They might see some interesting chemistry?
Clare - Yeah, I mean that the sediment that you get is due to the tannins. So part of the aging process with tannins is that eventually they go from being long chains, to sort of aggregates of short change and they then form sort of sediment and then the sediment drops out in the wine. But I think with we're sending a case for a year in space, the kind of other factors that alter the way a wine age as well as temperature, is vibration, there's clearly going to be a vibrational process taking the case in space but also radiation. So, one of the things you do when you store wines, you store in the dark because the UV light generates free radicals and that causes problems. So I presume you're thinking about the effects potentially of gravity or microgravity, different space radiation and what that might actually do to the way in which the wine ages.
Chris - There have been booze cruises into space before - haven't there? This isn't the first in that respect?
Clare - No it's not. The Russians when they went up to the 'Mir' space station, I believe it's called, used to smuggle vodka with them they'd actually lose weight and tuck the vodka in their suits and then take it with them when they went into space.
Chris - And the moon landings? They had a tipple as well?
Clare - The moon landings took the communion wine onto the moon with them and they poured it into the glass and it kind of climbed its way out of the glass!
Chris - So would you indulge? Would you worried at all by radiation - irradiated wine?
Clare - Well interestingly there's there was a really interesting presentation done by somebody at the Royal Society chemistry. They actually talked about the effects of radiation, and what it does to wine aging. The problem is that actually instead of not actually improving the wine, it can actually make it worse because it can generate a bunch of volatile sulphurs - and that gives it a nasty smell. So I'm not so sure that this is such a good idea!

20:50 - 3000-year-old wheat genetically sequenced
3000-year-old wheat genetically sequenced
Laura Botigué, Centre for Research in Agricultural Genomics, Barcelona
If you’ve ever left a box of cereal neglected in your cupboard for months, you’ll know that it takes a long time to go off. But will it last 3,000 years? It turns out the answer’s yes… or at least well enough to still read its genes, as scientists have just discovered, using grain harvested three more than 3 millennia ago. Phil Sansom…
Phil - This all started with an expedition to Egypt in the 1920s. There, English archaeologists uncovered some tiny grains of emmer wheat exquisitely preserved.
Laura - This was one of the first wheats that were domesticated in the Near East - around 10000 years ago.
Phil - The grains then moved to London to the Petrie museum of Egyptian archaeology. They sat there for 100 years until a scientists from UCL saw them featured on a BBC documentary. He and his colleagues got permission from the museum to try something new. Carbon date the wheat and then analyse its DNA.
Laura - We sequenced its genome of these like three thousand year old sample from Egypt. So it was very exciting to use a sample that had been in a museum and find nail results.
Phil - That is Laura Botigué, a geneticist who was part of a team sequencing the wheat samples.
Laura - What we sequenced were the husks, which are some sort of leaves that usually enclose the seed. So if you imagine a grain, a wheat grain that has this slightly around its shape which this beautiful golden colour.
Phil - Pairs of leaves each curled around the seed that had long since been eaten away by bugs.
Laura - The first step is to remove all the protein to free the DNA. You are in the blind when you do that because you follow the protocol pretty much like a recipe but you do not know the result until you sequence it. It's a bit challenging.
Phil - It works though and they produce the full genome of the ancient wheat. Every single one of its genes, they found that it had many of the helpful traits that crops have today, like big seeds that stayed on the plant when it's right. But the scientists main goal was to investigate how domesticated emmer wheat spread around the world. To do that they matched modern types of wheat, what are called 'land races' to the ancient genome to see which were most closely related.
Laura - At least the middle was surprising that we found that the closest relatives where land races that are or were cultivated in the Arabian Peninsula and in India and not the samples that used to be cultivated in the Mediterranean.
Phil - Why is that surprising. Well it's all about how early crop technology spread around the world.
Laura - This was a major transfusion in human history. It triggered a lot of changes in how populations lived and interactions between societies. This is sometimes called the Neolithic revolution and hunter gatherer populations were replaced by agricultural populations.
