How should we protect marine ecosystems? We examine the science behind Marine Protected Areas and find out how a new plan could protect the oceans around England. In the news, the sub-zero bacterial ecosystem surviving in an Antarctic lake and fibres inspired by slimy fish. Plus, will mixing spider and human DNA give us superpowers?
In this episode

09:35 - Protecting the Pitcairn Islands
Protecting the Pitcairn Islands
with Josh Reichert & Alistair Gammel, Pew environment group; Enric Sala, National Geographic; Pitcairn Islanders
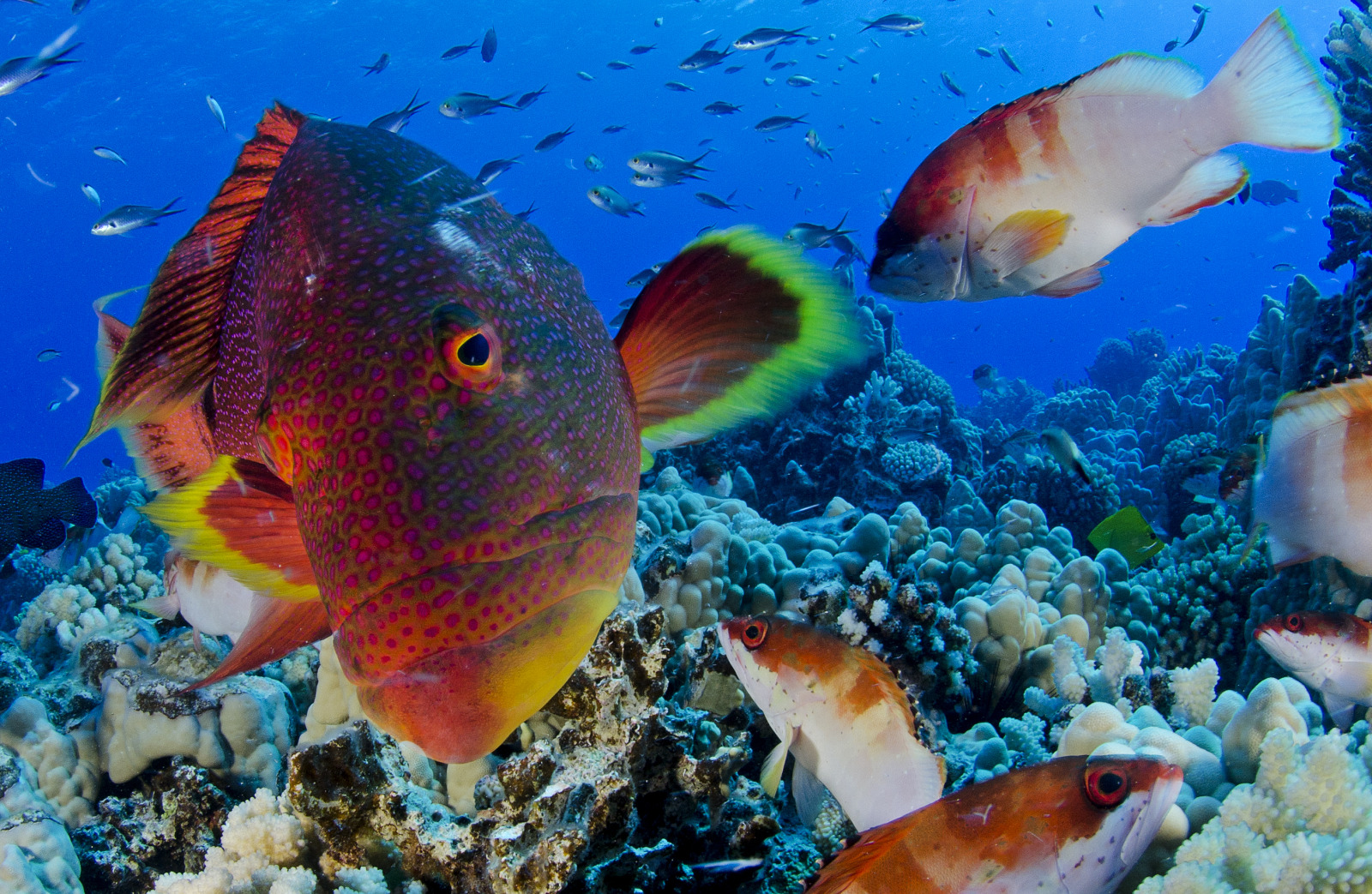
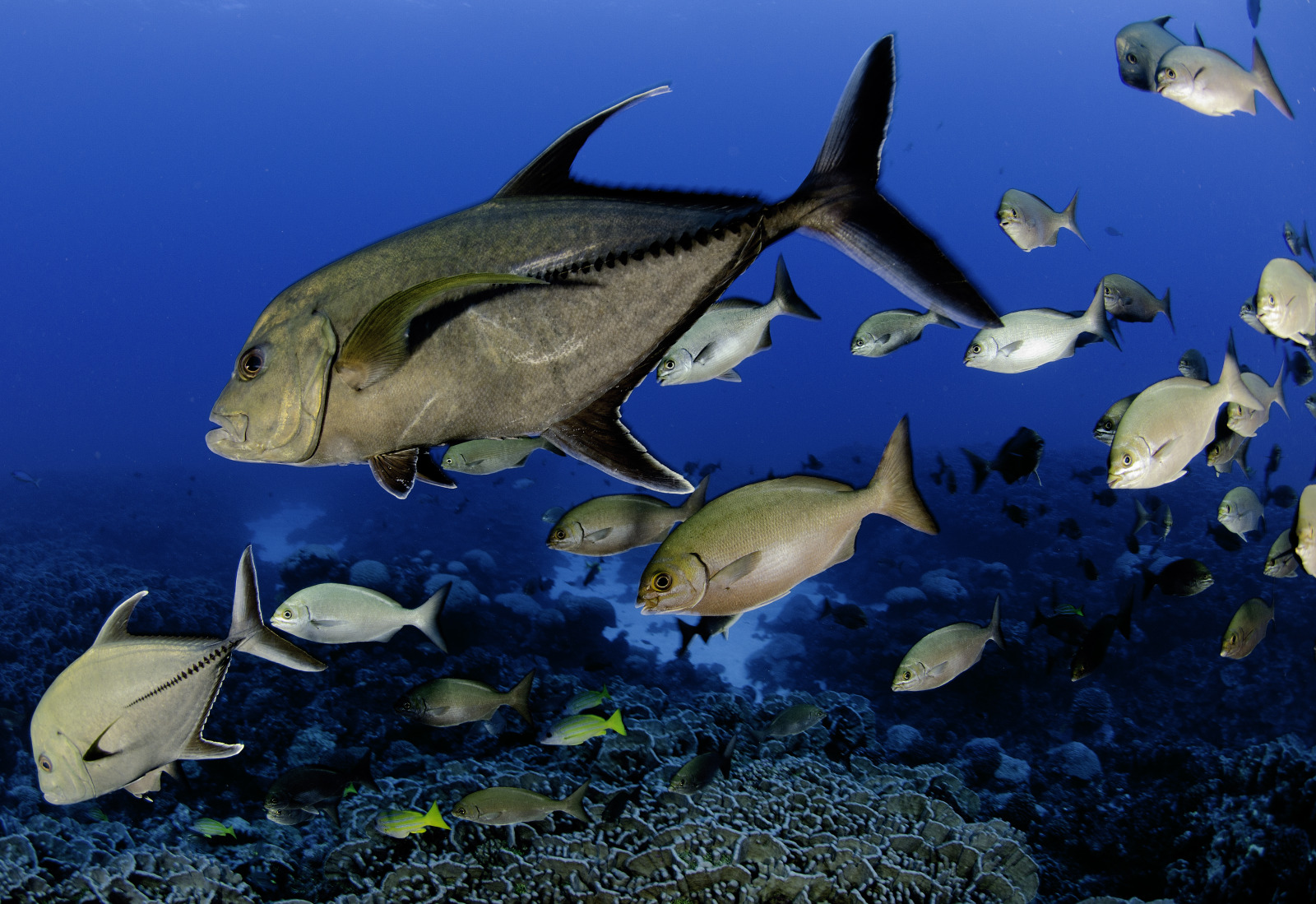