Phil - Scientists previously thought that when plants first got domesticated in places like the Middle East, the technology spread outwards in all directions at once to Europe, Africa and Asia. But this emmer wheat genome from Egypt was a way closer cousin to Asian land races than European ones. Maybe that means the technology came earlier to some areas than to others.
Laura - Probably there was a first wave of expansion towards the north and then Europe, and then there was a second wave of expansion towards the south of the Mediterranean and Asia.
Phil - The question is then why would technology have moved like that.There might be a European bias going on here but there might be something more.
Laura - One of the common problems is that usually Europe is better studied than other places. If you look at the archaeological sites I think that this theory of the later replacement matches the timing of the archaeological sites that you'll find. But because it's an understudied area (African nations) compared to Europe. I would say that at least it's intriguing to know why this happened at a later stage.
Phil - And the other big conclusion of the study is that lots of samples in museums might actually be treasure troves of DNA - even if they're not exactly going to make a healthy and nutritious breakfast.

Mailbox: Blindsight
We had Tony, one of our listeners in touch about a question in our recent QnA. One of our experts Keziah Latham was talking about what blind people actually see, especially give that 95% of people who are legally blind still have some visual perception. But Tony wanted to know more. Say they were born blind but with a working visual corext in the brain. Well, we went back to Keziah, who said...
Chris - The questioner is right to suggest that processing of vision in the eye and in the brain is not always exactly the same. One example of this is that it is relatively common for people with acquired sight loss (due to ocular problems, but with a working visual cortex) to experience visual hallucinations in their non-seeing areas of vision. This is called Charles Bonnet Syndrome. It is thought to be due to visual cortex being ‘bored’ by no longer receiving input from the eyes, and neural excitation creating images that the person can see. These hallucinations can be upsetting to people, and anyone who experiences visual hallucinations related to sight loss is encouraged to contact Esme’s Umbrella (https://www.charlesbonnetsyndrome.uk/) for advice and support.
Adam - On the other hand, some people who are blind due to cortical problems, but have no problems with the eyes themselves, appear to be able to respond to visual stimuli that they do not consciously see, which is termed ‘blindsight’. This can occur after stroke, which typically affects one side of the brain and thus one side of vision. People with blindsight can respond to stimuli presented in their blind field with an accuracy greater than chance, despite being unaware of seeing any visual stimuli. This suggests that some visual processing bypasses the usual visual pathway to the occipital cortex.
However, if someone is born with no sight, their occipital cortex will not develop the ability to ‘see’ – input from the eye to the brain is needed in the first years of life to fine tune visual processing ability through experience. If this is lacking, then the eye / pathway becomes amblyopic or ‘lazy’.
Chris - However, Tony Morland at York University has shown that there is some plasticity in the visual cortex, in that people born with only one type of photoreceptor in the eye can show signs of remapping of the visual cortex in compensation. This plasticity does not seem to occur if loss occurs later in life.
The power of phenomics
Sam Virtue, University of Cambridge, Jeremy Nicholson, Australian National Phenome Centre, Murdoch University
Last month a very special multi-million dollar facility opened at Murdoch University in Perth, Western Australia. It’s the Australian National Phenome Centre. It’s staffed by an international team, many of them recruited from the UK, and their aim is to do for biochemistry and human health what the Large Hadron Collider has done for physics, and potentially revolutionise the way we do medicine. The basic idea is to use very high-end analytical technology to sift through the thousands of different molecules in a human body to spot patterns - or fingerprint changes - in the levels of those different chemicals that predict diseases that a person hasn’t got yet but will go on to develop in years to come, as Adam Murphy and Chris Smith heard from Sam Virtue and Jeremy Nicholson...
Sam - It's important to realise that virtually every disease we now look at will have some component of metabolism to it. Metabolism is basically all of the biochemical processes that go on in our body to enable us to both live and grow. So we can think about, very basically, when we eat a meal there will be a whole series of biochemical processes that will go on. The food that we get into our body will be broken down and then rebuilt up into the molecules our body needs. So almost all diseases will lead to alterations in how the body processes nutrients, how it stores nutrients; and therefore, depending on how these nutrients change, we will be able to detect different metabolites. A metabolite is a breakdown product of a nutrient. So we could say, a metabolite of glucose, we could say... the ultimate one maybe is carbon dioxide and water, if we oxidise it. But there are lots of bits in between going from a glucose molecule to the carbon dioxide and water that comes out in our breath. And if we have diseases that impact on metabolism, we'll be able to detect different molecules. And those can be detected in blood, some can be detected in sweat, in urine, and also in breath.
Adam - So why does this field lay ahead of us now? Why are we now talking about smart toilets? Well, the Australian National Phenome Centre has recently opened, planning to analyse the chemicals in the bodies of millions of Australians. Chris jetted out there and got a tour of the facility and spoke to one of the people behind the whole thing, Jeremy Nicholson.
Jeremy - The work that we're going to do here will be measuring fundamental metabolic properties of humans, both in the general population and those who are patients. And the aim of that is to understand how genes and environment come together to create disease, and how that expresses itself in metabolism, so that we use that information to predict disease risks. And furthermore, when you have that sort of analytical capability to measure details, one can also use that to monitor therapeutic interventions in clinical situations to see if somebody is getting metabolically well, or getting worse, or nothing’s happened during what we call the patient journey. And we can use that type of monitoring approach to optimise therapies and to see what basically works for what people. So it's a personalised healthcare approach.
Chris - Why is this better than just reading my genome?
Jeremy - It's different to reading the genome. The genome tells quite a lot about... potentially about future disease risks, but it also tells you about particular defects related to different subtypes of disease. But most diseases have a huge environmental influence. Whether you get a disease or not will be partly depending on your genes and partly depending on how you you have your lifestyle: how well you exercise, what you eat. And the vast majority of common disease is related to gene-environment interaction. So genes are not enough to stratify patients on their own. And also, when you are looking at genomics, that's very good for classifying certain types of patients before you start the hospital journey, but your genome does not change during the hospital journey; whereas your phenome, your metabolism, physiology, does; and you can therefore use that output as a representation of the success, or not, of the therapy.
Chris - What's the strategy you're using here to actually establish the phenome of the average human in Western Australia?
Jeremy - By doing studies we can try and find out what normality is. I mean, what is normality, and what is health? A healthy profile is the target for any therapy. So what we're trying to do is take therapies in patients who are sick and see if they move you in the direction of health, that you can actually attribute to particular biochemical pathways. And therefore you can use that as a metric of responder versus non-responder, and how well that therapy is working for that person.
Chris - If one took a sort of visual analogy, if I drew a landscape, where the high and low points on that landscape correspond to how the different levels of different molecules in my body relate to each other, you'd know what the normal pattern of the landscape was for a ‘healthy person’. And if someone had Mount Everest in the middle of their landscape you'd know that something had gone wrong with a particular group of molecules. Could you therefore ask, “what do I have to change about that person's environment, their lifestyle, their diet - or do I give them a pill - in order to level Mount Everest, so their landscape resembles a healthy one”?
Jeremy - Yes. So the thing is that in your body, you hopefully are born healthy, and if you have a bad lifestyle - you drink too much, eat too much, whatever it is, don't exercise enough - you will move into a different physiological state. It’s what we call a pathophysiological state. It's not quite a disease, but it's in a different state, and it’s a state that is a pathway to disease. When you get a certain level of pathophysiology, actually abnormal physiology, it becomes very difficult to go back; and then you get a disease. And then what you're doing is treating a disease to try and eliminate that particular problem. The first part of this is trying to build a map of what human physiology looks like, which helps you understand where you need to get to for that population. The dream would be to take people through from birth, through the years, and build a map of their life for their biochemistry. And you know their background genomics. So that when somebody becomes poorly, you already know a hell of a lot about them and you would know what needs to be fixed. But furthermore, if you know those people in that detail, then you would also be able to prevent disease.