| The following amazing images are taken on or around the Pitcairn Islands.©Enric Sala, National Geographic. | |
|---|---|
 The twinspot snapper is an abundant predator on the pristine reefs of the Pitcairn Islands |  The regional endemic Hotumotua angelfish |
 |  |
 |  |

18:29 - Grand Canyon could be much older than previously thought
Grand Canyon could be much older than previously thought
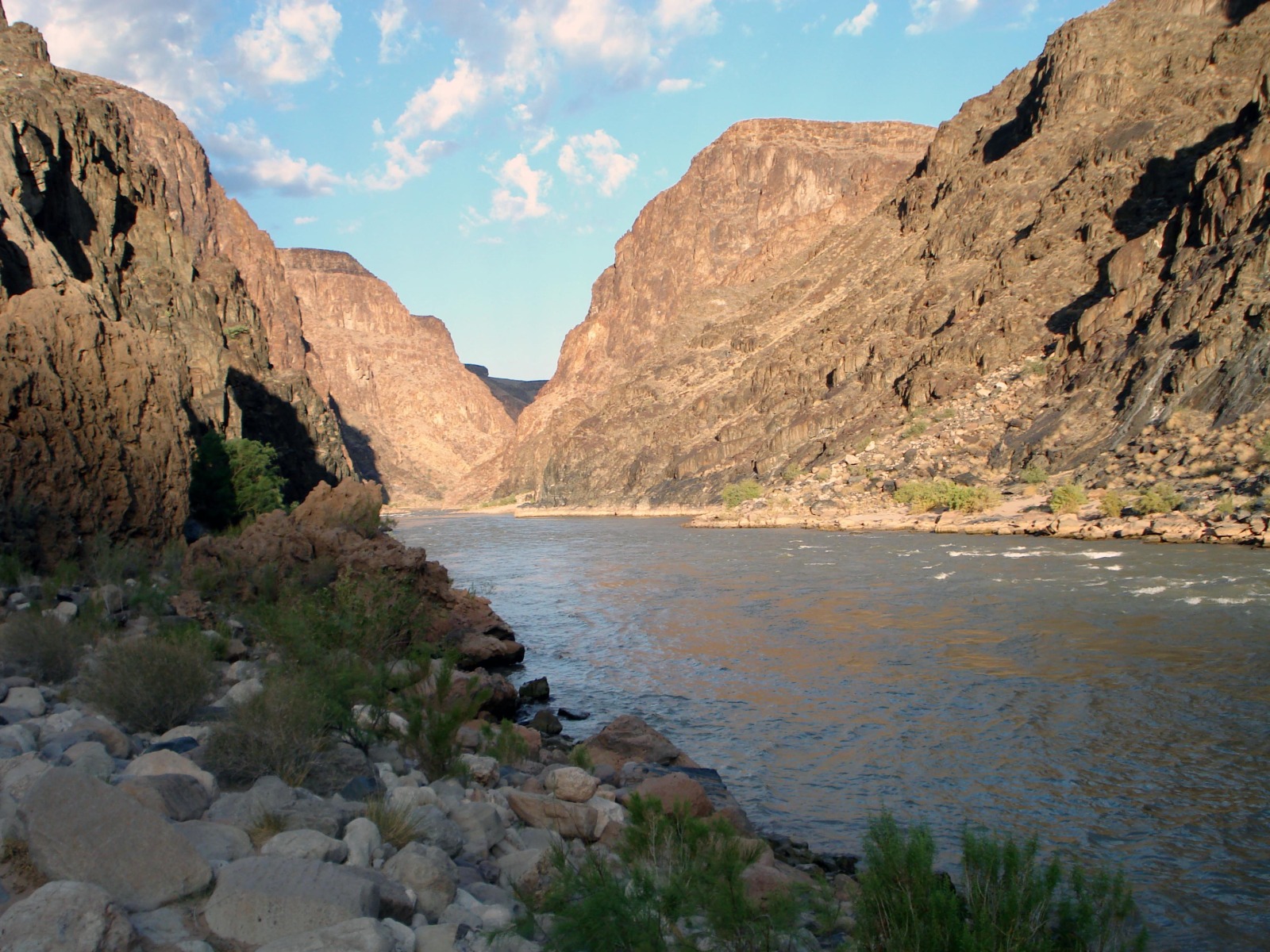
America's Grand Canyon could be a lot older than previously thought - ancient enough that dinosaurs could have strolled down it.
The spectacular Grand Canyon is 277 miles long, up to 18 miles across at its widest point, and a mile deep and until now, it's been generally accepted that it was formed by a river cutting through and eroding rocks around 5 to 6 million years ago.
Remains of phosphate crystals in the canyon push back the estimated formation date to around 70 million years ago.
 That's according to a new study published this week in the journal Science by Rebecca Flowers from the University of Colorado at Boulder and Kenneth A. Farley from Caltech.
That's according to a new study published this week in the journal Science by Rebecca Flowers from the University of Colorado at Boulder and Kenneth A. Farley from Caltech.