Chris - Is the ultimate aspiration, then, to - having used these very powerful analytical instruments you have here - to discover what these relationships are; you then build something which is a very small, very fast analytical device that could for instance sit in a chemist’s shop, or a doctor's office, or even a person's own bathroom?
Jeremy - Indeed so. So the trick is to know what it is to put in the device. So ultimately our discoveries within the large Phenome Centre will be translated into primary healthcare and potentially into the home with smaller devices to just measure the right things at low cost. So we do have a long term idea for the iToilet, which is where your toilet becomes intelligent and measures things that are about your health, and potentially tells you that, you know, you need a checkup at the doctor's. That would be the future. And that would have a massive change in population... not only the potential for detecting disease in population, but also potentially the way that people behave. Because people are often given advice by epidemiologists, or whatever it is; they say, you know, “don’t eat red meat, eat more fruit,” and things like that. That's fine. People are really very noncompliant about that. But if you have a machine in your toilet saying, “you're really not very well today, and for the following reasons,” you're much more likely to... or your children look as though they might have something, you are very much more likely to action that, and therefore that becomes a really major contribution to preventive medicine.
What is phenomics?
Sam Virtue, University of Cambridge
What exactly is phenomics? And what is a phenotype? To take us through the biological basics, Adam Murphy spoke to Sam Virtue, from the University of Cambridge...
Sam - Phenomics is the study of phenotypes. So you can have all sorts of different phenotypes. What we think of in terms of maybe phenomics and the phenotypes we're looking at, because we work on obesity and diabetes, would be things like the actual physical manifestation. So an obesity phenotype is how heavy you are, or maybe your BMI. A diabetes phenotype would be, do you have high blood glucose.
Adam - So if we've been understanding at a basic level phenomics, certain phenotypes, for a while now; why is something like this centre in Western Australia such a big deal?
Sam - Because with the advance of technology we've begun to be able to look at much more sophisticated phenotypes and in much more detail, and start looking at many, many, many different aspects of biology. So if we think about diabetes for an example, which is what I work on, the very first phenotypes for diabetes date back to the Egyptians, and it was the fact that people would have sweet-tasting urine. But diabetes is a complex disease with many, many different components interacting to lead to this ultimate, clinically-observable sign of high glucose. So with the proposal in Western Australia and the Phenomics Centre, what they will be able to do is not just measure the glucose - which simply tells you someone already has diabetes, and they may have had it for many years undetected - but to look at hundreds if not thousands of different molecules within the body which are all interacting as part of metabolism; to look at things like proteins, to look at things like fats, and see: are these things which can predict whether people will go on to develop diabetes? They can also then start to break apart this one overall classification of diabetes - ie, you have high glucose - into different types. And this then enables us to think about targeting, medically, different people with different phenotypes which are leading to diabetes with different medicines, and we may get better clinical outcomes.

How to study a phenome
Elaine Holmes, Sam Lodge, Australian National Phenome Centre
Phenomics involves tracking the levels of thousands of chemicals in the body to spot patterns that can predict diseases a person is at risk from. So how will these measurements be made? Elaine Holmes is an analytical chemist helping to lead the initiative at the new Australian National Phenome Centre. She took Chris Smith through how samples will be processed in the lab, including visits to researcher Sam Lodge…
Elaine - Here we are in the first stop on our sample journey. What you can see is a room, it's about your average size of a living room, and it's full of large magnets. And these magnets look... if you can imagine a giant tea urn. So each of these big tea urns contain greater than the whole of the Earth's magnetic field within the can. And so if you think in terms of the big magnets that would pull your car upwards in a scrap heap, this is way, way more powerful than those.
Chris - And that has the effect of doing what to the sample is it as it goes down inside the can?