Tracking back ancient erosion events is challenging because scientists aren't studying when sedimentary rocks were laid down, but when they were taken away.
The rocks of the Grand Canyon were laid down between 200 million and 2 billion years ago, much of them when this part of the western United States was a shallow tropical sea. A river then cut through exposing those rocks revealing one of the most complete and mind boggling sequences of rocks on earth.
The approach Flowers and Farley took was to analyse the composition of apatite crystals taken from the western end of the Canyon. Uranium and thorium molecules inside those crystals break down radioactively to form helium, which gets locked up inside the crystals as they cool at lower temperatures. As rocks erode and get closer to the earth's surface they cool down and the traces of helium leave a record of that cooling history and so can indicate when the canyon was cut through the rocks.
The team already used this technique to analyse rocks from the eastern end of the canyon, giving an erosion date of 55 million years ago.
And Rebecca Flowers thinks that the canyon was first formed by a predecessor of the Colorado River, that ran in the opposite direction.
These findings don't come without controversy. The age and formation of the Grand Canyon is something that scientists have disagreed about for over 150 years and it doesn't seem like they are going to stop now. There's no doubt that the canyon has a complicated history and that may it well not have formed all at the same time. And this isn't just about figuring out how the Grand Canyon itself was made but studies like this have really important implications for understanding other how landscapes form as scientists get to grips with topography, hydrology and tectonics.
And it shows that even with new techniques emerging there are still immense puzzles about the world that haven't yet been fully unravelled.
With this new idea of when the grand canyon was formed, researchers now want to dig deeper into the question of how it took shape.
22:18 - Overcoming cunning cancer cells
Overcoming cunning cancer cells
Drug resistance is a major problem when it comes to treating cancer...
In many cases, chemotherapy drugs work in the short term, but the tumours become resistant to treatment and keep growing. This is true not only for older drugs but for also for newer, targeted therapies too, and it happens because cancer cells evolve to overcome the effects of the medication.
There's a lot of research going on around the world aimed at understanding and overcoming this resistance. Now a team of European and US researchers, led by Rene Bernards at the Netherlands Cancer Institute, have identified an entirely new mechanism by which cancer cells can develop drug resistance. They've just published their results in the journal Cell.
The researchers started by looking at lung cancer cells carrying a specific genetic fault - a fusion between two genes called EML4 and ALK, which drives them to grow. They used a technique called RNA interference, testing 24,000 tiny fragments of RNA which switch off 8,000 different genes, one at a time, in cancer cells that have been treated with different drugs.
They were searching for cells that had become resistant to the drug treatment, then working out which gene had been switched off for that to happen. And they found one - a gene called MED12 - which was rather unexpected.
MED12 is part of something called the Mediator complex, which helps cells to switch genes on and off. The researchers discovered that switching off MED12 leads to the activation of a pathway called TGF-beta signalling, a kind of molecular escape route that keeps cancer cells growing out of control, even when they've been treated with drugs. The exciting thing is that are drugs that block TGF-beta signalling already in development, so maybe combining them with chemotherapy or targeted cancer treatments could help to tackle drug resistance and make the therapy more effective.
There's still a lot more to figure out about what's going on here, and this is just work on cells growing in the lab and doesn't take into account all the complicated interactions between cancer cells and the tissue around them, which we know are important in helping tumours deveop drug resistance. So it's still a long way from being something that could be turned into immediate benefits for cancer patients.
But it's certainly a significant and exciting finding that takes us a big step closer to understanding how cancers become resistant to therapy, and how it might be possible to overcome it...

35:35 - Becoming a Bat Detective - Planet Earth Online
Becoming a Bat Detective - Planet Earth Online
with Professor Kate Jones, University College London & Charlotte Walters, Zoological Society of London
Christmas is approaching, Halloween is long gone and those black rubber bats have been put away and stored for another year. Even real bats are currently in hibernation - but if you're interested in citizen science and learning more about how these mammals communicate, then stay tuned - as ecologists recently developed the first bat detector software that can be used throughout Europe.
It's called I-BatsID and with it we can all become bat detectives. Planet Earth podcast presenter Sue Nelson met up with Professor Kate Jones, Chair of Ecology and Biodiversity at the University College London, and Charlotte Walters - from the Zoological Society of London - to find out how it works.