Elaine - Well the magnetic field is pretty strong, and there are some chemicals, some atoms, that have a property we call spin. So they're like little bar magnets and they're spinning. And when you put them into a magnetic field they start to line up with the field. And if you then shoot some energy in, a radio frequency pulse, it makes these little bar magnets flip; and then as they relax again back to their relaxed position, if you like, they're emitting energy, and you pick this up. And because every chemical has different atoms, they interact with the magnetic field in a slightly different way. And these small differences we can pull apart and you end up with a series of peaks which we call our molecular fingerprint. This actually is a very quick technique. If you put your sample in you can have a spectrum, you can have your fingerprint, if you like, within five minutes.
Adam - So NMR, nuclear magnetic resonance, is a very speedy way to identify lots of molecules in a single sample and to do it very cheaply. But what sorts of things can the team look for? And what does the output from the machines actually look like? Sam Lodge.
Sam - So it results via a transformation into something called a spectrum, which has two axes. On the vertical axis will be intensity, so if there's something that's very concentrated, you have a high intensity. On the bottom axis is something called ppm; this is parts per million. This is the point at which it resonates.
Chris - Looking at this computer screen, this is an example of the sort of thing that the machine would generate. This looks like almost a sawtooth. So the height of each of those peaks corresponds to how much of the substance was in the sample that you put into the machine, and along the x-axis, these are all the different types of chemical that it's picking up.
Sam - Yeah, so it's a little bit more complicated than that. So each peak is essentially from a proton in a particular chemical environment. So one metabolite might have several different peaks, because you might get different compounds with different proton environments.
Chris - So how do you sort them all out then? How does… because that just looks like a really complicated sawtooth. How do you work out what chemicals that corresponds to?
Sam - We run standards, and these standards are one particular chemical, so we can get a chemical signature.
Chris - I see. So you run a bunch of known chemicals through, you know what pattern they would produce, and you just compare what comes from your sample to what you know it should look like, and then you can say, “oh, that's in there, that’s in there, that’s in there…” To give a practical example then, say the doctor puts me on antibiotics. At the moment the dose that we prescribe for people is just a standard dose for any adult. But my metabolism might be different to your metabolism, so the amount I'm taking might be different than the amount that you would actually need to take. Could you use this to work out how much antibiotic there is in my blood compared to, say, your blood, and therefore work out whether I'm metabolising it faster than yours, and therefore tailor my dose better?
Sam - Yes, you can exactly do that, because an MRI is actually quantitative. We can monitor any drug compound within a sample, measure the concentration, so essentially we can then tailor the dose to be perfect for that particular individual.
Chris - So it's not just your own body's own molecules, but you can look at things we put into our body from outside?
Sam - Yes. And it's not just drugs either. So we can look at different food compounds, so for example someone who eats a lot of meat would have a high amount of carnitine in their blood, in their urine. Someone that’s eaten a lot of fish will have a compound called TMAO, and that changes dependent on the time from consumption.
Chris - So it's almost like dietary forensics. You can work out whether someone's lying to you when they say they've eaten certain things, you can work out what they’ve really eaten and when they've eaten it.
Sam - Yes. Yes, we can pick up things like alcohol, caffeine; and every food has a different marker which we can identify, so we know exactly what someone's eaten.
Chris - If you're not actually actively looking for those things, will they nonetheless be present in these readouts so that you could go back and look for them later? If someone a researcher comes to you and says, “well actually Sam I'm doing a study on this substance in the blood,” and you happen to have screened a million people by then, could you go to your computer and just pull out a million people's worth of these traces, and look for that particular molecule that's interesting for that researcher?
Sam - Yes, you can. So the way the NMR data is actually stored is actually very powerful, because we can run something now or in five years time, and we can overlay them and compare them.
Adam - Sam Lodge. But what about the things that NMR can't tell you? Like substances present in only tiny amounts? Elaine Holmes again.
Elaine - So now we want to go a little bit deeper into the profile, find out a little bit more about what's in your sample, so we come to the second stage - which is our mass spectrometry laboratory.