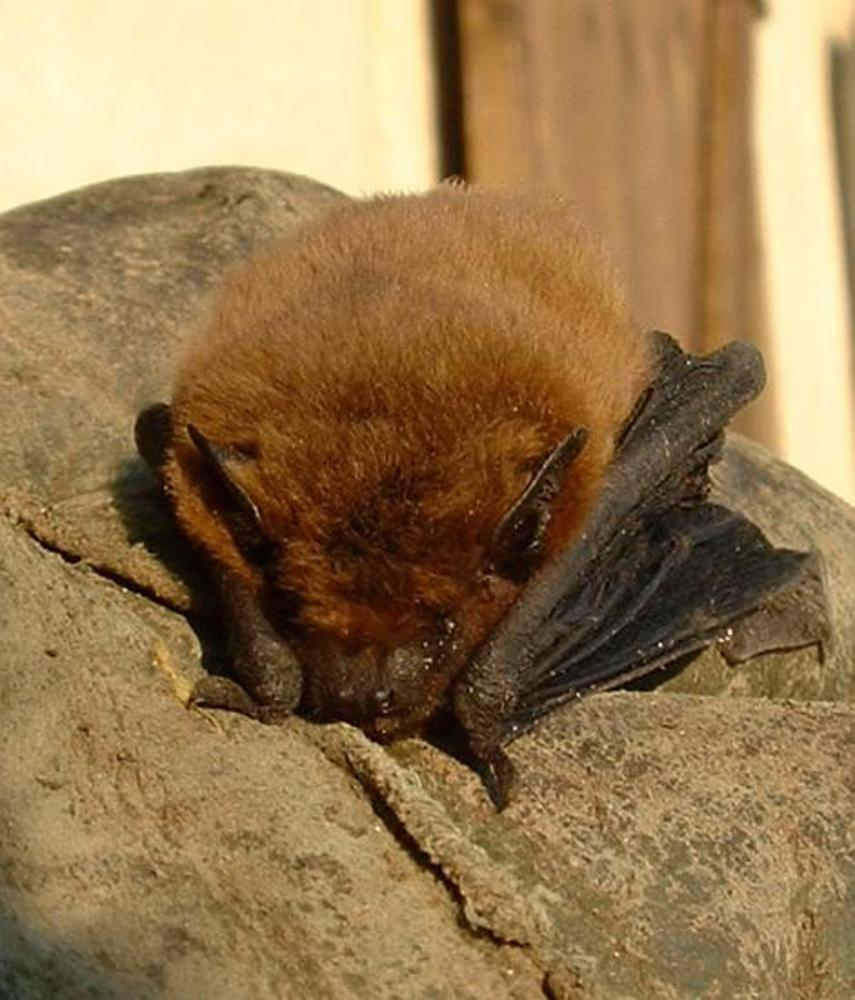 Charlotte - We collected an echolocation call library of calls from species all over Europe collected in lots of different habitats and in different situations and we used this to train the software to identify differences between the different calls. So then when we introduce unknown calls to the software it can tell us which species it is and with a probability of how certain it is.
Charlotte - We collected an echolocation call library of calls from species all over Europe collected in lots of different habitats and in different situations and we used this to train the software to identify differences between the different calls. So then when we introduce unknown calls to the software it can tell us which species it is and with a probability of how certain it is.
Sue - So each species of bat has its own unique identification?
Charlotte - Yes, they all have calls that vary slightly. Some species their calls are really different so we can play you some of the calls - if we play a Rhinolophus call it sounds like this...
Charlotte - And a different type of bat, Pipistrelles has calls which sound like this...
Sue - That sounded more like a bird, cheeping.
Charlotte - These calls are slowed down ten times, so you've got to imagine the actual sound that that is making is ten times faster and ten times higher pitched than these sounds, but you can tell there are differences between the calls that we can use for identification purposes.
Sue - Now, Kate, you are in charge today with your finger on the mouse playing us some of those calls, you can actually see on your computer screen what the echo looks like as well.
Kate - What we're using here is transforming the actual raw data, the raw sounds, so we're using an FFT transfer, so fast fourier transform of the data, which takes tiny bits out of each of the raw data so it becomes a much simpler signal. It turns out that when you do that with bat calls they tend to have a characteristic shape. The ones that your heard before, the kind of ee, ee, that were a constant frequency, they look very flat. But when you have calls which have got a more chirpy sound they have bits of frequency modulation, so they sound more chirpy, so that's where you get a kind of a vertical line straight down and then you may get a bit of constant frequency at the end.
So, if I just play these again you can hear that... And then these are the flat ones...
There are other calls that bats make as well. Some of them do searching calls which we've been showing you.
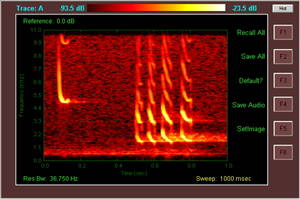 These are social calls which they make to attract mates to communicate with each other, which are more trill-like. You also get bats which when they're searching they'll emit a search phase call and get the echo back so that they can figure out where they are where their food is, but when they get closer to an insect or an object they need much more information about it. So they will do a feeding buzz. On some detectors it sounds like someone blowing a raspberry, but I will just play it for real...
These are social calls which they make to attract mates to communicate with each other, which are more trill-like. You also get bats which when they're searching they'll emit a search phase call and get the echo back so that they can figure out where they are where their food is, but when they get closer to an insect or an object they need much more information about it. So they will do a feeding buzz. On some detectors it sounds like someone blowing a raspberry, but I will just play it for real...
Sue - That sounds like a budgerigar! And there is now a way, a citizen science project, that people can help identify these bats as well.
Kate - We have all these volunteers that go out and collect data in a monitoring programme called iBats and we get the data back and we have over two million files that they send us. So what we try and do is go through it with some automatic filters but those filters aren't very good at the moment, so what we're trying to do is develop new ways of going through them automatically and trying to link it up to iBatsID so that it's all one tool.
At the moment, you go through and you find a call, you take the parameters and then you feed it into the software ID tool, but actually what we want to do is to integrate that all together. The first step on that is to look at all the files and try to label all that data.
We have launched a new site called
Bat Detective which anyone can log on and help us look at bats. So, you go onto the site, sign in, and then you can help us classify where these calls are and hunt for these calls in the recordings...

55:02 - Can we mix human and spider DNA?
Can we mix human and spider DNA?
Hannah Critchlow untangled this sticky question...
Hannah - This week we wonder if we can make a real life Spiderman, and Enriques got in touch with this question...
Enriques - If we put the DNA from a spider into a human what would happen.
Hannah - So can you successfully mix spider and human DNA and what could you create. If so we turn to a genetics professor working where Dolly the sheep was created.
Helen - Hello my name is Helen Sang; I work at the Roslin Institute which is part of the University of Edinburgh. It would be relatively easy to introduce spider genes into humans by adapting the methods we used to make genetically modified mice, for example, and combining this with the techniques used in in vitro fertilization - IVF - that are used to make test tube babies. We can introduce genes into specific tissues or organs; this is the basis of gene therapy. This is very new technology and it's only just becoming licensed as a medical treatment. And here we would introduce a gene into a specific tissue, for example muscle, and repair a mutation.
Hannah - So it is possible to mix the DNA of humans and spiders. For this scenario, let's ignore the strict regulations in place to ensure these methods are used responsibly, and ask instead could we actually make a real life Spiderman?
Helen - Spiderman had a lot of characteristics of a superhuman that he was hyperactive and an elite athlete: many characteristics that individuals have some of, but nobody has all these different characteristics. We have people who are sprinters, or people who are very energetic, but Spider man pulls a lot of these characteristics together and at the moment human geneticists are identifying variations in our genes that lead to these differences. So maybe Spiderman has just got the elite versions of many of these genes. We could think about, in the longer term, genetically modifying people so that they could sprint and were very active and very intelligent, but that would require us to make a lot of changes by genetic modification. And not only is that illegal but I think that's a bit too much of a challenge for today...
Hannah - And researchers have genetically tweaked a goat to produce spider silk protein in its milk. So maybe we could purify this goat's milk to produce a Spiderman Web shooter. Joseph Katz added via Facebook: "the trait that allows spiders to climb walls has a lot to do with their low mass to surface area ratio. Dense humans might be better off taking inspiration from geckos, whose wall-sticking ability is due to millions of microscopic Velcro-like hair pads on their feet. Now, sticking with the genre of genetics, we glide into this question:
Louis - Hi my name's Louis. I'm 15 and I come from London. I was just wondering. In foetuses, the cells multiply really quickly; but I've never heard of one having cancer. Is it possible?
Hannah - So can fetuses become cancerous? What do think about that one? Let us know by posting on the Naked scientist's Facebook page. You can e-mail Chris at the naked scientist dot com. You can join in the debate live on our forum which is that Naked Scientist stock com slash forum or you can tweet at Naked Scientists.










Comments
Have yall tried to put spider
Have yall tried to put spider DNA into human body?
Add a comment