Elaine - So this room's a little bit bigger, as you can see. And it's full of 16 different machines. Looks like a big box with a big stick coming out of the box. And these are all a type of mass spectrometer that we use to do screening. So we're trying to look at everything we can in your sample. We don't tell the machine, “I want to look at fats, I want to look at sugars.” We just put it in and we want to capture everything we can about the sample.
Chris - Why is this used and why is this different, or what does this do for you that we can't get out of the NMR machines next door?
Elaine - NMR machines are very reliable. So you can measure things very accurately. But it doesn't have the capacity to go to really, really low concentrations. Maybe for other diseases you want to look at your hormones, or things that are present in very tiny concentrations. And this is where mass spectrometry comes into its own.
Chris - And how do these machines work compared with what the NMR machines do?
Elaine - These machines still separate molecules out but they did in a slightly different way. So they're separated in two ways. The first is called chromatography, and that's where you put your blood sample or your urine sample onto a column, and different chemicals stick and different chemicals go straight through. And you can then run some liquid through this, and they'll start to bleed out of the column, but at a slightly different rate. And we can catch them as they come off. You can then separate them a little further by putting them into the mass spectrometer part, and this is really just a weighing machine. All you're looking at here is how much your molecule weighs and what its charge is.
Chris - How do you work out what the actual molecule is? Because if you just get a weight and you just get a charge, there are lots of different possible arrangements of atoms that could be that weight and that charge. So how do you sort them out?
Elaine - Well we have databases where we've looked at molecules, and standard molecules, chemicals you can buy, so we know what some are. But you don't always know what they are. In these cases, what you need to do is separate them even further so you've got a single chemical. And then you blast the chemical apart, you split it up and break it; and you look at the fragments, how much each little part of the molecule weighs. And like a jigsaw puzzle you pull them all back together, add all the weights up, to make sense of the whole picture.
Adam - Elaine Holmes.
What could phenomics mean?
Nicola Grey, Luke Wiley, Ruey Leng Loo, Australian National Phenome Centre, Murdoch University
What sorts of questions might phenomics answer? From inside our bodies to the food we eat, phenomics has the potential to change a lot of things. Chris found out more when he visited the Australian National Phenome Centre and spoke to researchers Nicola Grey, Luke Wiley and Ruey Leng Loo. And to finish, a few closing words from Jeremy Nicholson...
Nicola - So in terms of using these state of the art platforms, we'll be looking initially to develop methodologies, making sure that those methodologies are very robust so that we can use them not only now but in years to come, and they'll be able to run many many thousands of samples and always give us the same reliable data that we need. We rely on that information to then make our inferences as to how a disease is progressing in a particular population. We're also looking at the mechanisms behind why particular diseases occur, why particular populations are more susceptible to developing a disease, and we'll be looking at how environments, how our lifestyle, what we eat, affects that disease risk. By doing that we can hopefully prevent a lot of these diseases from occurring. We're looking at diseases such as dementia, obesity, type 2 diabetes; big global health problems that we know are largely affected by our environment. So it's really trying to pinpoint exactly how the environment increases those risks, enabling us to reduce them and developing better therapies to reduce those disease risks.
Adam - Luke?
Luke - My research is going to be looking at the way people age and the way people have disease through age, in particular looking at how that impacts our cognitive health and our brain health. So what we can do is we can look at people's biological profiles and see if there are any trends in people's aging, and if we can spot trends, perhaps we can give policy advice of what can help healthy aging. Perhaps we can identify things that are causative of neurodegeneration or cognitive problems as people age. So I'm very interested in looking at a variety of samples of tissue, and for example urine and blood samples, and just really trying to understand how people age and how their metabolism changes as people age.
Chris - Because one of the goals is what we call a healthy or a longer health span, rather than just lifespan, because it's more a quality over quantity thing, isn't it? We've become very good in the modern era at making people live for what feels like forever for some people, but unfortunately they spend a significant amount of that time in ill health, and we want to minimise that.
Luke - Yes, exactly. So if we can improve the quality of life - an example would be Alzheimer's disease - if we can understand the disease more and slow down the progression of the disease, people are going to have a higher quality of life for longer, and we can really make real gains in people's enjoyment of life as they age.
Chris - How do you go from a bunch of molecules on a graph, which is what you're going to see from these machines in here, to practical advice for somebody? “You need to eat more bananas, your potassium is a bit low.” How do you actually get those sorts of conclusions out of the cocktail of chemicals that emerge from these machines?
Nicola - It's very complex, and one of the biggest challenges that we will face. So once we've actually identified these molecules, we then would have to validate that. What we would potentially do is to look at that in a more controlled environment. It might potentially be an intervention study where participants would be in a very controlled environment, we would give them a very specific diet and we would look to see how that would affect their metabolites and potentially disease risk markers.
Chris - So you’re hunting through the haystack here, biochemical needles in haystacks, that might be indicative of certain diseases, certain risks for diseases. And then once you think you've found them, then you're going to intervene in people and do a proper controlled experiment. So, “if we change those things, will we change outcomes”.
Luke - Exactly. The initial work that we complete is certainly at the discovery end of the lifespan and as we progress, and as we start to understand more about that metabolism and how metabolic systems are behaving in health and disease, then we can take that forward for validation and for real intervention and make the impact. So, the long term plan is to be involved at all stages of that process.
Adam - Luke Wiley, and before him, Nicola Gray. But it's not just about what goes on inside our bodies. You can point the powerful finger of phenomics at the food we eat too, and explore how different cooking techniques will affect the nutrient quality of a meal.
Ruey - My name is Ruey Leng Loo. I’m the Premier Early to Mid Career Fellow. Currently, we don't know too much about the chemical composition of the food. But when you cook them, how you cook them, that is going to change the chemical composition again. By doing all these different experiments, we can cook it differently before we put it through the instrument, then it can give us the full picture of what’s the kind of vitamin that's lost. If we knew that chemical composition that have a health benefit to us, how we can maintain those molecules, that's what we want to do, really.
Chris - So when I barbecue my chicken leg, we know it has a risk of colon cancer but chemically we don't know why, and you're saying if we feed it into machines like this, we can work out what the molecules are that might be linked to those disease states?
Ruey - Yes, so that we can see what are the changes on the molecule when it’s fresh, when we haven't done anything to that, and compared to if we barbecue the meat, what are the changes on those.
Chris - And going a step further, are you also going to ask the question: if I cook it and eat it, what effect do these different cooking methods have on my biochemistry?
Ruey - That’s right. So because all of us have a very unique set of gut microbacteria, they all have a specific function. If I eat this piece of chicken boiled that’s probably better for me because of my gut bacteria compared to somebody else. That's what we wanted to differentiate so that we can be able to precisely tell you whether you're better off eating a piece of chicken boiled, or maybe a piece of fruit because that would be better for them.
Chris - So you can debunk some of these claims about foods being extra good for you, blueberries being superfoods and all this kind of thing, and actually say “yes, look, I've got plausible scientific evidence” or “actually, this is bunkum, don't waste your money”.
Ruey - Hopefully, that's the goal. We wanted to be able to demythify some of these claims, really.
Chris - And will this enable you to prescribe what actually constitutes a healthy diet one day, then?
Ruey - If I can do that, that would be great. That would be my aim.
Adam - And a laudable and very realistic one at that. So, it's clear we are moving into a very exciting era for medicine. One that will enable us to keep more people living well for longer. But it's also clear that the hard work is only just beginning. We'll leave you with this from Jeremy Nicholson.
Jeremy - Genes and environment come together to create your disease risk. They create you. The things that we want to do is understand how those come together exactly by studying the biochemistry of the body, so that we understand the origins of disease and therefore can inform future healthcare policy. And when somebody becomes ill, we use the same technologies for moving someone from an unhealthy space into a healthy space, which we've already defined by the biochemistry of the whole population.
Related Content
- Previous What's in the water?
- Next Inclusivity in academia










Comments
Add a comment